-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Archaea in and on the Human Body: Health Implications and Future Directions
article has not abstract
Published in the journal: . PLoS Pathog 11(6): e32767. doi:10.1371/journal.ppat.1004833
Category: Pearls
doi: https://doi.org/10.1371/journal.ppat.1004833Summary
article has not abstract
Although they are abundant and even dominant members of animal microbiomes (microbiotas), from sponges and termites to mice and cattle, archaea in our own microbiomes have received much less attention than their bacterial counterparts. The fact that human-associated archaea have been relatively little-studied may be at least partially attributed to the lack of any established archaeal human pathogens [1,2]. Clinically oriented microbiology courses often do not mention archaea at all, and most medical school and biology students are only aware of archaea as exotic extremophiles that have strange and eukaryotic-like molecular machinery. Since archaea have been known to be associated with the human gut for several decades, one would think that human microbiome studies may unravel new facets of archaea–human interactions. However, adequate universal primers that amplify both bacterial and archaeal small 16S rRNA genes but not any host rRNA genes were only published in mid-2011 [3], and thus, many studies chose to focus on bacteria alone rather than multiply effort and expense to cover taxa that are considered secondary in importance, if not altogether rare. Here, we provide a brief overview of what is currently known about archaea in and on the human body and their potential effects on human health (for additional reviews on archaea and their potential involvement in human disease, see [4–8]).
Archaea in the Human Gut
The human large intestine (colon), in healthy individuals, has extremely low oxygen concentrations, and over 90% of its microbiota are strict anaerobes. Researchers taking metagenomic fecal microbiota surveys of adult Europeans could assign about 0.8% of the genes in their dataset to archaea [9], and similar numbers (0.2%–0.3%) were reported for Amerindians and Malwaians [10], while North Americans had much lower fractions (<0.05%). With the exception of a single report indicating the presence of halophilic archaea in biopsies of inflammatory bowel disease patients [11], archaea that reside in the human colon are nearly always methanogens. Most of these strict anaerobes belong to the order Methanobacteriales (Fig 1), the most common genera being the closely related Methaonbrevibacter and Methanosphaera. Methanobrevibacter (previously called Methanobacterium) was first isolated from human stool as early as 1968 [12], followed nearly 15 years later by the discovery that such fecal isolates belonged to the species Methanobrevibacter smithii. M. smithii has been shown to be present in up to 95.7% of human subjects [13], and to be the most abundant methanogen in the human gut by several studies, comprising up to as much as 10% of all anaerobes found in a healthy individual's colon [14–16]. Remarkably, its abundance appears to remain stable over time, even following radical dietary changes [17], and it is highly heritable, meaning that monozygotic twins are more concordant for its presence, or absence, than dizygotic twins [18,19]. Importantly, substrates for methanogenesis, such as H2, methanol and acetate, are mostly derived from the end products of bacterial fermentation. The second most abundant methanogen in this environment is Methanosphaera stadtmanae. This organism, which has the most restricted energy metabolism of all known methanogenic archaea, is totally dependent on acetate as a carbon source, and its methane production requires methanol and hydrogen [20]. The human colon and other mammalian intestines are dominated by hundreds of bacterial species [14], and it is therefore not surprising to observe that the genomes of M. smithii and M. stadtmanae appear to be very rich in inter-domain lateral gene transfers, especially relating to glycosyltransferases and ABC transporters in both species and adhesin-like proteins in M. smithii [21,22]. These laterally acquired genes are thought to have played a significant role in these organisms' initial adaptation to mammalian hosts [21,22]. Both of these species have been recently shown to induce monocyte-derived dendritic cell maturation, and M. stadtmanae also induced a strong pro-inflammatory cytokine release from these cells [23] and is more prevalent in patients with inflammatory bowel disease [24].
