-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
R-loops Associated with Triplet Repeat Expansions Promote Gene Silencing in Friedreich Ataxia and Fragile X Syndrome
Friedreich ataxia and Fragile X syndrome are among 40 human diseases associated with expansion of repeated sequences. In both disorders repeat expansion leads to gene silencing, the molecular mechanism of which is not well understood, impeding the development of specific therapies to treat these disorders. It is proposed that formation of unusual DNA structures (RNA/DNA hybrids, or R-loops) over repeat regions may play a role, but their molecular function has not been investigated in vivo. We show that R-loops form on expanded repeats of FXN and FMR1 genes in cells from FRDA and FXS patients. These R-loops are stable, correlate with repressive chromatin marks and hinder FXN transcription in patient cells. We studied the relationship between repressive chromatin and R-loops. Decrease in the amount of repressive chromatin has no effect on R-loop levels. In contrast, increase in the R-loops leads to transcriptional silencing of FXN gene and formation of repressive chromatin, providing a direct molecular link between R-loops and pathology of expansion diseases. This discovery is important for understanding the basic molecular mechanism underlying the pathology of expansion diseases. The ability of R-loops to trigger transcriptional silencing makes them an attractive target for future therapeutic approaches to treat these diseases.
Published in the journal: . PLoS Genet 10(5): e32767. doi:10.1371/journal.pgen.1004318
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1004318Summary
Friedreich ataxia and Fragile X syndrome are among 40 human diseases associated with expansion of repeated sequences. In both disorders repeat expansion leads to gene silencing, the molecular mechanism of which is not well understood, impeding the development of specific therapies to treat these disorders. It is proposed that formation of unusual DNA structures (RNA/DNA hybrids, or R-loops) over repeat regions may play a role, but their molecular function has not been investigated in vivo. We show that R-loops form on expanded repeats of FXN and FMR1 genes in cells from FRDA and FXS patients. These R-loops are stable, correlate with repressive chromatin marks and hinder FXN transcription in patient cells. We studied the relationship between repressive chromatin and R-loops. Decrease in the amount of repressive chromatin has no effect on R-loop levels. In contrast, increase in the R-loops leads to transcriptional silencing of FXN gene and formation of repressive chromatin, providing a direct molecular link between R-loops and pathology of expansion diseases. This discovery is important for understanding the basic molecular mechanism underlying the pathology of expansion diseases. The ability of R-loops to trigger transcriptional silencing makes them an attractive target for future therapeutic approaches to treat these diseases.
Introduction
Around forty human diseases are associated with expanded repeat sequences [1]. Friedreich ataxia (FRDA) is the most frequent autosomal recessive ataxia (2–4 cases/100,000), caused by a GAA expansion in the first intron of the frataxin (FXN) gene, which encodes a mitochondrial protein involved in iron-sulfur cluster biogenesis [2], [3]. The GAA expansion leads to reduced levels of FXN mRNA and protein [4]–[6]. Several mechanisms mediating FXN transcriptional silencing have been proposed, including the formation of unusual DNA structures (triplex DNA and RNA/DNA hybrids) and repressive heterochromatin over expanded repeats [5]–[10].
RNA/DNA hybrids (R-loops) are formed during transcription, when nascent RNA hybridizes to the DNA template behind the elongating RNA polymerase (Pol II). R-loops are detected in organisms from bacteria to humans and implicated in many processes [11]. In mammalian cells, R-loops were originally discovered in the immuno-globulin class switch regions, essential for generating the antibody diversity in mouse activated B cells [12], [13]. R-loops also accumulate in cells depleted of the key splicing factor SRSF1, resulting in genome instability and appearance of double-strand breaks [14]. Recent studies demonstrated that R-loops are enriched over CpG promoters and may be involved in protection of these regions from DNA methylation and maintaining the hypomethylated state of CpG promoters [15]. We recently showed that R-loops formed over the G-rich pause sites downstream of the polyA signal in human genes are essential for the process of transcriptional termination of RNA Pol II [16]. Furthermore RNA/DNA hybrids are induced at GAA repeats following in vitro transcription and in bacteria [17], [18]. Also R-loops formed on plasmids containing CTG/CAG repeats in E.coli and mini-gene constructs in human cells promoted repeat instability, pointing towards their role in disease pathology [19], [20]. However, the direct involvement of R-loops on endogenous expanded alleles in the pathology of FRDA has not yet been investigated in vivo.
