-
Články
Top novinky
Reklama- Vzdělávání
- Časopisy
Top články
- Témata
Top novinky
Reklama- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Volná místa
Doporučené pozice
Reklama- Praxe
Top novinky
ReklamaA Genome-Wide Association Meta-Analysis of Circulating Sex Hormone–Binding Globulin Reveals Multiple Loci Implicated in Sex Steroid Hormone Regulation
Sex hormone-binding globulin (SHBG) is a glycoprotein responsible for the transport and biologic availability of sex steroid hormones, primarily testosterone and estradiol. SHBG has been associated with chronic diseases including type 2 diabetes (T2D) and with hormone-sensitive cancers such as breast and prostate cancer. We performed a genome-wide association study (GWAS) meta-analysis of 21,791 individuals from 10 epidemiologic studies and validated these findings in 7,046 individuals in an additional six studies. We identified twelve genomic regions (SNPs) associated with circulating SHBG concentrations. Loci near the identified SNPs included SHBG (rs12150660, 17p13.1, p = 1.8×10−106), PRMT6 (rs17496332, 1p13.3, p = 1.4×10−11), GCKR (rs780093, 2p23.3, p = 2.2×10−16), ZBTB10 (rs440837, 8q21.13, p = 3.4×10−09), JMJD1C (rs7910927, 10q21.3, p = 6.1×10−35), SLCO1B1 (rs4149056, 12p12.1, p = 1.9×10−08), NR2F2 (rs8023580, 15q26.2, p = 8.3×10−12), ZNF652 (rs2411984, 17q21.32, p = 3.5×10−14), TDGF3 (rs1573036, Xq22.3, p = 4.1×10−14), LHCGR (rs10454142, 2p16.3, p = 1.3×10−07), BAIAP2L1 (rs3779195, 7q21.3, p = 2.7×10−08), and UGT2B15 (rs293428, 4q13.2, p = 5.5×10−06). These genes encompass multiple biologic pathways, including hepatic function, lipid metabolism, carbohydrate metabolism and T2D, androgen and estrogen receptor function, epigenetic effects, and the biology of sex steroid hormone-responsive cancers including breast and prostate cancer. We found evidence of sex-differentiated genetic influences on SHBG. In a sex-specific GWAS, the loci 4q13.2-UGT2B15 was significant in men only (men p = 2.5×10−08, women p = 0.66, heterogeneity p = 0.003). Additionally, three loci showed strong sex-differentiated effects: 17p13.1-SHBG and Xq22.3-TDGF3 were stronger in men, whereas 8q21.12-ZBTB10 was stronger in women. Conditional analyses identified additional signals at the SHBG gene that together almost double the proportion of variance explained at the locus. Using an independent study of 1,129 individuals, all SNPs identified in the overall or sex-differentiated or conditional analyses explained ∼15.6% and ∼8.4% of the genetic variation of SHBG concentrations in men and women, respectively. The evidence for sex-differentiated effects and allelic heterogeneity highlight the importance of considering these features when estimating complex trait variance.
Published in the journal: . PLoS Genet 8(7): e32767. doi:10.1371/journal.pgen.1002805
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1002805Summary
Sex hormone-binding globulin (SHBG) is a glycoprotein responsible for the transport and biologic availability of sex steroid hormones, primarily testosterone and estradiol. SHBG has been associated with chronic diseases including type 2 diabetes (T2D) and with hormone-sensitive cancers such as breast and prostate cancer. We performed a genome-wide association study (GWAS) meta-analysis of 21,791 individuals from 10 epidemiologic studies and validated these findings in 7,046 individuals in an additional six studies. We identified twelve genomic regions (SNPs) associated with circulating SHBG concentrations. Loci near the identified SNPs included SHBG (rs12150660, 17p13.1, p = 1.8×10−106), PRMT6 (rs17496332, 1p13.3, p = 1.4×10−11), GCKR (rs780093, 2p23.3, p = 2.2×10−16), ZBTB10 (rs440837, 8q21.13, p = 3.4×10−09), JMJD1C (rs7910927, 10q21.3, p = 6.1×10−35), SLCO1B1 (rs4149056, 12p12.1, p = 1.9×10−08), NR2F2 (rs8023580, 15q26.2, p = 8.3×10−12), ZNF652 (rs2411984, 17q21.32, p = 3.5×10−14), TDGF3 (rs1573036, Xq22.3, p = 4.1×10−14), LHCGR (rs10454142, 2p16.3, p = 1.3×10−07), BAIAP2L1 (rs3779195, 7q21.3, p = 2.7×10−08), and UGT2B15 (rs293428, 4q13.2, p = 5.5×10−06). These genes encompass multiple biologic pathways, including hepatic function, lipid metabolism, carbohydrate metabolism and T2D, androgen and estrogen receptor function, epigenetic effects, and the biology of sex steroid hormone-responsive cancers including breast and prostate cancer. We found evidence of sex-differentiated genetic influences on SHBG. In a sex-specific GWAS, the loci 4q13.2-UGT2B15 was significant in men only (men p = 2.5×10−08, women p = 0.66, heterogeneity p = 0.003). Additionally, three loci showed strong sex-differentiated effects: 17p13.1-SHBG and Xq22.3-TDGF3 were stronger in men, whereas 8q21.12-ZBTB10 was stronger in women. Conditional analyses identified additional signals at the SHBG gene that together almost double the proportion of variance explained at the locus. Using an independent study of 1,129 individuals, all SNPs identified in the overall or sex-differentiated or conditional analyses explained ∼15.6% and ∼8.4% of the genetic variation of SHBG concentrations in men and women, respectively. The evidence for sex-differentiated effects and allelic heterogeneity highlight the importance of considering these features when estimating complex trait variance.
Introduction
Sex hormone-binding globulin (SHBG) is a protein secreted mainly by the liver that binds to the sex steroids, testosterone, dihydrotestosterone, and estradiol, transports them in the circulation, and influences their action in target tissues by regulating their bioavailability. SHBG thereby influences the expression of sex hormone sensitive phenotypes including sexual characteristics and reproductive function in men and women [1]. In addition to regulating sex steroid hormone effects, SHBG may exert independent effects through its own receptor [2]. Variation in SHBG concentration has also been associated with various chronic diseases including cancers [3], polycystic ovary syndrome (PCOS) [4], [5] and type 2 diabetes (T2D) [6], [7]. Although SHBG is estimated to have a heritable component (∼50%) [8], little is known about the genetic regulation of SHBG. Polymorphisms at the SHBG gene locus have been associated with SHBG concentrations [9], [10], but much remains unknown about specific genetic variants that may determine circulating SHBG concentrations. Identifying genetic factors that influence SHBG may provide insights into the biology of sex steroid hormone regulation, metabolism and tissue effects that underlie their relationship with chronic diseases such as T2D as well as hormone-sensitive cancers such as breast and prostate cancer.
Results
We identified nine loci associated with SHBG concentrations at the genome-wide significance threshold of p = 5×10−8 (Table 1 and Figure 1) in a genome-wide association study (GWAS) meta-analysis of circulating SHBG concentrations in 21,791 men and women from 10 studies (Table S1). All nine lead SNPs at these loci had effects in the same direction (seven with p<0.05) in the validation dataset of 7,046 men and women from six additional studies (Table S2). The strongest association was within the SHBG locus (rs12150660, p = 2×10−106). Together, these nine lead SNPs explained 7.2% of the genetic variance (assuming 50% heritability) in SHBG concentrations.