Fig. 1. The distribution of human-associated archaea in the phylogenetic tree of the domain Archaea. 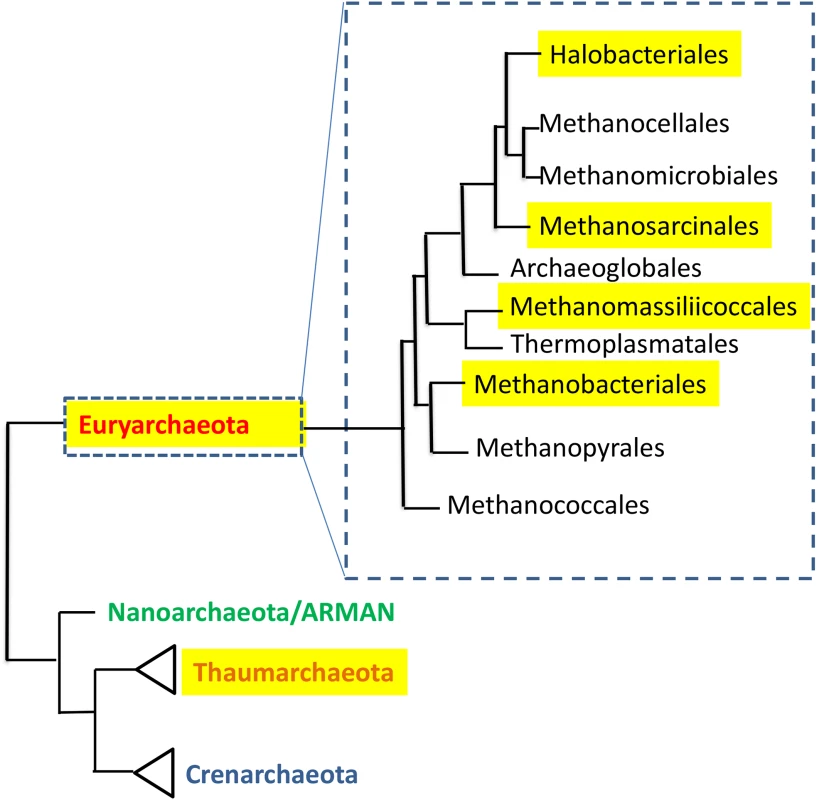
Tree is based on [63], and [64]. Highlighted are groups that contain human-associated members. In ruminants, presence of the methanogen Methanobrevibacter ruminantium can result in loss of up to 6% of all ingested energy [25]. In contrast, it has been suggested, based primarily on mouse studies [26,27], that gut methanogens contribute to human obesity. Indeed, methanogens are capable of syntrophic interactions with bacteria that enhance production of short-chain fatty acids, which provide a considerable caloric contribution to the host. However, more recent evidence from several large human studies strongly supports an association of M. smithii with leanness [19,28–30]. Future research may determine more precisely the roles that methanogens play in host metabolism in order to enable new microbiota-based approaches for weight management.
Another possible connection between gut methanogens and human health is the strong association between methanogen presence and chronic constipation [31]. Methane was shown to slow intestinal transit time by 59% [32], and thus may contribute substantially to constipation. However, a shorter intestinal transit time probably selects against the presence of methanogens, since they tend to have generation times that are longer than those of many gut bacteria, even when grown in the most favorable, state-of-the-art culture media [33]. In agreement with these in vitro data, human studies have shown a lower prevalence of methanogens (determined by methane excretion) in patients that tend to have diarrhea episodes (such as those with inflammatory bowel diseases) compared to healthy individuals [34]. Taken together, these findings indicate that in individuals with already slow intestinal transit, methanogens may bloom and promote further constipation.
Until recently, there were only six known orders of methanogens, only one of which (the above-mentioned Methanobacteriales) was represented in the human body. However, recent reports show that there is a seventh order, Methanomassiliicoccales, which includes the gut-residing methanogens Methanomassiliicoccus luminyensis and Candidatus Methanomethylophilus alvus [35]. Both these organisms are methylotrophic archaea isolated from human feces [36,37]. Although M. luminyensis has been shown to require H2 for methanogesis, it can utilize not just methanol but also trimethylamine for growth in the presence of H2, which has important implications for human health [38]. Trimethylamine is a metabolite produced from host dietary choline [39] or carnitine [40], by the gut microbiota, which is later oxidized to trimethylamine N-oxide (TMAO) by the host enzyme flavin monooxygenases. TMAO has been shown to promote atherosclerosis in mice and to be a strong biomarker for human cardiovascular disease [39]. Thus, having an archaeal community that may remediate not just methanol toxicity, but also prevent trimethylamine accumulation and TMAO production could be highly desirable. Accordingly it has been suggested that these archaea can be used as "archaebiotics," for the prevention of cardiovascular disease as well as trimethylaminuria, a hereditary deficiency in flavin monooxygenases activity that results in an unpleasant fishy odor in breath and sweat [38]. Whether this novel and exciting therapeutic concept can be tested in animals and subsequently translated into the clinic remains to be seen.