Our study shows that RNA/DNA hybrids (R-loops) form on expanded repeats of endogenous FXN and FMR1 genes, associated with Friedreich ataxia and Fragile X (FXS) disorders, in patient cells. These transcription-dependent R-loops are resistant to cellular degradation and co-localise with repressive H3K9me2 chromatin marks, characteristic of these diseases. Using nascent nuclear run-on analysis we show that R-loops over expanded repeats impede RNA Polymerase II transcription of the FXN gene in patient cells. We investigated the interplay between repressive chromatin marks and R-loops on the FXN gene. We show that a decrease in repressive H3K9me2 chromatin mark has no effect on R-loop levels and FXN transcription. In contrast, increasing R-loop levels leads to transcriptional repression of FXN gene, providing a direct molecular link between R-loops and pathology of FRDA. These data suggest that R-loops formed over expanded repeats act as an initial trigger to promote FXN and FMR1 silencing, and represent a common feature of nucleotide expansion diseases, contributing to their pathology in vivo.
Results
FXN transcriptional initiation and elongation defect in FRDA cells
We examined transcriptional regulation of the FXN gene in immortalized lymphoblastoid cells derived from FRDA patients, where FXN mRNA expression is reduced by ∼80% (Figure 1A–C). Pol II chromatin immuno-precipitation (ChIP) analysis in these cells showed that Pol II is enriched over the exon 1, positioned at the major transcriptional start site (TSS2) in lymphoblasts, correlating with the promoter-specific histone H3 depleted region [4] (Figure 1D, S1). Pol II levels over exon 1 were significantly reduced in FRDA cells. Similarly, using quantitative RT-PCR (RT-qPCR) in three independent control and three FRDA cell lines, a dramatic reduction in nascent RNA was detected over exon 1 in FRDA cells, further confirming a defect in transcription initiation (Figure 1E, S2A and S2B left panels). We also observed ∼10-fold reduction in the nascent RNA downstream of the expansion in regions D–G in FRDA cells. Overall Pol II ChIP and RT-qPCR results suggest transcriptional initiation and elongation defects triggered by expanded repeats, in line with previous reports [4], [6], [21].
Fig. 1. R-loops are formed over expanded repeats of FXN gene in FRDA cells. 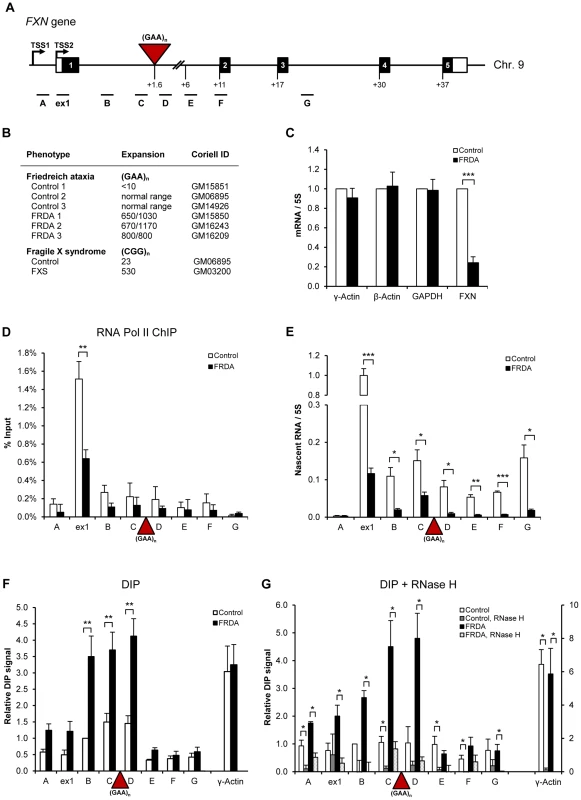
A. Diagram of FXN gene. Black boxes are exons, white boxes are 5′ and 3′UTRs, lines are introns, red triangle is (GAA)n expansion. TSS2 is the major transcriptional start site in lymphoblastoid cells. qPCR amplicons are shown below the diagram. Numbers indicate the distances to TSS2 in kilobases. B. Cell lines used in the study. The repeat sizes are indicated. C. RT-qPCR analysis of γ-actin, β-actin, GAPDH and FXN mRNAs in control (GM15851) and FRDA (GM15850) cells. Values are normalised to 5S rRNA and relative to control cells. D. RNA Pol II ChIP in control (GM15851) and FRDA (GM15850) cells. E. RT-qPCR analysis of FXN nascent RNA in control (GM15851) and FRDA (GM15850) cells, normalised to 5S rRNA and relative to ex1 RNA in control cells. F. DIP on endogenous FXN gene in control (GM15851) and FRDA (GM15851) cells. γ-actin is positive control. G. R-loops are sensitive to RNase H digestion. DIP samples were treated with 25 U of recombinant E.coli RNase H (NEB, M0297S) for 6 hours at 37°C. γ-actin is positive control. Bars in C–G are average values +/− SEM (n>3). R-loops are formed over expanded repeat regions of FXN gene in vivo