Fig. 1. Manhattan plot of the autosomal SNPs identified in the GWA meta-analysis. 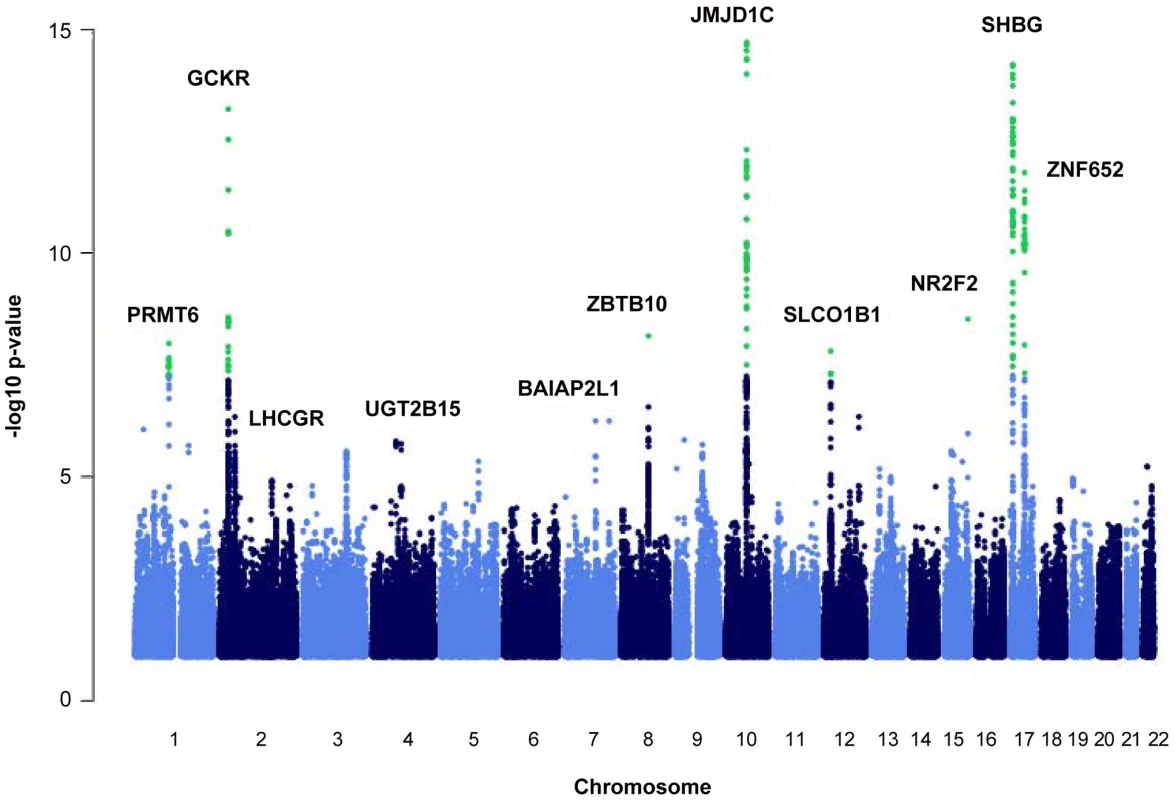
The Manhattan plot depicts the SNPs identified in the GWAS analysis labeled with the nearest gene on the plot. The SNP identified on the X chromosome, rs1573036, at Xq22.3, is not included in this figure. Tab. 1. SNPs representing loci associated with circulating SHBG concentrations. 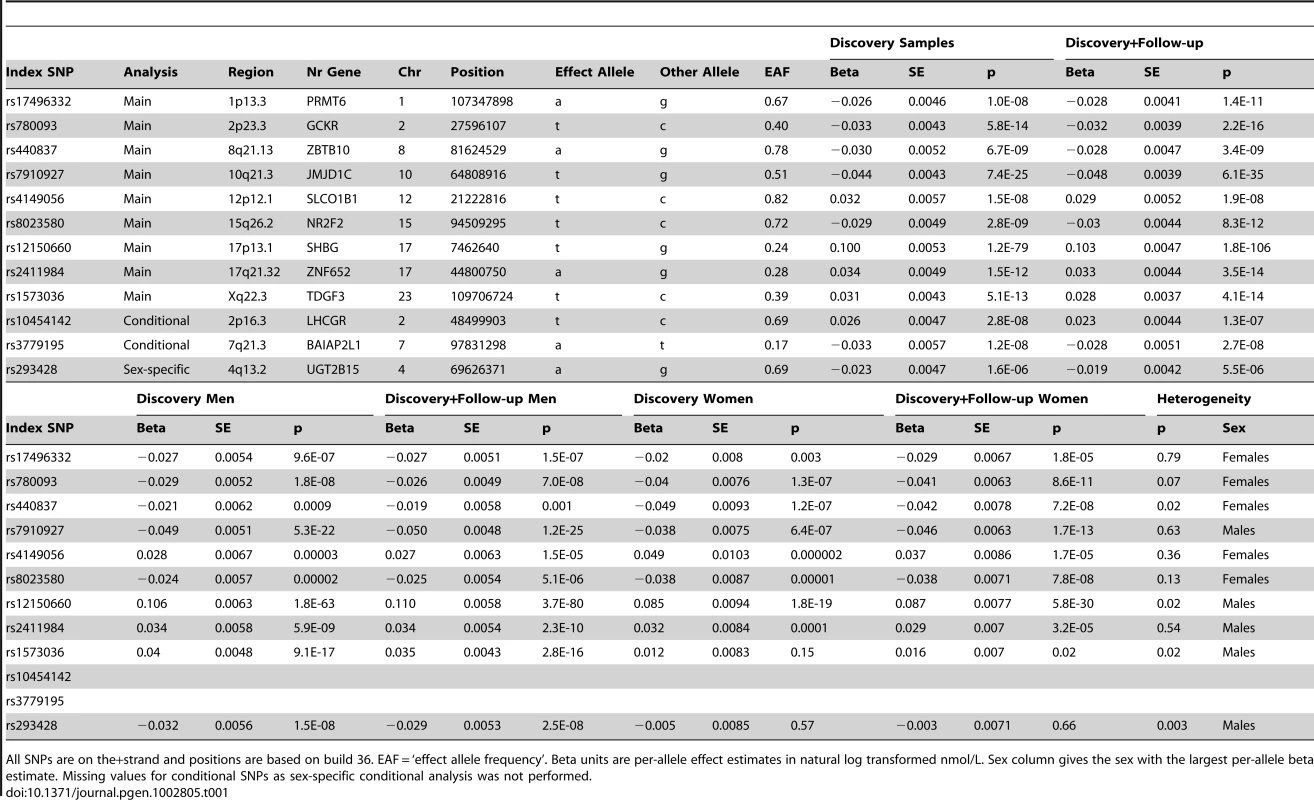
All SNPs are on the+strand and positions are based on build 36. EAF = ‘effect allele frequency’. Beta units are per-allele effect estimates in natural log transformed nmol/L. Sex column gives the sex with the largest per-allele beta estimate. Missing values for conditional SNPs as sex-specific conditional analysis was not performed. We next performed a series of additional analyses to explain more of the phenotypic variance (Figure 2). First, we hypothesized that genetic effects may be different in men and women, as SHBG concentrations are >50% higher in females than males, and may be differentially regulated between sexes. In a sex stratified analysis, three of the nine loci showed evidence of sex-differentiated effects at p<0.02 when we would not expect any signals to have reached this level of significance by chance. The associations at the 17p13.1-SHBG and Xq22.3 loci were stronger in males whereas the association at the 8q21.13 locus was stronger in females. To investigate the apparent differential sex effect for the X chromosome further we ran a recessive regression model for the X chromosome SNP rs1573036 in women in the Framingham Heart Study and found no association with SHBG suggesting the sex-differentiated effect is not the result of a recessive inheritance pattern. Sex stratified GWAS identified one novel signal in men, which showed no association in women (4q13.2: men p = 2.5×10−8, women p = 0.66, heterogeneity p = 0.003).
Fig. 2. Summary of the analytic plan. 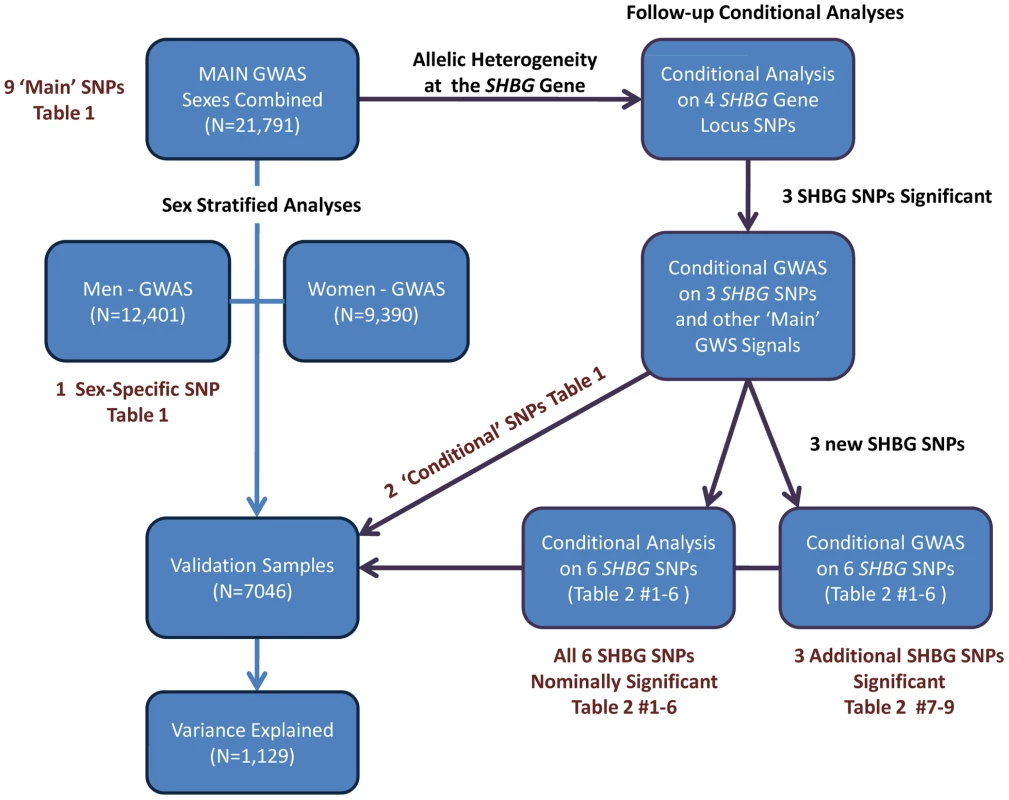
A series of conditional analyses were performed to identify statistically independent signals. At the SHBG locus, three apparently independent additional signals separate from the main index SNP were observed, based on low (r2<0.05) pairwise correlations in HapMap (rs6258 p = 2.7×10−46, rs1625895 p = 1.2×10−14 and rs3853894 p = 2.5×10−11). A series of iterative conditional analyses (Table 2) involving SNPs at the SHBG locus generated a final regression model including six statistically independent SHBG SNPs. Four of these SNPs (#1–4 Table 2) retained GWS when conditioned against the other five, and two were nominally associated (SNP#5 p = 0.0001, SNP#6 p = 0.01). Re-running the GWAS meta-analysis adjusting for these six SNPs revealed evidence for three additional statistically independent (pairwise HapMap r2<0.01) signals at the SHBG locus (SNP#7 p = 1.5×10−7, SNP#8 p = 4.6×10−5, SNP#9 p = 9.9×10−6) (Figure 3). There were also two additional trans signals located at 2p16.3 and 7q21.3 (Table 1). Although the 2p16.3 signal dropped below GWS when combined with follow-up samples (p = 1×10−7), the index SNP at 2p16.3 is ∼300 kb away from a strong candidate gene, the luteinizing hormone receptor gene (LHCGR).