Subgingival Archaea
Methanogenic Archaea have been reported in subgingival dental plaque as early as 1987 [41]. To date, three genera have been successfully isolated from subgingival plaque: Methanobrevibacter [42,43], Methanosphaera (based on weak antigenic similarity) [44], and Methanosarcina (based on physiology and staining) [45]. Additionally, 16S rRNA gene amplicon sequencing studies detected archaea related to Thermoplasmata [46,47] (which, in retrospect, probably belong to the seventh order of methanogens; see gut methanogens section, above), as well as members of the Methanobacterium genus [48,49]. In general, it appears that the genetic diversity of archaea of the human subgingival dental plaques is low, much as is the case for the gut methanogens, and that Methanobrevibacter oralis is by far the most prevalent methanogen found in this environment. In a recent review, Nguyen-Hieu et al. pooled the data from several studies of methanogens in the oral cavity and concluded that M. oralis is significantly associated with periodontal disease both in terms of abundance comparisons between patients and controls and between diseases and healthy sites within the same patient [50]. Furthermore, they concluded that indirect evidence supports the contribution of that methanogen to periodontal disease and that this contribution likely stems from syntrophic interactions with sulfate-reducing bacteria. Thus, a mixed infection may be required for a direct causal demonstration of the pathogenic contribution of M. oralis in an animal model of periodontal disease. Unlike many antibiotics that do not target archaea (because they do not have a peptidoglycan cell wall and have ribosomes that are more eukaryotic like [51]), metronidazole, which is commonly used to treat periodontitis, is highly effective against M. oralis [52] and, thus, suppression of M. oralis could contribute to its efficacy [50]. Statins that inhibit the activity of 3-hydroxy-3-methylglutaryl coenzyme A reductases of eukaryotes and archaea lower blood cholesterol in humans, but they also effectively inhibit archaeal growth because they block the synthesis of their main membrane lipids [53]. If indeed M. oralis is a "co-pathogen," it would be interesting to examine the effects of "archaea-specific" drugs such as statins on periodontal disease, for example, by examining the periodontal pockets of patients who have recently been prescribed statins before and after several months of statin use.
Archaea on the Human Skin
Archaea on the human skin have been discovered only in recent years. A 16S rRNA gene amplicon sequencing study focusing on the navel found rare occurrence of Methanobrevibacter in several individuals. Even more rare were phylotypes belonging to the halophilic archaea (family Halobacteriaceae), which were only present in a single individual, who abstained from showers or baths for several years prior to sampling [54]. A large metagenomic survey detected reads that matched archaea in most individual samples, but all archaeal sequences combined did not exceed 2.3 × 10–5 of the reads in any sample [55]. A recent study, using archaeal-specific 16S rRNA gene primers, found archaea to be present on the skin of 13/13 volunteers, with relative abundances that exceeded 4% in one individual. Five out of five individuals that were more closely studied displayed human-associated archaea that were not methanogens, as may be expected in such an aerobic niche. Instead, the dominant skin-associated archaea belonged to the phylum Thaumarchaeota [56]. In a study that continuously sampled the skin (left and right palm) of one male and one female over several months, the male had only transient Thaumarchaeota, while the female had persistent, albeit low, presence of these archaea on her right palm [57], indicating there is likely to be high inter-individual variation in skin colonization by these archaea. Like other members of the Thaumarchaeota phylum, skin phylotypes are thought to be chemolithotrophic ammonium oxidizers and encode characteristic amoA gene homologs [56]. Whether the relatively small amounts of ammonium in sweat are sufficient to sustain such metabolism in the human skin is unclear, but ammonium release in sweat was shown to increase during physical exercise [58] and could reach several mM [59]. Thus, people who sweat and/or exercise more could harbor larger communities of these archaea.
Concluding Remarks
The availability of reference genomes from previously unrepresented groups, such as the Methanomassiliicoccales for metagenomic analysis, as well as better 16S rRNA gene primers, should improve the detection of archaea in human microbiome studies. This improvement is highly timely, since archaea are still an under-detected and little-studied part of the human microbiome, and their contributions to human health or disease remain mostly unknown. This knowledge gap should be addressed in the near future to inform clinicians, many of whom are totally unaware of these organisms. While no human clinical study studying the in vivo effects of statins on archaea in our microbiomes has been published, in vitro results [60] strongly suggest that these drugs could inhibit the growth of archaea in the human body. While the inhibitory concentrations reported for archaea in vitro (4 mg/L, about 10 μmol/L for lovastatin [60]) are much higher than their level in circulation (9.4 nmol/L [61]), their levels in the gut may be very much higher. Moreover, in highly competitive niches, such as the colon, even partial growth inhibition may cause extinction. In developed countries, such as the United States, statin use is on the rise, and over a third of people over 65 use these drugs for their cholesterol-lowering effects, unaware that at the same time they are taking a broad-spectrum anti-archaeal agent. At the moment, there is little evidence of whether eradication of human-associated archaea (and potentially their bacterial syntrophs) will be beneficial or harmful for human health, with the possible exception of periodontal disease. Thus, before archaea become part of the "disappearing human microbiota" [62] we should at least know if we are going to miss them when they are gone.