Recently we established the DNA immuno-precipitation (DIP) method, which allows detection of R-loops on endogenous human genes in vivo using S9.6 antibody which recognizes RNA/DNA hybrids [16], [22]. Here we employed DIP to investigate R-loop distribution on the FXN gene. As a positive control we used the intron 1 region of the γ-actin gene, where high levels of R-loops are detected [15]. Significantly, we observed ∼3-fold enrichment of R-loops over regions B, C and D in the FXN intron 1 in FRDA cells, compared to control cells (Figure 1F). R-loops were concentrated over the expanded repeat region and were low in the downstream regions E–G. γ-actin R-loop levels were similar in control and FRDA cells (Figure 1F). Similar R-loop enrichment over expanded GAA repeats was detected in two additional independent FRDA cell lines (Figure S2A–B, right panels). Interestingly, when we compared the DIP data from all control and FRDA cell lines we observed that the level of R-loops correlates with expansion length (Figure S2C). To confirm the specificity of the DIP signal, we treated the samples with RNase H, which specifically degrades the RNA in RNA/DNA hybrids, prior to immuno-precipitation. Following RNase H digestion, the signal was strongly reduced for control γ-actin and FXN regions, suggesting that genuine R-loops are formed over the expanded GAA repeats (Figure 1G). High level of R-loops detected in FRDA cells may also suggest that these structures are particularly stable over FXN expanded repeats. Therefore, R-loops could act in cis to affect FXN gene expression in FRDA cells.
R-loops are stable and impede Pol II transcription on FXN gene
To understand the function of R-loops in FRDA pathology, we further characterized these structures over the expanded FXN allele. In particular, we treated cells with the transcriptional inhibitor actinomycin D. Following this treatment for 21 hours we observed an ∼80% reduction in γ-actin nascent RNA and R-loop signal, suggesting that γ-actin R-loops are quickly turned over in the cell (Figure 2A, B). In contrast, although nascent FXN RNA decreased following actinomycin D treatment, no change in R-loop levels was detected. However, we did finally observe a significant decrease in the level of R-loops over expanded repeats following prolonged treatment with actinomycin D for 48 hours (Figure S3). Overall these results suggest that R-loops associated with FXN expanded repeats are resistant to degradation. This may relate to the expanded GAA repeat property of transcriptional repression and repeat instability. We also observed that enrichment of R-loops correlated with highest peaks of the repressive histone modification H3K9me2 over the FXN regions B–D in FRDA cells (Figure 2C and 1F). This suggests that R-loops over expanded GAA repeats may functionally associate with repressive heterochromatin and be involved in mediating transcriptional repression.