Fig. 3. Allelic heterogeneity at the SHBG gene locus. 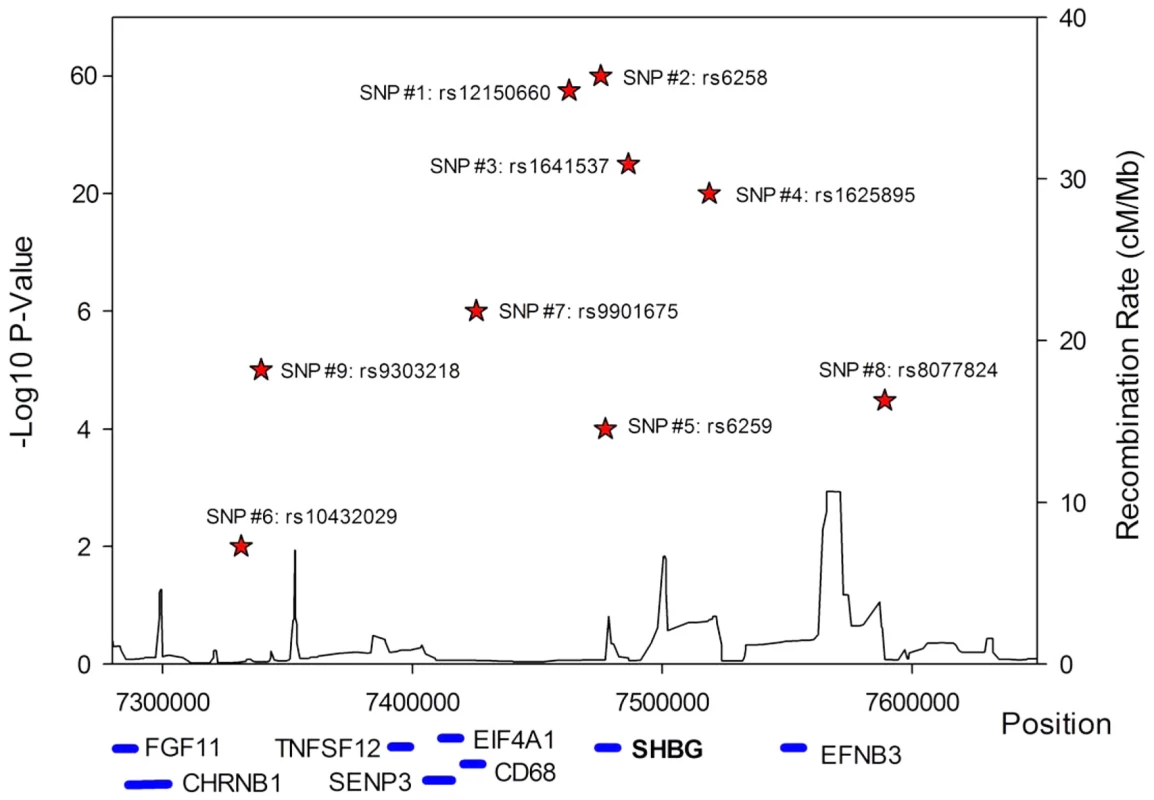
There was significant allelic heterogeneity at the SHBG gene locus. The nine independent signals identified in the SHBG gene are shown in relation to their position within the gene. All positions based on build 36. Not all genes are shown. Tab. 2. Statistically independent signals at the SHBG gene locus. 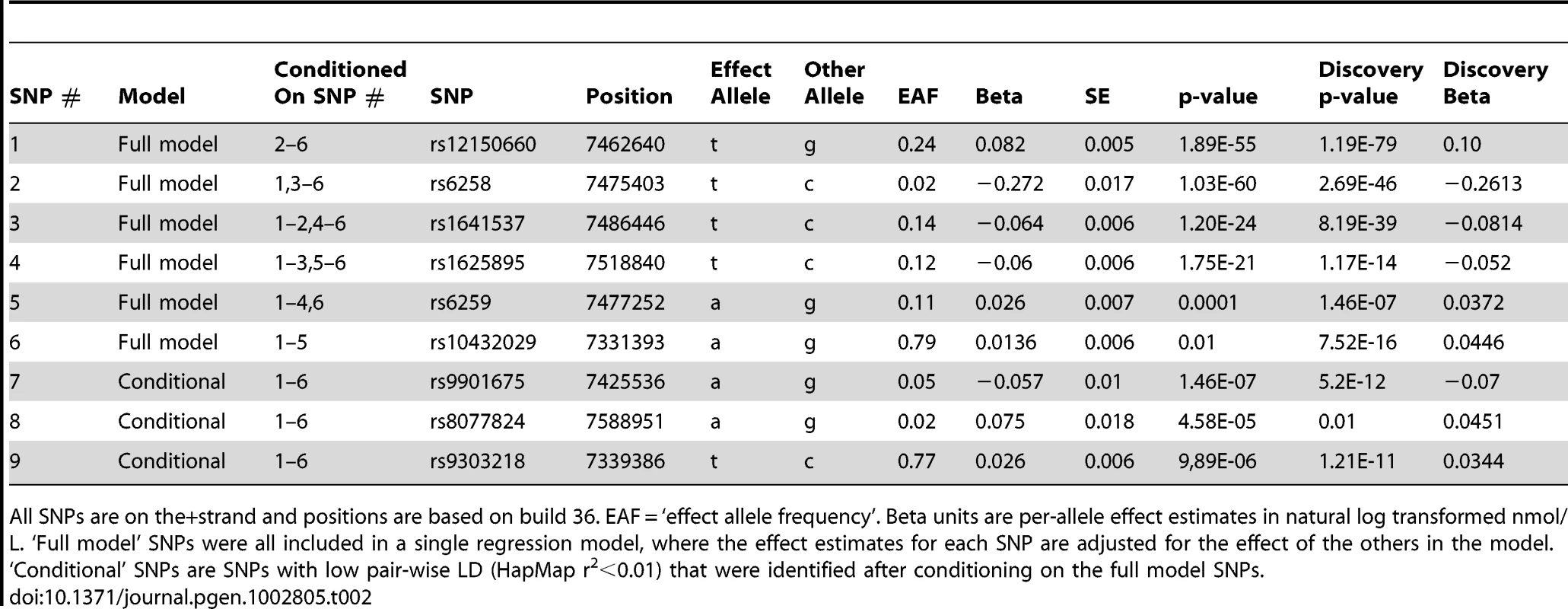
All SNPs are on the+strand and positions are based on build 36. EAF = ‘effect allele frequency’. Beta units are per-allele effect estimates in natural log transformed nmol/L. ‘Full model’ SNPs were all included in a single regression model, where the effect estimates for each SNP are adjusted for the effect of the others in the model. The majority of pair-wise correlations for the nine SHBG locus SNPs highlighted by our conditional analyses showed very low HapMap r2 values. However, the pairwise D′ values are often high (Table S3) indicating that no or few recombination events have occurred between some SNPs, and that combinations of SNPs may be tagging un-typed variants on a common haplotype. To investigate this possibility, we performed more extensive analyses in a single study (NFBC1966, n = 4467). We used a denser set of SNPs imputed from the June 2011 version of the 1000 Genomes data and performed model selection analyses. Model selection identifies a set of SNPs that best explain phenotypic variation, while simultaneously penalizing each SNP included in this set, and therefore correlated SNPs tend to be excluded from the final model. These analyses consistently included at least seven SNPs in the model, although it is hard to estimate the false-negative rate of using the reduced sample size. While we are underpowered to accurately pinpoint the exact number of independent signals, these analyses support the results of the conditional analysis and suggest that multiple variants at the SHBG locus are independently associated with SHBG concentrations.