Zdroje
1. Gill EE, Brinkman FS (2011) The proportional lack of archaeal pathogens: Do viruses/phages hold the key? Bioessays 33 : 248–254. doi: 10.1002/bies.201000091 21328413
2. Cavicchioli R, Curmi PM, Saunders N, Thomas T (2003) Pathogenic archaea: do they exist? Bioessays 25 : 1119–1128. 14579252
3. Walters WA, Caporaso JG, Lauber CL, Berg-Lyons D, Fierer N, et al. (2011) PrimerProspector: de novo design and taxonomic analysis of barcoded polymerase chain reaction primers. Bioinformatics 27 : 1159–1161. doi: 10.1093/bioinformatics/btr087 21349862
4. Eckburg PB, Lepp PW, Relman DA (2003) Archaea and their potential role in human disease. Infect Immun 71 : 591–596. 12540534
5. Horz HP, Conrads G (2010) The discussion goes on: What is the role of Euryarchaeota in humans? Archaea 2010 : 967271. doi: 10.1155/2010/967271 21253553
6. Gaci N, Borrel G, Tottey W, O'Toole PW, Brugere JF (2014) Archaea and the human gut: new beginning of an old story. World J Gastroenterol 20 : 16062–16078. doi: 10.3748/wjg.v20.i43.16062 25473158
7. Dridi B, Raoult D, Drancourt M (2011) Archaea as emerging organisms in complex human microbiomes. Anaerobe 17 : 56–63. doi: 10.1016/j.anaerobe.2011.03.001 21420503
8. Dridi B (2012) Laboratory tools for detection of archaea in humans. Clin Microbiol Infect 18 : 825–833. doi: 10.1111/j.1469-0691.2012.03952.x 22897827
9. Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, et al. (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464 : 59–65. doi: 10.1038/nature08821 20203603
10. Yatsunenko T, Rey FE, Manary MJ, Trehan I, Dominguez-Bello MG, et al. (2012) Human gut microbiome viewed across age and geography. Nature 486 : 222–227. doi: 10.1038/nature11053 22699611
11. Oxley AP, Lanfranconi MP, Wurdemann D, Ott S, Schreiber S, et al. (2010) Halophilic archaea in the human intestinal mucosa. Environ Microbiol 12 : 2398–2410. doi: 10.1111/j.1462-2920.2010.02212.x 20438582
12. Nottingham PM, Hungate RE (1968) M. J Bacteriol 96 : 2178–2179. 4881707
13. Dridi B, Henry M, El Khechine A, Raoult D, Drancourt M (2009) High prevalence of Methanobrevibacter smithii and Methanosphaera stadtmanae detected in the human gut using an improved DNA detection protocol. PLoS One 4: e7063. doi: 10.1371/journal.pone.0007063 19759898
14. Eckburg PB, Bik EM, Bernstein CN, Purdom E, Dethlefsen L, et al. (2005) Diversity of the human intestinal microbial flora. Science 308 : 1635–1638. 15831718
15. Samuel BS, Hansen EE, Manchester JK, Coutinho PM, Henrissat B, et al. (2007) Genomic and metabolic adaptations of Methanobrevibacter smithii to the human gut. Proc Natl Acad Sci U S A 104 : 10643–10648. 17563350
16. Miller TL, Wolin MJ, de Macario EC, Macario AJ (1982) Isolation of Methanobrevibacter smithii from human feces. Appl Environ Microbiol 43 : 227–232. 6798932
17. Miller TL, Wolin MJ (1983) Stability of Methanobrevibacter smithii populations in the microbial flora excreted from the human large bowel. Appl Environ Microbiol 45 : 317–318. 6824322
18. Hansen EE, Lozupone CA, Rey FE, Wu M, Guruge JL, et al. (2011) Pan-genome of the dominant human gut-associated archaeon, Methanobrevibacter smithii, studied in twins. Proc Natl Acad Sci U S A 108 Suppl 1 : 4599–4606. doi: 10.1073/pnas.1000071108 21317366
19. Goodrich JK, Waters JL, Poole AC, Sutter JL, Koren O, et al. (2014) Human genetics shape the gut microbiome. Cell 159 : 789–799. doi: 10.1016/j.cell.2014.09.053 25417156
20. Fricke WF, Seedorf H, Henne A, Kruer M, Liesegang H, et al. (2006) The genome sequence of Methanosphaera stadtmanae reveals why this human intestinal archaeon is restricted to methanol and H2 for methane formation and ATP synthesis. J Bacteriol 188 : 642–658. 16385054
21. Lurie-Weinberger MN, Peeri M, Tuller T, Gophna U (2012) Extensive Inter-Domain Lateral Gene Transfer in the Evolution of the Human Commensal Methanosphaera stadtmanae. Front Genet 3 : 182. doi: 10.3389/fgene.2012.00182 23049536
22. Lurie-Weinberger MN, Peeri M, Gophna U (2012) Contribution of lateral gene transfer to the gene repertoire of a gut-adapted methanogen. Genomics 99 : 52–58. doi: 10.1016/j.ygeno.2011.10.005 22056789