Fig. 2. R-loops are stable and impede Pol II transcription on FXN gene. 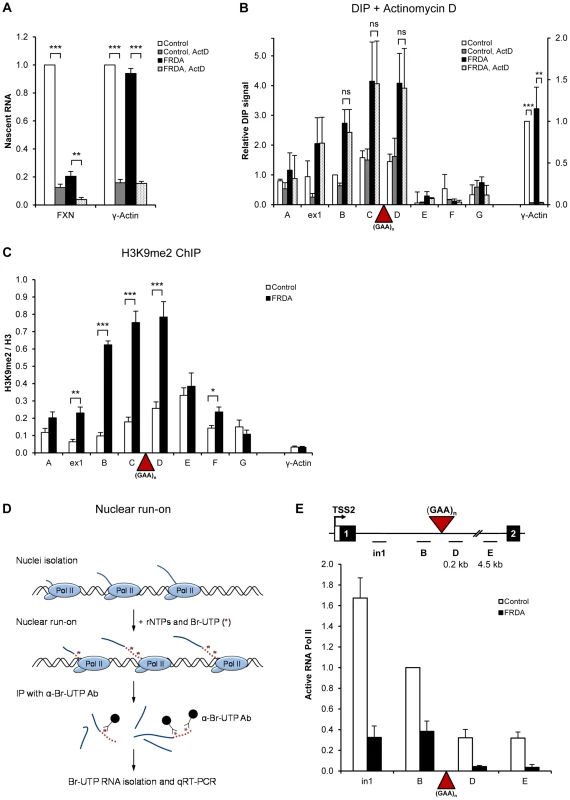
A. RT-qPCR analysis of nascent γ-actin and FXN RNA from control and FRDA cells treated with 5 µg/ml of actinomycin D for 21 hours. Values are relative to untreated control cells. B. DIP on FXN gene in control and FRDA cells treated with 5 µg/ml of actinomycin D for 21 hours. γ-actin is positive control. C. H3K9me2 ChIP in control and FRDA cells. H3K9me2 levels were normalized to the total H3 levels. γ-actin is used as background control. D. Diagram depicting the Br-UTP nuclear run-on (NRO) method. E. Br-UTP nuclear run-on in two control (GM15851, GM14926) and two FRDA (GM15850 and GM16243) cells, normalised to the region B in control cells. Bars in A–C and E are average values +/− SEM (n>3). Previously we showed that R-loops formed at the 3′ends of human genes promote transcriptional termination of RNA Pol II [16]. We therefore investigated if R-loops over FXN expanded repeats affect Pol II elongation. Here we employed nuclear run-on (NRO) analysis with Br-UTP labelled nucleotide [16], which measures actively transcribing Pol II (Figure 2D), in contrast to ChIP, which detects the total Pol II level on the gene. Using NRO we observed a substantial decrease in active transcription upstream of the GAA expansion (regions in1 and B) in FRDA cells, confirming our Pol II ChIP results (Figure 2E and 1D). In addition, we also detected ∼3-fold reduction in active transcription in FRDA cells over FXN regions D and E, positioned 210 nt and 4.5 kb downstream of expansion, respectively (Figure 2E). This elongation defect is not due to the increased distance caused by GAA expansion (∼3 kb), since we observed no decrease in active transcription between regions D to E, separated by ∼4.3 kb, in both cell lines. This suggests that expanded repeats directly interfere with active Pol II transcription in FRDA cells.
R-loops on expanded repeats of integrated FXN gene are sensitive to RNase H1 over-expression in HEK293 cells
To test if expanded GAA repeats trigger the formation of R-loops in a different genomic location, we employed HEK293 cells, containing a copy of the FXN gene with either six (FXN-Luc) or ∼310 (FXN-GAA-Luc) GAA repeats, fused to the luciferase gene, integrated on chromosome 1, while the endogenous FXN gene lies on chromosome 9 (Figure 3A) [23]. We confirmed the presence of the GAA expansion using PCR on genomic DNA extracted from these cells. As demonstrated in Figure 3B, the FXN-GAA-Luc cell line indeed contains ∼310 expanded repeats. The presence of GAA repeats caused a reduction of ∼37% in FXN-luciferase nascent RNA levels as determined by RT-qPCR (Figure 3C) and an increase in the repressive histone modification H3K9me2 (Figure S4A), recapitulating repression of gene expression seen in FRDA cells. This smaller reduction in the RNA levels in FXN-GAA-Luc cells compared to FRDA lymphoblastoid cells can be explained by low number of repeats (only ∼310) on the integrated FXN copy.