Data from an independent study, the InCHIANTI study, was used to calculate the proportion of genetic variance in SHBG concentrations explained when accounting for sex specific effects, the multiple signals of association at the SHBG locus, and the additional trans signals identified post conditional analysis. In men and women we explained ∼15.6% and ∼8.4% of the heritable component respectively. The SHBG locus accounted for ∼10% and ∼6.6% of the genetic variation in men and women respectively with the lead SNP in isolation accounting for ∼7.8% and ∼3.3% of the variation in men and women, respectively.
We identified genes near the associated SNPs and explored their biologic relevance to SHBG. The genes associated with identified SNPs included the SHBG locus (rs12150660, 17p13.1, p = 1.8×10−106), PRMT6 (rs17496332, 1p13.3, p = 1.4×10−11), GCKR (rs780093, 2p23.3, p = 2.2×10−16), ZBTB10 (rs440837, 8q21.13, p = 3.4×10−09), JMJD1C (rs7910927, 10q21.3, p = 6.1×10−35), SLCO1B1 (rs4149056, 12p12.1, p = 1.9×10−08), NR2F2 (rs8023580, 15q26.2, p = 8.3×10−12), ZNF652 (rs2411984, 17q21.32, p = 3.5×10−14), TDGF3 (rs1573036, Xq22.3, p = 4.1×10−14), LHCGR (rs10454142, 2p16.3, p = 1.3×10−07), BAIAP2L1 (rs3779195, 7q21.3, p = 2.7×10−08), and UGT2B15 (rs293428, 4q13.2, p = 5.5×10−06) (Figure 1).
We used the online tool STRING (www.string-db.org) to perform pathway analyses to explore possible interactions between the SHBG gene and the proteins encoded by the 11 most plausible genes nearest the 11 SNPs listed above. There was an interaction noted between GCKR and JMJD1C which were associated with the lipoprotein fractions VLDL and HDL, respectively [11]. In an expanded analysis, we assessed protein interactions among SHBG and 67 genes within 500 kb of our 11 identified SNPs and uncovered additional protein interaction pathways. An interaction between two proteins encoded by GTF2A1L and STON1 was found; these proteins are co-expressed in testicular germ cells in the mouse [12]. An interaction between LHCGR and BRI3 encoded proteins that are associated with the G-protein coupled receptor complex in the human luteinizing hormone receptor was also identified [13]. Finally, an interaction between LHCGR and IAPP (amylin) proteins which are components of a ligand/G-protein receptor/G-protein alpha subunit complex was found (database: www.reactome.com).
Targeted analysis of two strong candidate genes, hepatocyte nuclear factor-4α (HNF4α) and peroxisome-proliferating receptor γ (PPARγ) did not identify any SNPs at HNF4α but did identify one SNP, rs2920502, at PPARγ that reached statistical significance (p = 9.9×10−5) and a second SNP at PPARγ, rs13081389, that reached nominal significance (p = 0.01).
Discussion
In total, we identified 12 genomic regions associated with circulating SHBG concentrations, including extensive allelic heterogeneity at the SHBG locus itself. Conditional meta-analyses carried out at the SHBG locus, identified nine genome-wide significant SNPs with low correlation (r2<0.01) between them. Two of these signals (rs6258 [10] and rs6259) are missense variants and two are low frequency variants (MAF ∼2%). Furthermore, rs12150660 is highly correlated (r2>0.95) [10] with a pentanucleotide repeat, which affects SHBG expression in-vitro [14]. To our knowledge, the magnitude of secondary signals observed at this locus are the largest seen for any complex trait.
The proportion of genetic variance in SHBG serum concentrations explained when accounting for sex specific effects, the multiple signals of association at the SHBG locus, and the additional trans signals identified post conditional analysis was ∼15.6% in men and ∼8.4% in women. The SHBG locus accounted for ∼10% and ∼6.6% of the genetic variance in men and women, respectively, with the lead SNP explaining most of the genetic variation at ∼7.8% for men and ∼3.3% for women. Thus additional signals at the SHBG locus identified through conditional analyses approximately doubled the variance of the trait explained. While we provide evidence for multiple variants associated with SHBG concentrations, further studies are needed to pinpoint the causal loci and functional variants. For the 11 regions outside the SHBG locus, most have biologically plausible related genes within 300 kb.
Biology of Plausible Genes near Identified SNPs
Several genes near the identified SNPs regulate sex steroid production and function. The NR2F2 locus (15q26.2) encodes a nuclear receptor important in testicular Leydig cell function, the primary source of gonadal testosterone production [15], and has been linked to male infertility [16]. NR2F2 has also been associated with estrogen receptor alpha (ERα) signaling and may influence hormone responsivity in breast cancer [17]. PRMT6 (1p13.3) also encodes a nuclear receptor regulatory protein that mediates estrogen signaling as a co-activator of the estrogen receptor [18]. LHCGR (2p16.3) encodes the luteinizing hormone receptor which was associated with polycystic ovary syndrome (PCOS) in a recent GWAS [19], [20]. PCOS is both a reproductive and metabolic disorder characterized by higher testosterone serum concentrations as well as an increased prevalence of obesity, insulin resistance, and T2D in women. Inappropriate secretion of luteinizing hormone leads to increased ovarian production of testosterone. Coincident lower SHBG concentrations contribute to increased bioavailable testosterone concentrations and the expression of both reproductive and metabolic phenotypes in PCOS [21], [22], [23].
The SLCO1B1 locus encodes a liver-specific transporter of thyroid hormone as well as estrogens which impact liver production of SHBG [24]. JMJD1C (10q21.3), also known as TRIP 8 (thyroid hormone receptor interactor protein 8 [25]), may impact SHBG concentrations via thyroid hormone effects on liver protein production. Thyroid hormone may alter SHBG production through effects on HNF4α which is known to regulate SHBG transcription [26], [27].
Many of the genes identified are involved in carbohydrate and lipid metabolism and liver function. The GCKR locus (2p23.3) encodes a protein that regulates glucokinase activity and has been associated with T2D in several ethnic populations [28], [29], [30], [31]. GCKR has been associated with metabolic and inflammatory traits including triglyceride concentrations and other lipid fractions [30], [32], fasting plasma glucose [33], [34], insulin concentrations, uric acid, c-reactive protein (CRP), and non-alcoholic fatty liver disease which are all characteristic of the metabolic syndrome and T2D [28], [35], [36], [37], [38], [39], [40], [41], [42]. The SLCO1B1 locus (12p12.1) codes for a protein, hepatocyte protein anion-transporting polypeptide 1B1, involved in liver metabolism of both endogenous and exogenous compounds [43]. Consistent with SLCO1B1's role in liver metabolism, the same SNP (rs4149056) has been associated with circulating bilirubin concentrations in previous GWAS [44]. BAIAP2L1 (7q21.3) encodes a protein important in cytoskeleton organization [45] that has been associated with the inflammatory marker CRP in patients with arthritis [46]. BAIAP2L1 is also known as IRTKS (insulin receptor tyrosine kinase substrate) which is involved in insulin receptor signaling [47] and may relate to insulin resistant states including obesity and T2D [48], [49], [50], [51], [52], [53], [54]. We conducted a targeted analysis of PPARγ, a gene that influences SHBG gene expression in the liver [1], [55] and is associated with T2D [56], [57]. Our analysis identified one significant SNP (rs2920502, p = 9.9×10−5) and a second nominally significant SNP (rs13081389, p = 0.01) at PPARγ. Some of the identified genes involved in hepatic metabolism of lipids and carbohydrates may be affect SHBG concentrations indirectly through effects on the SHBG transcription regulator HNF4α although HNF4α itself was not identified in this meta-analyses [27], [58], [59], [60].