23. Bang C, Weidenbach K, Gutsmann T, Heine H, Schmitz RA (2014) The intestinal archaea Methanosphaera stadtmanae and Methanobrevibacter smithii activate human dendritic cells. PLoS One 9: e99411. doi: 10.1371/journal.pone.0099411 24915454
24. Blais Lecours P, Marsolais D, Cormier Y, Berberi M, Hache C, et al. (2014) Increased prevalence of Methanosphaera stadtmanae in inflammatory bowel diseases. PLoS One 9: e87734. doi: 10.1371/journal.pone.0087734 24498365
25. Leahy SC, Kelly WJ, Altermann E, Ronimus RS, Yeoman CJ, et al. (2010) The genome sequence of the rumen methanogen Methanobrevibacter ruminantium reveals new possibilities for controlling ruminant methane emissions. PLoS One 5: e8926. doi: 10.1371/journal.pone.0008926 20126622
26. Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, et al. (2006) An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444 : 1027–1031. 17183312
27. Samuel BS, Gordon JI (2006) A humanized gnotobiotic mouse model of host-archaeal-bacterial mutualism. Proc Natl Acad Sci U S A 103 : 10011–10016. 16782812
28. Million M, Angelakis E, Maraninchi M, Henry M, Giorgi R, et al. (2013) Correlation between body mass index and gut concentrations of Lactobacillus reuteri, Bifidobacterium animalis, Methanobrevibacter smithii and Escherichia coli. Int J Obes (Lond) 37 : 1460–1466. doi: 10.1038/ijo.2013.20 23459324
29. Le Chatelier E, Nielsen T, Qin J, Prifti E, Hildebrand F, et al. (2013) Richness of human gut microbiome correlates with metabolic markers. Nature 500 : 541–546. doi: 10.1038/nature12506 23985870
30. Schwiertz A, Taras D, Schafer K, Beijer S, Bos NA, et al. (2010) Microbiota and SCFA in lean and overweight healthy subjects. Obesity (Silver Spring) 18 : 190–195. doi: 10.1038/oby.2009.167 19498350
31. Pimentel M, Gunsalus RP, Rao SS, Zhang H (2012) Methanogens in Human Health and Disease. Am J Gastroenterol Suppl 6 : 28–33.
32. Pimentel M, Lin HC, Enayati P, van den Burg B, Lee HR, et al. (2006) Methane, a gas produced by enteric bacteria, slows intestinal transit and augments small intestinal contractile activity. Am J Physiol Gastrointest Liver Physiol 290: G1089–1095. 16293652
33. Khelaifia S, Raoult D, Drancourt M (2013) A versatile medium for cultivating methanogenic archaea. PLoS One 8: e61563. doi: 10.1371/journal.pone.0061563 23613876
34. McKay LF, Eastwood MA, Brydon WG (1985) Methane excretion in man—a study of breath, flatus, and faeces. Gut 26 : 69–74. 3965369
35. Borrel G, O'Toole PW, Harris HM, Peyret P, Brugere JF, et al. (2013) Phylogenomic data support a seventh order of Methylotrophic methanogens and provide insights into the evolution of Methanogenesis. Genome Biol Evol 5 : 1769–1780. doi: 10.1093/gbe/evt128 23985970
36. Borrel G, Harris HM, Tottey W, Mihajlovski A, Parisot N, et al. (2012) Genome sequence of "Candidatus Methanomethylophilus alvus" Mx1201, a methanogenic archaeon from the human gut belonging to a seventh order of methanogens. J Bacteriol 194 : 6944–6945. doi: 10.1128/JB.01867-12 23209209
37. Gorlas A, Robert C, Gimenez G, Drancourt M, Raoult D (2012) Complete genome sequence of Methanomassiliicoccus luminyensis, the largest genome of a human-associated Archaea species. J Bacteriol 194 : 4745. doi: 10.1128/JB.00956-12 22887657
38. Brugere JF, Borrel G, Gaci N, Tottey W, O'Toole PW, et al. (2014) Archaebiotics: proposed therapeutic use of archaea to prevent trimethylaminuria and cardiovascular disease. Gut Microbes 5 : 5–10. doi: 10.4161/gmic.26749 24247281
39. Wang Z, Klipfell E, Bennett BJ, Koeth R, Levison BS, et al. (2011) Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472 : 57–63. doi: 10.1038/nature09922 21475195
40. Koeth RA, Wang Z, Levison BS, Buffa JA, Org E, et al. (2013) Intestinal microbiota metabolism of L-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat Med 19 : 576–585. doi: 10.1038/nm.3145 23563705
41. Brusa T, Conca R, Ferrara A, Ferrari A, Pecchioni A (1987) The presence of methanobacteria in human subgingival plaque. J Clin Periodontol 14 : 470–471. 3308971
42. Ferrari A, Brusa T, Rutili A, Canzi E, Biavati B (1994) Isolation and characterization ofMethanobrevibacter oralis sp. nov. Current Microbiology 29 : 7–12.