Fig. 3. Over-expressed RNase H1 resolves R-loops formed on FXN expanded repeats in HEK293 cells. 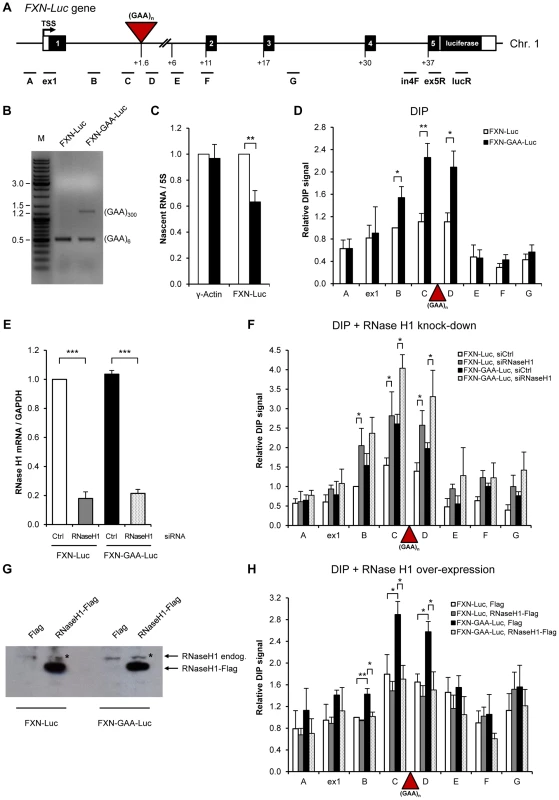
A. Diagram of the FXN-Luc gene, containing 6 (FXN-Luc) or 310 GAA repeats (FXN-GAA-Luc), integrated on the chromosome 1 of HEK293 cells. Frataxin gene was fused to the luciferase at the beginning of the FXN exon 5. Black boxes are exons, white boxes are 5′ and 3′UTRs, lines are introns, red triangle is (GAA)n expansion. TSS is the transcriptional start site. qPCR amplicons are shown below the diagram. Numbers indicate the distances to TSS in kilobases. B. Size of GAA expansion determined by PCR analysis on genomic DNA from FXN-Luc and FXN-GAA-Luc cell lines, using GAA104F and GAA629R primers. PCR products were run on 1% agarose gel. M denotes the marker lane. FXN-Luc and FXN-GAA-Luc cells contain endogenous wild type FXN gene [23], giving rise to the PCR product of 0.5 kb. C. FXN and γ-actin nascent RNA levels in FXN-Luc (white bars) and FXN-GAA-Luc (black bars) HEK293 cells, determined by RT-qPCR and normalised to 5S rRNA. The level of FXN and γ-actin nascent RNA in FXN-Luc cells was taken as 1. LucR primer was used for the reverse transcription reaction. qPCR was carried using in4F and ex5R primers, shown in A. D. DIP analysis on FXN-Luc gene in FXN-Luc (white bars) and FXN-GAA-Luc (black bars) HEK293 cells using RNA/DNA hybrid-specific S9.6 antibody. E. RT-qPCR analysis of RNase H1 mRNA from FXN-Luc and FXN-GAA-Luc cells, treated with control and RNase H1 siRNAs. Values are normalised to GAPDH mRNA and are relative to FXN-Luc cells, treated with control siRNA. F. DIP analysis on FXN-Luc gene in FXN-Luc and FXN-GAA-Luc HEK293 cells, treated with control and RNase H1 siRNAs. G. Western blot analysis of 50 µg of protein extracts obtained from FXN-Luc and FXN-GAA-Luc cells transfected with Flag and RNase H1-Flag expression plasmids. Western blot was probed with anti-RNase H1 antibody. * denotes endogenous RNase H1 protein. H. DIP analysis on FXN-Luc gene in FXN-Luc and FXN-GAA-Luc HEK293 cells transfected with Flag or RNase H1-Flag expression plasmids. Bars in C–F and H represent the average values from at least three independent experiments +/− SEM. Next we investigated if R-loops are formed on expanded repeats of the integrated FXN copy in HEK293 cells using DIP analysis (Figure 3D). Similar to patient-derived FRDA lymphoblast cells, we observed 2–3-fold increase in the level of R-loops over expanded repeats region (amplicons B, C and D) in FXN-GAA-Luc cells. This suggests that R-loops are formed on transcribed expanded GAA repeats, independently of their genomic location. To confirm the specificity of this R-loop signal, we employed RNAi to knock down endogenous RNase H1 enzyme, which specifically degrades the RNA in RNA/DNA hybrids [24]. Following depletion of RNase H1 in HEK293 cells, we observed a significant increase in the R-loop signal, suggesting that endogenous RNase H1 can degrade R-loops formed over FXN gene in vivo (Figure 3E, F).
Next we wanted to test if RNase H1 over-expression can resolve R-loops formed on expanded GAA repeats. To this end we over-expressed Flag and RNase H1-Flag constructs in FXN-Luc and FXN-GAA-Luc cells. High level of RNase H1 over-expression was confirmed by RT-qPCR (Figure S4B) and western blot analysis (Figure 3G). Interestingly, following RNase H1 over-expression, we observed a reduction of R-loop signal over the GAA expansion (Figure 3H). In line with these observations, RNase H1 over-expression resulted in up-regulation of FXN transcription from the expanded allele (Figure S4C). This suggests that R-loops formed over expanded repeats in vivo can be resolved by over-expressed RNase H1 and removal of R-loops leads to increase in FXN gene expression.