The UGT2B15 locus (4q13.2) was significantly associated with SHBG concentrations in men but not women in this meta-analysis. UGT2B15 belongs to a family of genes (the UGT2B gene family) that code for enzymes involved in the metabolism of sex hormones through glucuronidation which allows for excretion of sex steroids through the kidney and the gut via bile excretion [61], [62], primary clearance mechanisms for sex steroids [63]. UGT2B15 is involved in the conjugation and inactivation of testosterone [64]. An association between rs293428 in the UGT2B15 locus and circulating SHBG concentrations in men is supported by a previous study demonstrating that a non-synonymous SNP in UGT2B15 (rs1902023; D85Y) is associated with serum SHBG concentrations in younger adult men [65]. UGT2B15 is thought to play a significant role in local tissue inactivation of androgens in androgen dependent prostate cancer [66], [67]. The mechanism behind the influence of genetic variants in UGT2B15 on SHBG concentrations is unknown, but one may speculate that UGT2B15 affects the local androgenic environment in selected tissues, which in turn results in regulation of SHBG concentrations.
In addition to UGT2B15, three other genes near the identified SNPs are associated with carcinogenesis, particularly in the prostate and breast. ZBTB10 (8q21.13), has been linked to breast cancer [68]. In breast cancer cell lines ZBTB10 is suppressed by ROS-microRNA27a thereby enhancing ERα alpha expression and mediating estrogen effects [17]. The ZNF652 (17q21.32) locus codes for a DNA binding protein thought to act as a tumor suppressor gene in breast cancer [69], [70], [71] that is also co-expressed with the androgen receptor in prostate cancer [72]. TDGF3, teratocarcinoma derived growth factor 3, is the only significant region identified on the X chromosome ((Xq22.3). TDGF3 is a pseudogene of TDGF1 located on chromosome 3p23-p21 that has been associated with testicular germ cell tumors [73].
Strengths and Limitations
This GWAS meta-analysis incorporated data from approximately 22,000 men and women from 16 epidemiologic cohorts. The overall size of the study yields power but the meta-analysis of data from different epidemiologic studies requires the inclusion of different laboratory methods. The different studies used a variety of assay methodologies to measure serum SHBG concentrations although the vast majority were immunoassays (Tables S1 and S2, Text S1) with similar methodologies. Variation introduced by the use of different SHBG assays would result in loss of statistical power and likely bias toward the null. Additionally, the majority of women were post-menopausal as ascertained by self-report in all studies (Table S1). SHBG concentrations, like testosterone, decline only slightly across the menopause [74] so adjustment for menopause status is not necessary. SHBG may also increase with ovulation and be slightly higher in the luteal versus the follicular phase of the menstrual cycle in premenopausal women, but most studies did not collect data on menstrual phase at the time of SHBG measurement so adjustment for menstrual phase was not possible [75]. Finally, individuals were not excluded based on health status, therefore some individuals with chronic conditions that may affect hepatic production of or clearance of proteins including SHBG such as liver disease, renal disease, or severe malnutrition, may have been included in this analysis.
Conclusion
SHBG synthesis in the liver is known to be affected directly or indirectly by estrogens, androgens and thyroid hormones and has been observed to be inversely associated with the higher insulin concentrations characteristic of insulin resistant states such as T2D [1], [6]. In summary, the results of this GWAS reflect these influences. Three regions map to proteins related to hepatic function (12p12.1-SLCO1B1 [76], 2p23.3-GCKR [77] and 10q21.3-JMJD1C [77]). In addition, 2p23.3-GCKR and 7q21.3-BAIAP2L1 [alias insulin receptor tyrosine kinase substrate (IRTKS)] are involved in susceptibility to T2D [48] and insulin signaling [47], respectively. Two signals also mapped to loci involved in thyroid hormone regulation (10q21.3-JMJD1C and 12p12.1-SLCO1B1). One signal mapped to the receptor for luteinizing hormone 2p16.3-LHCGR [20], the hormone that stimulates testosterone production. Five regions mapped to genes previously implicated in androgen and estrogen signaling (1p13.3-PRMT6 [18], 8q21.13-ZBTB10 [17], 12p12.1-SLCO1B1 [76], 15q26.2-NR2F2 [78], 4q13.2-UGT2B15 [63]).
We have combined a conventional GWAS approach with detailed additional analyses, including sex stratification, conditional analysis and imputation from 1000 Genomes. Our results demonstrate that these approaches can lead to an appreciable gain in heritable variance explained. It does however highlight the complexity of elucidating individual variant causality through statistical approaches. In addition to the extensive allelic heterogeneity at the SHBG locus, our data identify loci with a role in sex steroid hormone metabolism, which may help elucidate the role of sex steroid hormones in disease, particularly T2D and hormone-sensitive cancers.
Methods
We performed a genome wide association study (GWAS) meta-analysis of 21,791 individuals (Table S1: 9,390 women, 12,401 men) from ten observational studies. Data from an additional six studies totaling 7,046 individuals (Table S2: 4,509 women; 2,537 men) were used for validation. The proportion of variance explained was estimated in an independent study (InCHIANTI, n = 1,129). The individual study protocols were approved by their respective institution's ethics committee/institutional review board and all participants provided informed consent prior to participation. Individuals known to be taking hormonal contraceptives or hormone replacement therapy at time of SHBG measurement were excluded from analysis. Age, sex and body mass index (BMI) were included as covariates. After applying standard quality control measures, imputed genotypes were available for approximately 2.5 M SNPs. See Figure 2 for an overview of the analytic plan and the Text S1 for further information for individual studies included in this meta-analysis.
GWAS Conditional Meta-Analysis Steps
Conditional analysis #1
The initial starting point for the conditional analysis was the four SHBG locus SNPs that all showed low Hapmap LD (r2<0.05) with each other: rs12150660 (lead SNP Table 1), rs6258 p = 2.7×10−46, rs1625895 p = 1.2×10−14 and rs3853894 p = 2.5×10−11. Each cohort fitted a single regression model, fitting SHBG concentrations against these four genome-wide significant SHBG locus SNPs (rs12150660, rs6258, rs1625895 and rs3853894), in addition to age, sex and BMI. After meta-analyzing the results from all cohorts, three of the SNPs retained genome wide significance when regressed against each other, with the fourth SNP narrowly missing that threshold (rs3853894, p = 4.1×10−6).
Conditional GWAS #1 (Table 1, conditional analysis)
We next performed a conditional GWAS meta-analysis, where each study included, as additional covariates to the original analysis plan, the ten genome-wide significant autosomal SNPs (the eight ‘Main’ signals from Table 1 and the two unique SHBG locus signals described above in addition to the lead SNP rs12150660: rs6258 and rs1625895). Three additional signals (independence based on HapMap r2<0.05) at the SHBG locus reached genome-wide significance (rs1641537 p = 7.8×10−32, rs6259 p = 1.5×10−12 and rs10432029 p = 3×10−8), giving a total of six independent signals in this gene region. In addition, two novel signals reached genome-wide significance in the conditional analysis, at 7q21.3 (rs3779195 p = 1×10−8) and 2p16.3 (rs10454142 p = 3×10−8). After replication, only rs3779195 at the BAIAP2L1 locus retained genome-wide significance.
Conditional analysis #2 (Table 2, full model)
Given the six signals observed at the SHBG locus (three through conditional analysis #1 rs12150660, rs6258, rs1625895, three through LD estimates from conditional GWAS #1: rs1641537, rs6259, rs10432029), we sought to confirm which of these six were truly independent by a second round of conditional analysis. All discovery and replication cohorts fitted a single regression model of the six SNPs (SNPs # 1–6, Table 2) against SHBG concentrations, using the same parameters and covariates as conditional analysis #1. Four of the six SNPs (#1–4: rs12150660, rs6258, rs1641537, and rs1625895) retained genome-wide significance when conditioned against each other, with two showing nominal evidence of association (SNP #5 rs6259, p = 0.0001; SNP #6 rs10432029, p = 0.01).
Conditional GWAS #2 (Table 2, conditional model)
Finally, we performed a second conditional GWAS analysis, adjusting for the six SHBG locus SNPs which had evidence of association from conditional analysis #2. All the discovery cohorts were used in this analysis, in addition to three replication cohorts (total sample size 24,354). This analysis revealed evidence for a further three independent signals at the SHBG locus (based on HapMap r2<0.01), SNP #7 rs9901675 p = 1.5×10−7, SNP #8 rs8077824 p = 4.6×10−5, and SNP #9 rs9393218 p = 9.9×10−6.