43. Kulik EM, Sandmeier H, Hinni K, Meyer J (2001) Identification of archaeal rDNA from subgingival dental plaque by PCR amplification and sequence analysis. FEMS Microbiol Lett 196 : 129–133. 11267768
44. Belay N, Johnson R, Rajagopal BS, Conway de Macario E, Daniels L (1988) Methanogenic bacteria from human dental plaque. Appl Environ Microbiol 54 : 600–603. 3355146
45. Robichaux M, Howell M, Boopathy R (2003) Methanogenic activity in human periodontal pocket. Curr Microbiol 46 : 53–58. 12432465
46. Li CL, Liu DL, Jiang YT, Zhou YB, Zhang MZ, et al. (2009) Prevalence and molecular diversity of Archaea in subgingival pockets of periodontitis patients. Oral Microbiol Immunol 24 : 343–346. doi: 10.1111/j.1399-302X.2009.00514.x 19572899
47. Horz HP, Seyfarth I, Conrads G (2012) McrA and 16S rRNA gene analysis suggests a novel lineage of Archaea phylogenetically affiliated with Thermoplasmatales in human subgingival plaque. Anaerobe 18 : 373–377. doi: 10.1016/j.anaerobe.2012.04.006 22561061
48. Matarazzo F, Ribeiro AC, Feres M, Faveri M, Mayer MP (2011) Diversity and quantitative analysis of Archaea in aggressive periodontitis and periodontally healthy subjects. J Clin Periodontol 38 : 621–627. doi: 10.1111/j.1600-051X.2011.01734.x 21539593
49. Faveri M, Goncalves LF, Feres M, Figueiredo LC, Gouveia LA, et al. (2011) Prevalence and microbiological diversity of Archaea in peri-implantitis subjects by 16S ribosomal RNA clonal analysis. J Periodontal Res 46 : 338–344. doi: 10.1111/j.1600-0765.2011.01347.x 21338359
50. Nguyen-Hieu T, Khelaifia S, Aboudharam G, Drancourt M (2013) Methanogenic archaea in subgingival sites: a review. APMIS 121 : 467–477. doi: 10.1111/apm.12015 23078250
51. Allers T, Mevarech M (2005) Archaeal genetics—the third way. Nat Rev Genet 6 : 58–73. 15630422
52. Dridi B, Fardeau ML, Ollivier B, Raoult D, Drancourt M (2011) The antimicrobial resistance pattern of cultured human methanogens reflects the unique phylogenetic position of archaea. J Antimicrob Chemother 66 : 2038–2044. doi: 10.1093/jac/dkr251 21680581
53. Lam WL, Doolittle WF (1989) Shuttle vectors for the archaebacterium Halobacterium volcanii. Proc Natl Acad Sci U S A 86 : 5478–5482. 2748598
54. Hulcr J, Latimer AM, Henley JB, Rountree NR, Fierer N, et al. (2012) A jungle in there: bacteria in belly buttons are highly diverse, but predictable. PLoS One 7: e47712. doi: 10.1371/journal.pone.0047712 23144827
55. Oh J, Byrd AL, Deming C, Conlan S, Kong HH, et al. (2014) Biogeography and individuality shape function in the human skin metagenome. Nature 514 : 59–64. doi: 10.1038/nature13786 25279917
56. Probst AJ, Auerbach AK, Moissl-Eichinger C (2013) Archaea on human skin. PLoS One 8: e65388. doi: 10.1371/journal.pone.0065388 23776475
57. Caporaso JG, Lauber CL, Costello EK, Berg-Lyons D, Gonzalez A, et al. (2011) Moving pictures of the human microbiome. Genome Biol 12: R50. doi: 10.1186/gb-2011-12-5-r50 21624126
58. Czarnowski D, Gorski J (1991) Sweat ammonia excretion during submaximal cycling exercise. J Appl Physiol (1985) 70 : 371–374.
59. Sato K, Kang WH, Saga K, Sato KT (1989) Biology of sweat glands and their disorders. I. Normal sweat gland function. Journal of the American Academy of Dermatology 20 : 537–563. 2654204
60. Miller TL, Wolin MJ (2001) Inhibition of growth of methane-producing bacteria of the ruminant forestomach by hydroxymethylglutaryl-SCoA reductase inhibitors. J Dairy Sci 84 : 1445–1448. 11417704
61. Bjorkhem-Bergman L, Lindh JD, Bergman P (2011) What is a relevant statin concentration in cell experiments claiming pleiotropic effects? Br J Clin Pharmacol 72 : 164–165. doi: 10.1111/j.1365-2125.2011.03907.x 21223360
62. Blaser MJ, Falkow S (2009) What are the consequences of the disappearing human microbiota? Nat Rev Microbiol 7 : 887–894. doi: 10.1038/nrmicro2245 19898491
63. Spang A, Martijn J, Saw JH, Lind AE, Guy L, et al. (2013) Close Encounters of the Third Domain: The Emerging Genomic View of Archaeal Diversity and Evolution. Archaea 2013 : 12.