Sensitivity Analysis—Allelic Heterogeneity at the SHBG Locus
We performed a sensitivity analysis using samples from the 1966 Northern Finland Birth Cohort (NFBC1966) study to further investigate allelic heterogeneity at the SHBG locus (Text S1). The conditional meta-analysis showed evidence for up to nine signals at the SHBG locus, but it is possible that these signals could be explaining a much smaller number of causal variants in the region. Since 1000 Genomes imputation allows us to assess the genetic variation associated with a phenotype across a much denser set of markers, it increases our power to detect allelic heterogeneity within a region. Therefore, 1000 Genomes imputation was carried out on all the samples in the NFBC1966 study and forward selection was used to identify the set of SNPs that best explain the variation in the SHBG phenotype. 1000 Genomes imputation was carried out using IMPUTE2. The mean genotype probabilities for each SNP were calculated and used in the model selection step. Only SNPs 250 kb upstream and 250 kb downstream from the SHBG locus (7283453–7786700 bp) were used in the analysis. All SNPs with MAF <0.1% or an imputation quality score less than 0.4 were excluded from the analysis. In total, 1978 SHBG region SNPs measured or imputed in 4467 samples from the NFBC1966 study were used in the sensitivity analysis. Forward selection was implemented in R (version 2.13.0) using the stepAIC package to estimate the Akaikie Information Criterion (AIC), an inclusion parameter. Given the high degree of correlation between the SNPs in this region, we increased the penalty (k) on the number of terms included in the model to 12 (where it is usually two), to minimize possible over fitting. The final model included seven SNPs, adjusted for sex and BMI.
Pathway Analysis
We examined potential interactions among the proteins encoded by the SHBG locus and the proteins encoded by the 11 genes (ZBT10, TDGF1, ZNF652, PRMT6, JMJD1C, GCKR, BAIAP2L1, LHCGR, SLCO1B1, UGT2B15, NR2F2) closest to the 11 identified SNPs using pathway analysis with Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) Pathways Analysis (www.string-db.org). The interactions explored by STRING include direct (physical) and indirect (functional) associations. We then expanded the analysis to examine protein interactions among the SHBG gene and the proteins encoded by 67 genes within 500 kb of the 11 identified SNPs.
Targeted Candidate Gene Analysis
We conducted targeted analysis of two strong candidate genes, hepatocyte nuclear factor-4α (HNF4α) and peroxisome-proliferating receptor γ (PPARγ). Statistical significance thresholds were set correcting for the number of SNPs tested in each gene region (±100 kb).
Supporting Information
Zdroje
1. HammondGL 2011 Diverse roles for sex hormone-binding globulin in reproduction. Biol Reprod 85 431 441
2. HammesAAndreassenTKSpoelgenRRailaJHubnerN 2005 Role of endocytosis in cellular uptake of sex steroids. Cell 122 751 762
3. ThompsonDJHealeyCSBaynesCKalmyrzaevBAhmedS 2008 Identification of common variants in the SHBG gene affecting sex hormone-binding globulin levels and breast cancer risk in postmenopausal women. Cancer Epidemiol Biomarkers Prev 17 3490 3498
4. WassellJMichailMSolimanNWardlePG 2011 The value of sex hormone binding globulin (SHBG) in predicting treatment response in polycystic ovary syndrome (PCOS). Clin Lab 57 95 98
5. XitaNTsatsoulisAChatzikyriakidouAGeorgiouI 2003 Association of the (TAAAA)n repeat polymorphism in the sex hormone-binding globulin (SHBG) gene with polycystic ovary syndrome and relation to SHBG serum levels. J Clin Endocrinol Metab 88 5976 5980
6. DingELSongYMansonJEHunterDJLeeCC 2009 Sex hormone-binding globulin and risk of type 2 diabetes in women and men. N Engl J Med 361 1152 1163
7. PerryJRWeedonMNLangenbergCJacksonAULyssenkoV 2010 Genetic evidence that raised sex hormone binding globulin (SHBG) levels reduce the risk of type 2 diabetes. Hum Mol Genet 19 535 544
8. CovielloADZhuangWVLunettaKLBhasinSUlloorJ 2011 Circulating testosterone and SHBG concentrations are heritable in women: the Framingham Heart Study. J Clin Endocrinol Metab 96 E1491 1495
9. MelzerDPerryJRHernandezDCorsiAMStevensK 2008 A genome-wide association study identifies protein quantitative trait loci (pQTLs). PLoS Genet 4 e1000072 doi:10.1371/journal.pgen.1000072
10. OhlssonCWallaschofskiHLunettaKLStolkLPerryJR 2011 Genetic determinants of serum testosterone concentrations in men. PLoS Genet 7 e1002313 doi:10.1371/journal.pgen.1002313
11. ChasmanDIPareGMoraSHopewellJCPelosoG 2009 Forty-three loci associated with plasma lipoprotein size, concentration, and cholesterol content in genome-wide analysis. PLoS Genet 5 e1000730 doi:10.1371/journal.pgen.1000730
12. HanSYZhouLUpadhyayaALeeSHParkerKL 2001 TFIIAalpha/beta-like factor is encoded by a germ cell-specific gene whose expression is up-regulated with other general transcription factors during spermatogenesis in the mouse. Biol Reprod 64 507 517
13. KudoMOsugaYKobilkaBKHsuehAJ 1996 Transmembrane regions V and VI of the human luteinizing hormone receptor are required for constitutive activation by a mutation in the third intracellular loop. J Biol Chem 271 22470 22478
14. HogeveenKNTalikkaMHammondGL 2001 Human sex hormone-binding globulin promoter activity is influenced by a (TAAAA)n repeat element within an Alu sequence. J Biol Chem 276 36383 36390
15. MartinLJTremblayJJ 2010 Nuclear receptors in Leydig cell gene expression and function. Biol Reprod 83 3 14
16. HuZXiaYGuoXDaiJLiH 2011 A genome-wide association study in Chinese men identifies three risk loci for non-obstructive azoospermia. Nat Genet 44 183 186
17. LiXMertens-TalcottSUZhangSKimKBallJ 2010 MicroRNA-27a Indirectly Regulates Estrogen Receptor {alpha} Expression and Hormone Responsiveness in MCF-7 Breast Cancer Cells. Endocrinology 151 2462 2473
18. HarrisonMJTangYHDowhanDH 2010 Protein arginine methyltransferase 6 regulates multiple aspects of gene expression. Nucleic Acids Res 38 2201 2216
19. FranksSGharaniNMcCarthyM 2001 Candidate genes in polycystic ovary syndrome. Hum Reprod Update 7 405 410
20. ChenZJZhaoHHeLShiYQinY 2011 Genome-wide association study identifies susceptibility loci for polycystic ovary syndrome on chromosome 2p16.3, 2p21 and 9q33.3. Nat Genet 43 55 59
21. CousinPCalemard-MichelLLejeuneHRaverotGYessaadN 2004 Influence of SHBG gene pentanucleotide TAAAA repeat and D327N polymorphism on serum sex hormone-binding globulin concentration in hirsute women. J Clin Endocrinol Metab 89 917 924
22. EhrmannDA 2005 Polycystic ovary syndrome. N Engl J Med 352 1223 1236
23. FranksS 1995 Polycystic ovary syndrome. N Engl J Med 333 853 861
24. AbeTKakyoMTokuiTNakagomiRNishioT 1999 Identification of a novel gene family encoding human liver-specific organic anion transporter LST-1. J Biol Chem 274 17159 17163
25. LeeJWChoiHSGyurisJBrentRMooreDD 1995 Two classes of proteins dependent on either the presence or absence of thyroid hormone for interaction with the thyroid hormone receptor. Mol Endocrinol 9 243 254
26. SelvaDMHammondGL 2009 Thyroid hormones act indirectly to increase sex hormone-binding globulin production by liver via hepatocyte nuclear factor-4alpha. J Mol Endocrinol 43 19 27
27. JanneMHammondGL 1998 Hepatocyte nuclear factor-4 controls transcription from a TATA-less human sex hormone-binding globulin gene promoter. J Biol Chem 273 34105 34114
28. ReilingEvan 't RietEGroenewoudMJWelschenLMvan HoveEC 2009 Combined effects of single-nucleotide polymorphisms in GCK, GCKR, G6PC2 and MTNR1B on fasting plasma glucose and type 2 diabetes risk. Diabetologia 52 1866 1870
29. QiQWuYLiHLoosRJHuFB 2009 Association of GCKR rs780094, alone or in combination with GCK rs1799884, with type 2 diabetes and related traits in a Han Chinese population. Diabetologia 52 834 843
30. LingYLiXGuQChenHLuD 2011 Associations of common polymorphisms in GCKR with type 2 diabetes and related traits in a Han Chinese population: a case-control study. BMC Med Genet 12 66
31. TanakaDNagashimaKSasakiMYamadaCFunakoshiS 2011 GCKR mutations in Japanese families with clustered type 2 diabetes. Mol Genet Metab 102 453 460
32. VarboABennMTybjaerg-HansenAGrandePNordestgaardBG 2011 TRIB1 and GCKR polymorphisms, lipid levels, and risk of ischemic heart disease in the general population. Arterioscler Thromb Vasc Biol 31 451 457
33. TakeuchiFKatsuyaTChakrewarthySYamamotoKFujiokaA 2010 Common variants at the GCK, GCKR, G6PC2-ABCB11 and MTNR1B loci are associated with fasting glucose in two Asian populations. Diabetologia 53 299 308
34. OnumaHTabaraYKawamotoRShimizuIKawamuraR 2010 The GCKR rs780094 polymorphism is associated with susceptibility of type 2 diabetes, reduced fasting plasma glucose levels, increased triglycerides levels and lower HOMA-IR in Japanese population. J Hum Genet 55 600 604
35. HadaritsFKisfaliPMohasMMaaszADugaB 2011 Common functional variants of APOA5 and GCKR accumulate gradually in association with triglyceride increase in metabolic syndrome patients. Mol Biol Rep
36. HuCZhangRWangCYuWLuJ 2010 Effects of GCK, GCKR, G6PC2 and MTNR1B variants on glucose metabolism and insulin secretion. PLoS ONE 5 e11761 doi:10.1371/journal.pone.0011761
37. KozianDHBarthelACousinEBrunnhoferRAnderkaO 2010 Glucokinase-activating GCKR polymorphisms increase plasma levels of triglycerides and free fatty acids, but do not elevate cardiovascular risk in the Ludwigshafen Risk and Cardiovascular Health Study. Horm Metab Res 42 502 506
38. MohasMKisfaliPJaromiLMaaszAFeherE 2010 GCKR gene functional variants in type 2 diabetes and metabolic syndrome: do the rare variants associate with increased carotid intima-media thickness? Cardiovasc Diabetol 9 79
39. Orho-MelanderMMelanderOGuiducciCPerez-MartinezPCorellaD 2008 Common missense variant in the glucokinase regulatory protein gene is associated with increased plasma triglyceride and C-reactive protein but lower fasting glucose concentrations. Diabetes 57 3112 3121
40. Perez-MartinezPDelgado-ListaJGarcia-RiosAMc MonagleJGulsethHL 2011 Glucokinase regulatory protein genetic variant interacts with omega-3 PUFA to influence insulin resistance and inflammation in metabolic syndrome. PLoS ONE 6 e20555 doi:10.1371/journal.pone.0020555
41. TamCHMaRCSoWYWangYLamVK 2009 Interaction effect of genetic polymorphisms in glucokinase (GCK) and glucokinase regulatory protein (GCKR) on metabolic traits in healthy Chinese adults and adolescents. Diabetes 58 765 769
42. YangZWenJTaoXLuBDuY 2011 Genetic variation in the GCKR gene is associated with non-alcoholic fatty liver disease in Chinese people. Mol Biol Rep 38 1145 1150
43. SeithelAKleinKZangerUMFrommMFKonigJ 2008 Non-synonymous polymorphisms in the human SLCO1B1 gene: an in vitro analysis of SNP c.1929A>C. Mol Genet Genomics 279 149 157
44. BuchSSchafmayerCVolzkeHSeegerMMiquelJF 2010 Loci from a genome-wide analysis of bilirubin levels are associated with gallstone risk and composition. Gastroenterology 139 1942 1951 e1942
45. MiyaharaAOkamura-OhoYMiyashitaTHoshikaAYamadaM 2003 Genomic structure and alternative splicing of the insulin receptor tyrosine kinase substrate of 53-kDa protein. J Hum Genet 48 410 414
46. GalliganCLBaigEBykerkVKeystoneECFishEN 2007 Distinctive gene expression signatures in rheumatoid arthritis synovial tissue fibroblast cells: correlates with disease activity. Genes Immun 8 480 491
47. MillardTHDawsonJMacheskyLM 2007 Characterisation of IRTKS, a novel IRSp53/MIM family actin regulator with distinct filament bundling properties. J Cell Sci 120 1663 1672
48. DupuisJLangenbergCProkopenkoISaxenaRSoranzoN 2010 New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat Genet 42 105 116
49. WangLMuromotoNHayashiHMitaniYUeharaH 1997 Hyperinsulinemia but no diabetes in transgenic mice homozygously expressing the tyrosine kinase-deficient human insulin receptor. Biochem Biophys Res Commun 240 446 451
50. HotamisligilGSBudavariAMurrayDSpiegelmanBM 1994 Reduced tyrosine kinase activity of the insulin receptor in obesity-diabetes. Central role of tumor necrosis factor-alpha. J Clin Invest 94 1543 1549
51. KusariJTakataYHatadaEFreidenbergGKoltermanO 1991 Insulin resistance and diabetes due to different mutations in the tyrosine kinase domain of both insulin receptor gene alleles. J Biol Chem 266 5260 5267
52. BlockNEKomoriKRobinsonKADuttonSLLamCF 1991 Diabetes-associated impairment of hepatic insulin receptor tyrosine kinase activity: a study of mechanisms. Endocrinology 128 312 322
53. OdawaraMKadowakiTYamamotoRShibasakiYTobeK 1989 Human diabetes associated with a mutation in the tyrosine kinase domain of the insulin receptor. Science 245 66 68
54. ArnerPPollareTLithellHLivingstonJN 1987 Defective insulin receptor tyrosine kinase in human skeletal muscle in obesity and type 2 (non-insulin-dependent) diabetes mellitus. Diabetologia 30 437 440
55. SelvaDMHammondGL 2009 Peroxisome-proliferator receptor gamma represses hepatic sex hormone-binding globulin expression. Endocrinology 150 2183 2189
56. AltshulerDHirschhornJNKlannemarkMLindgrenCMVohlMC 2000 The common PPARgamma Pro12Ala polymorphism is associated with decreased risk of type 2 diabetes. Nat Genet 26 76 80
57. HakonarsonHGrantSF 2011 GWAS and its impact on elucidating the etiology of diabetes. Diabetes Metab Res Rev
58. SelvaDMHogeveenKNInnisSMHammondGL 2007 Monosaccharide-induced lipogenesis regulates the human hepatic sex hormone-binding globulin gene. J Clin Invest 117 3979 3987
59. ShihDQBussenMSehayekEAnanthanarayananMShneiderBL 2001 Hepatocyte nuclear factor-1alpha is an essential regulator of bile acid and plasma cholesterol metabolism. Nat Genet 27 375 382
60. YinLMaHGeXEdwardsPAZhangY 2011 Hepatic hepatocyte nuclear factor 4alpha is essential for maintaining triglyceride and cholesterol homeostasis. Arterioscler Thromb Vasc Biol 31 328 336
61. WilsonW3rdPardo-Manuel de VillenaFLyn-CookBDChatterjeePKBellTA 2004 Characterization of a common deletion polymorphism of the UGT2B17 gene linked to UGT2B15. Genomics 84 707 714
62. TurgeonDCarrierJSLevesqueEHumDWBelangerA 2001 Relative enzymatic activity, protein stability, and tissue distribution of human steroid-metabolizing UGT2B subfamily members. Endocrinology 142 778 787
63. MenardVEapOHarveyMGuillemetteCLevesqueE 2009 Copy-number variations (CNVs) of the human sex steroid metabolizing genes UGT2B17 and UGT2B28 and their associations with a UGT2B15 functional polymorphism. Hum Mutat 30 1310 1319
64. LevesqueEBeaulieuMGreenMDTephlyTRBelangerA 1997 Isolation and characterization of UGT2B15(Y85): a UDP-glucuronosyltransferase encoded by a polymorphic gene. Pharmacogenetics 7 317 325
65. SwansonCMellstromDLorentzonMVandenputLJakobssonJ 2007 The uridine diphosphate glucuronosyltransferase 2B15 D85Y and 2B17 deletion polymorphisms predict the glucuronidation pattern of androgens and fat mass in men. J Clin Endocrinol Metab 92 4878 4882
66. ChouinardSBarbierOBelangerA 2007 UDP-glucuronosyltransferase 2B15 (UGT2B15) and UGT2B17 enzymes are major determinants of the androgen response in prostate cancer LNCaP cells. J Biol Chem 282 33466 33474
67. ChouinardSPelletierGBelangerABarbierO 2004 Cellular specific expression of the androgen-conjugating enzymes UGT2B15 and UGT2B17 in the human prostate epithelium. Endocr Res 30 717 725
68. PathiSSJutooruIChadalapakaGSreevalsanSAnandS 2011 GT-094, a NO-NSAID, inhibits colon cancer cell growth by activation of a reactive oxygen species-microRNA-27a: ZBTB10-specificity protein pathway. Mol Cancer Res 9 195 202
69. KumarRManningJSpendloveHEKremmidiotisGMcKirdyR 2006 ZNF652, a novel zinc finger protein, interacts with the putative breast tumor suppressor CBFA2T3 to repress transcription. Mol Cancer Res 4 655 665
70. KumarRSelthLSchulzRBTayBSNeilsenPM 2011 Genome-wide mapping of ZNF652 promoter binding sites in breast cancer cells. J Cell Biochem
71. KumarRCheneyKMNeilsenPMSchulzRBCallenDF 2010 CBFA2T3-ZNF651, like CBFA2T3-ZNF652, functions as a transcriptional corepressor complex. FEBS Lett 584 859 864
72. CallenDFRicciardelliCButlerMStapletonAStahlJ 2010 Co-expression of the androgen receptor and the transcription factor ZNF652 is related to prostate cancer outcome. Oncol Rep 23 1045 1052
73. BaldassarreGRomanoAArmenanteFRambaldiMPaolettiI 1997 Expression of teratocarcinoma-derived growth factor-1 (TDGF-1) in testis germ cell tumors and its effects on growth and differentiation of embryonal carcinoma cell line NTERA2/D1. Oncogene 15 927 936
74. BurgerHGDudleyECCuiJDennersteinLHopperJL 2000 A prospective longitudinal study of serum testosterone, dehydroepiandrosterone sulfate, and sex hormone-binding globulin levels through the menopause transition. J Clin Endocrinol Metab 85 2832 2838
75. SchijfCPvan der MoorenMJDoesburgWHThomasCMRollandR 1993 Differences in serum lipids, lipoproteins, sex hormone binding globulin and testosterone between the follicular and the luteal phase of the menstrual cycle. Acta Endocrinol (Copenh) 129 130 133
76. LinkEParishSArmitageJBowmanLHeathS 2008 SLCO1B1 variants and statin-induced myopathy–a genomewide study. N Engl J Med 359 789 799
77. ChambersJCZhangWSehmiJLiXWassMN 2011 Genome-wide association study identifies loci influencing concentrations of liver enzymes in plasma. Nat Genet 43 1131 1138
78. LeeDKKuriharaIJeongJWLydonJPDeMayoFJ 2010 Suppression of ERalpha activity by COUP-TFII is essential for successful implantation and decidualization. Mol Endocrinol 24 930 940
Štítky
Genetika Reprodukční medicína
Článek Allelic Heterogeneity and Trade-Off Shape Natural Variation for Response to Soil MicronutrientČlánek The Chicken Frizzle Feather Is Due to an α-Keratin () Mutation That Causes a Defective RachisČlánek A Trans-Species Missense SNP in Is Associated with Sex Determination in the Tiger Pufferfish, (Fugu)
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2012 Číslo 7- Akutní intermitentní porfyrie
- Transthyretinová amyloidóza z pohledu neurologa a kardiologa aneb jak se vyhnout „misdiagnostice“?