64. Borrel G, Parisot N, Harris HM, Peyretaillade E, Gaci N, et al. (2014) Comparative genomics highlights the unique biology of Methanomassiliicoccales, a Thermoplasmatales-related seventh order of methanogenic archaea that encodes pyrrolysine. BMC Genomics 15 : 679. doi: 10.1186/1471-2164-15-679 25124552
Štítky
Hygiena a epidemiologie Infekční lékařství Laboratoř
Článek Clearance of Pneumococcal Colonization in Infants Is Delayed through Altered Macrophage TraffickingČlánek An Model of Latency and Reactivation of Varicella Zoster Virus in Human Stem Cell-Derived NeuronsČlánek Protective mAbs and Cross-Reactive mAbs Raised by Immunization with Engineered Marburg Virus GPsČlánek Specific Cell Targeting Therapy Bypasses Drug Resistance Mechanisms in African TrypanosomiasisČlánek Peptidoglycan Branched Stem Peptides Contribute to Virulence by Inhibiting Pneumolysin ReleaseČlánek HIV Latency Is Established Directly and Early in Both Resting and Activated Primary CD4 T CellsČlánek Sequence-Specific Fidelity Alterations Associated with West Nile Virus Attenuation in Mosquitoes
Článek vyšel v časopisePLOS Pathogens
Nejčtenější tento týden
2015 Číslo 6- Stillova choroba: vzácné a závažné systémové onemocnění
- Perorální antivirotika jako vysoce efektivní nástroj prevence hospitalizací kvůli COVID-19 − otázky a odpovědi pro praxi
- Diagnostika virových hepatitid v kostce – zorientujte se (nejen) v sérologii
- Jak souvisí postcovidový syndrom s poškozením mozku?
- Familiární středomořská horečka
-
Všechny články tohoto čísla
- Introducing “Research Matters”
- Exploring Host–Pathogen Interactions through Biological Control
- Analysis of Bottlenecks in Experimental Models of Infection
- Expected and Unexpected Features of the Newly Discovered Bat Influenza A-like Viruses
- Clearance of Pneumococcal Colonization in Infants Is Delayed through Altered Macrophage Trafficking
- Recombinant Murine Gamma Herpesvirus 68 Carrying KSHV G Protein-Coupled Receptor Induces Angiogenic Lesions in Mice
- TRIM30α Is a Negative-Feedback Regulator of the Intracellular DNA and DNA Virus-Triggered Response by Targeting STING
- Targeting Human Transmission Biology for Malaria Elimination
- Two Cdc2 Kinase Genes with Distinct Functions in Vegetative and Infectious Hyphae in
- An Model of Latency and Reactivation of Varicella Zoster Virus in Human Stem Cell-Derived Neurons
- Protective mAbs and Cross-Reactive mAbs Raised by Immunization with Engineered Marburg Virus GPs
- Virulence Factors of Induce Both the Unfolded Protein and Integrated Stress Responses in Airway Epithelial Cells
- Peptide-MHC-I from Endogenous Antigen Outnumber Those from Exogenous Antigen, Irrespective of APC Phenotype or Activation
- Specific Cell Targeting Therapy Bypasses Drug Resistance Mechanisms in African Trypanosomiasis
- An Ultrasensitive Mechanism Regulates Influenza Virus-Induced Inflammation
- The Role of Human Transportation Networks in Mediating the Genetic Structure of Seasonal Influenza in the United States
- Host Delivery of Favorite Meals for Intracellular Pathogens
- Complement-Opsonized HIV-1 Overcomes Restriction in Dendritic Cells
- Inter-Seasonal Influenza is Characterized by Extended Virus Transmission and Persistence
- A Critical Role for CLSP2 in the Modulation of Antifungal Immune Response in Mosquitoes
- Twilight, a Novel Circadian-Regulated Gene, Integrates Phototropism with Nutrient and Redox Homeostasis during Fungal Development
- Surface-Associated Lipoproteins Link Virulence to Colitogenic Activity in IL-10-Deficient Mice Independent of Their Expression Levels
- Latent Membrane Protein LMP2A Impairs Recognition of EBV-Infected Cells by CD8+ T Cells
- Bank Vole Prion Protein As an Apparently Universal Substrate for RT-QuIC-Based Detection and Discrimination of Prion Strains
- Neuronal Subtype and Satellite Cell Tropism Are Determinants of Varicella-Zoster Virus Virulence in Human Dorsal Root Ganglia Xenografts
- Molecular Basis for the Selective Inhibition of Respiratory Syncytial Virus RNA Polymerase by 2'-Fluoro-4'-Chloromethyl-Cytidine Triphosphate
- Structure of the Virulence Factor, SidC Reveals a Unique PI(4)P-Specific Binding Domain Essential for Its Targeting to the Bacterial Phagosome
- Activated Brain Endothelial Cells Cross-Present Malaria Antigen
- Fungal Morphology, Iron Homeostasis, and Lipid Metabolism Regulated by a GATA Transcription Factor in