- Prof. Petr Urbánek: Potřebujeme najít pacienty s nediagnostikovanou akutní intermitentní porfyrií
- S MUDr. Evou Flachsovou, Ph.D., o nové éře cílené terapie měnící prognózu pacientů s primární hyperoxalurií
- Vitamín B6 jako prevence kolorektálního karcinomu
-
Všechny články tohoto čísla
- Functional Evolution of Mammalian Odorant Receptors
- Oocyte Family Trees: Old Branches or New Stems?
- Allelic Heterogeneity and Trade-Off Shape Natural Variation for Response to Soil Micronutrient
- Guidelines for Genome-Wide Association Studies
- GWAS Identifies Novel Susceptibility Loci on 6p21.32 and 21q21.3 for Hepatocellular Carcinoma in Chronic Hepatitis B Virus Carriers
- DNA Methyltransferases Are Required to Induce Heterochromatic Re-Replication in Arabidopsis
- Genomic Data Reveal a Complex Making of Humans
- Let-7b/c Enhance the Stability of a Tissue-Specific mRNA during Mammalian Organogenesis as Part of a Feedback Loop Involving KSRP
- The Secreted Immunoglobulin Domain Proteins ZIG-5 and ZIG-8 Cooperate with L1CAM/SAX-7 to Maintain Nervous System Integrity
- RsfA (YbeB) Proteins Are Conserved Ribosomal Silencing Factors
- Gene Conversion Occurs within the Mating-Type Locus of during Sexual Reproduction
- The Chicken Frizzle Feather Is Due to an α-Keratin () Mutation That Causes a Defective Rachis
- Meta-Analysis of Genome-Wide Scans for Total Body BMD in Children and Adults Reveals Allelic Heterogeneity and Age-Specific Effects at the Locus
- Balancing Selection at the Tomato Guardee Gene Family Maintains Variation in Strength of Pathogen Defense
- Large-Scale Introgression Shapes the Evolution of the Mating-Type Chromosomes of the Filamentous Ascomycete
- OSD1 Promotes Meiotic Progression via APC/C Inhibition and Forms a Regulatory Network with TDM and CYCA1;2/TAM
- Intact p53-Dependent Responses in miR-34–Deficient Mice
- FANCJ/BACH1 Acetylation at Lysine 1249 Regulates the DNA Damage Response
- CED-10/Rac1 Regulates Endocytic Recycling through the RAB-5 GAP TBC-2
- Histone H2A Mono-Ubiquitination Is a Crucial Step to Mediate PRC1-Dependent Repression of Developmental Genes to Maintain ES Cell Identity
- F-Box Protein Specificity for G1 Cyclins Is Dictated by Subcellular Localization
- The Gene Encodes a Nuclear Protein That Affects Alternative Splicing
- A Key Role for Chd1 in Histone H3 Dynamics at the 3′ Ends of Long Genes in Yeast
- Genome-Wide Association Analysis in Asthma Subjects Identifies as a Novel Bronchodilator Response Gene
- GRHL3/GET1 and Trithorax Group Members Collaborate to Activate the Epidermal Progenitor Differentiation Program
- Brain-Specific Rescue of Reveals System-Driven Transcriptional Rhythms in Peripheral Tissue
- Recent Loss of Self-Incompatibility by Degradation of the Male Component in Allotetraploid
- Pregnancy-Induced Noncoding RNA () Associates with Polycomb Repressive Complex 2 and Regulates Mammary Epithelial Differentiation
- The HEI10 Is a New ZMM Protein Related to Zip3
- The SCF Ubiquitin E3 Ligase Ubiquitylates Sir4 and Functions in Transcriptional Silencing
- Induction of Cytoprotective Pathways Is Central to the Extension of Lifespan Conferred by Multiple Longevity Pathways
- Role of Architecture in the Function and Specificity of Two Notch-Regulated Transcriptional Enhancer Modules
- Loss of ATRX, Genome Instability, and an Altered DNA Damage Response Are Hallmarks of the Alternative Lengthening of Telomeres Pathway
- A Regulatory Loop Involving PAX6, MITF, and WNT Signaling Controls Retinal Pigment Epithelium Development
- The Three Faces of Riboviral Spontaneous Mutation: Spectrum, Mode of Genome Replication, and Mutation Rate
- Unmet Expectations: miR-34 Plays No Role in p53-Mediated Tumor Suppression In Vivo
- A Genome-Wide Association Meta-Analysis of Circulating Sex Hormone–Binding Globulin Reveals Multiple Loci Implicated in Sex Steroid Hormone Regulation
- The Role of Rice HEI10 in the Formation of Meiotic Crossovers
- A Trans-Species Missense SNP in Is Associated with Sex Determination in the Tiger Pufferfish, (Fugu)
- Influences Bone Mineral Density, Cortical Bone Thickness, Bone Strength, and Osteoporotic Fracture Risk
- Evidence of Inbreeding Depression on Human Height
- Comparative Genomics of Plant-Associated spp.: Insights into Diversity and Inheritance of Traits Involved in Multitrophic Interactions
- Detecting Individual Sites Subject to Episodic Diversifying Selection
- Regulates Rhodopsin-1 Metabolism and Is Required for Photoreceptor Neuron Survival
- Identification of Chromatin-Associated Regulators of MSL Complex Targeting in Dosage Compensation
- Three Dopamine Pathways Induce Aversive Odor Memories with Different Stability
- TDP-1/TDP-43 Regulates Stress Signaling and Age-Dependent Proteotoxicity in
- Rapid Turnover of Long Noncoding RNAs and the Evolution of Gene Expression
- The Yeast Rab GTPase Ypt1 Modulates Unfolded Protein Response Dynamics by Regulating the Stability of RNA
- Histone H2B Monoubiquitination Facilitates the Rapid Modulation of Gene Expression during Arabidopsis Photomorphogenesis
- Cellular Variability of RpoS Expression Underlies Subpopulation Activation of an Integrative and Conjugative Element
- Genetic Variants in , , and Influence Male Recombination in Cattle
- Differential Impact of the HEN1 Homolog HENN-1 on 21U and 26G RNAs in the Germline of
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Guidelines for Genome-Wide Association Studies
- The Role of Rice HEI10 in the Formation of Meiotic Crossovers
- Identification of Chromatin-Associated Regulators of MSL Complex Targeting in Dosage Compensation
- GWAS Identifies Novel Susceptibility Loci on 6p21.32 and 21q21.3 for Hepatocellular Carcinoma in Chronic Hepatitis B Virus Carriers
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Revma Focus: Spondyloartritidy
nový kurz
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MUDr. Irena Krčmová, CSc.
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