- Peptidoglycan Branched Stem Peptides Contribute to Virulence by Inhibiting Pneumolysin Release
- A Macrophage Subversion Factor Is Shared by Intracellular and Extracellular Pathogens
- A Novel AT-Rich DNA Recognition Mechanism for Bacterial Xenogeneic Silencer MvaT
- Reovirus FAST Proteins Drive Pore Formation and Syncytiogenesis Using a Novel Helix-Loop-Helix Fusion-Inducing Lipid Packing Sensor
- The Role of ExoS in Dissemination of during Pneumonia
- IRF-5-Mediated Inflammation Limits CD8 T Cell Expansion by Inducing HIF-1α and Impairing Dendritic Cell Functions during Infection
- Discordant Impact of HLA on Viral Replicative Capacity and Disease Progression in Pediatric and Adult HIV Infection
- Crystal Structure of USP7 Ubiquitin-like Domains with an ICP0 Peptide Reveals a Novel Mechanism Used by Viral and Cellular Proteins to Target USP7
- HIV Latency Is Established Directly and Early in Both Resting and Activated Primary CD4 T Cells
- HPV16 Down-Regulates the Insulin-Like Growth Factor Binding Protein 2 to Promote Epithelial Invasion in Organotypic Cultures
- The νSaα Specific Lipoprotein Like Cluster () of . USA300 Contributes to Immune Stimulation and Invasion in Human Cells
- RSV-Induced H3K4 Demethylase KDM5B Leads to Regulation of Dendritic Cell-Derived Innate Cytokines and Exacerbates Pathogenesis
- Leukocidin A/B (LukAB) Kills Human Monocytes via Host NLRP3 and ASC when Extracellular, but Not Intracellular
- Border Patrol Gone Awry: Lung NKT Cell Activation by Exacerbates Tularemia-Like Disease
- The Curious Road from Basic Pathogen Research to Clinical Translation
- From Cell and Organismal Biology to Drugs
- Adenovirus Tales: From the Cell Surface to the Nuclear Pore Complex
- A 21st Century Perspective of Poliovirus Replication
- Is Development of a Vaccine against Feasible?
- Waterborne Viruses: A Barrier to Safe Drinking Water
- Battling Phages: How Bacteria Defend against Viral Attack
- Archaea in and on the Human Body: Health Implications and Future Directions
- Degradation of Human PDZ-Proteins by Human Alphapapillomaviruses Represents an Evolutionary Adaptation to a Novel Cellular Niche
- Natural Variants of the KPC-2 Carbapenemase have Evolved Increased Catalytic Efficiency for Ceftazidime Hydrolysis at the Cost of Enzyme Stability
- Potent Cell-Intrinsic Immune Responses in Dendritic Cells Facilitate HIV-1-Specific T Cell Immunity in HIV-1 Elite Controllers
- The Mammalian Cell Cycle Regulates Parvovirus Nuclear Capsid Assembly
- Host Reticulocytes Provide Metabolic Reservoirs That Can Be Exploited by Malaria Parasites
- The Proteome of the Isolated Containing Vacuole Reveals a Complex Trafficking Platform Enriched for Retromer Components
- NK-, NKT- and CD8-Derived IFNγ Drives Myeloid Cell Activation and Erythrophagocytosis, Resulting in Trypanosomosis-Associated Acute Anemia
- Successes and Challenges on the Road to Cure Hepatitis C
- BRCA1 Regulates IFI16 Mediated Nuclear Innate Sensing of Herpes Viral DNA and Subsequent Induction of the Innate Inflammasome and Interferon-β Responses
- A Structural and Functional Comparison Between Infectious and Non-Infectious Autocatalytic Recombinant PrP Conformers
- Phosphorylation of the Peptidoglycan Synthase PonA1 Governs the Rate of Polar Elongation in Mycobacteria
- Human Immunodeficiency Virus Type 1 Nef Inhibits Autophagy through Transcription Factor EB Sequestration
- Sequence-Specific Fidelity Alterations Associated with West Nile Virus Attenuation in Mosquitoes
- EBV BART MicroRNAs Target Multiple Pro-apoptotic Cellular Genes to Promote Epithelial Cell Survival
- Single-Cell and Single-Cycle Analysis of HIV-1 Replication
- TRIM32 Senses and Restricts Influenza A Virus by Ubiquitination of PB1 Polymerase
- The Herpes Simplex Virus Protein pUL31 Escorts Nucleocapsids to Sites of Nuclear Egress, a Process Coordinated by Its N-Terminal Domain
- Host Transcriptional Response to Influenza and Other Acute Respiratory Viral Infections – A Prospective Cohort Study
- PLOS Pathogens
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- HIV Latency Is Established Directly and Early in Both Resting and Activated Primary CD4 T Cells
- Battling Phages: How Bacteria Defend against Viral Attack
- A 21st Century Perspective of Poliovirus Replication
- Adenovirus Tales: From the Cell Surface to the Nuclear Pore Complex
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MUDr. Irena Krčmová, CSc.
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