R-loops trigger FXN transcriptional repression in vivo
Previous studies have demonstrated that expanded FXN GAA repeats are associated with increased levels of heterochromatin marks [4], [6], [7]. To investigate the relationship between R-loops and repressive heterochromatin marks formed on the expanded FXN allele, we employed histone methyltransferase inhibitor BIX-01294, previously shown to reduce the level of H3K9me2 over repeat regions [21]. Following BIX-01294 treatment, we observed a significant reduction in the levels of H3K9me2 chromatin mark (Figure 4A), similar to previous reports [21]. Significantly, following BIX-01294 treatment the level of R-loops over the expanded repeat region remained unchanged (Figure 4B). Similarly, reduction in H3K9me2 chromatin mark had no effect on FXN nascent RNA level (Figure 4C). These data suggest that H3K9me2 chromatin modification is not directly responsible for the FXN transcriptional repression and is likely to be a consequence of the reduced transcription or caused by R-loop formation.
Fig. 4. R-loops are not affected by changes in H3K9 dimethylation. 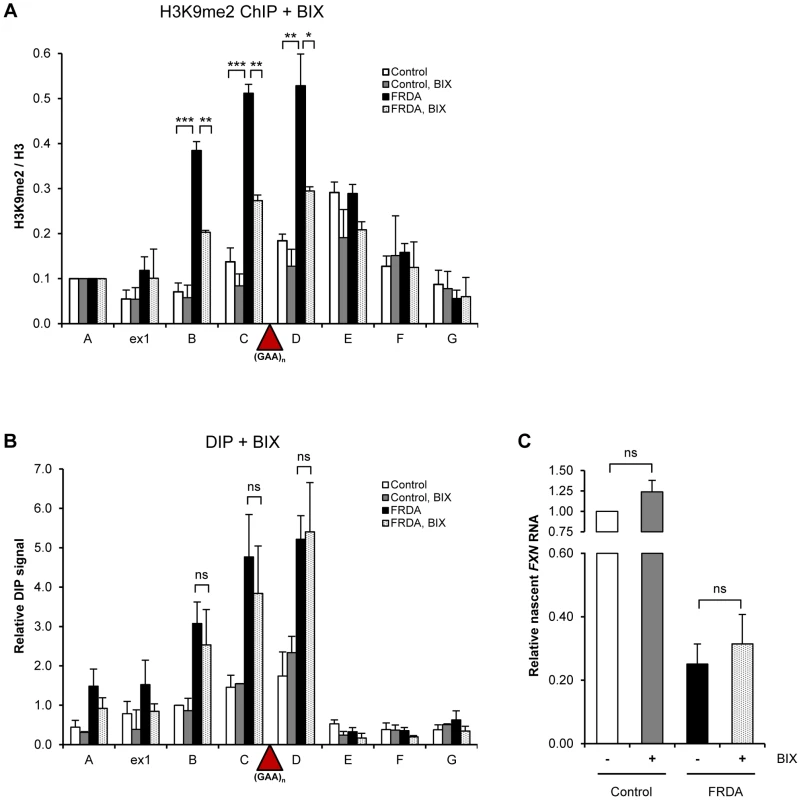
A. H3K9me2 ChIP in control and FRDA cells, treated with 4 µM BIX-01294 for 72 h. H3K9me2 levels were normalized to the total H3 levels and relative to amplicon FXN A, not affected by the treatment. B. DIP analysis in control and FRDA cells, treated with 4 µM BIX-01294 for 72 h. C. RT-qPCR analysis of FXN nascent RNA in control and FRDA cells, treated with 4 µM BIX-01294 for 72 h. Values are relative to untreated control cells and normalized to γ-actin nascent RNA. Bars in A–C are average values +/− SEM (n>3). To investigate the ability of R-loops to directly trigger FXN transcriptional repression, we took advantage of camptothecin (CPT), a specific inhibitor of DNA Topoisomerase I (Top1), an enzyme which relieves transcription-induced DNA supercoiling. Loss of Top1 enhances R-loop formation in yeast and human cells [25], [26]. Following CPT treatment, we observed an increase in R-loop formation over the expanded repeat region in FRDA cells while R-loop levels in FXN regions E–G remained unchanged (Figure 5A). This effect was consistent between different patient-derived cell lines (Figure S5A). We detected no effect on R-loop levels in control cells, demonstrating the specificity of CPT treatment to expanded repeats (Figure 5A and S5A). The ability of CPT to increase R-loop levels was not due to CPT-induced covalent links between Top1 and DNA, since Top1 knock-down also resulted in R-loop accumulation (Figure 5E, F).
Fig. 5. R-loops trigger transcriptional repression of FXN gene. 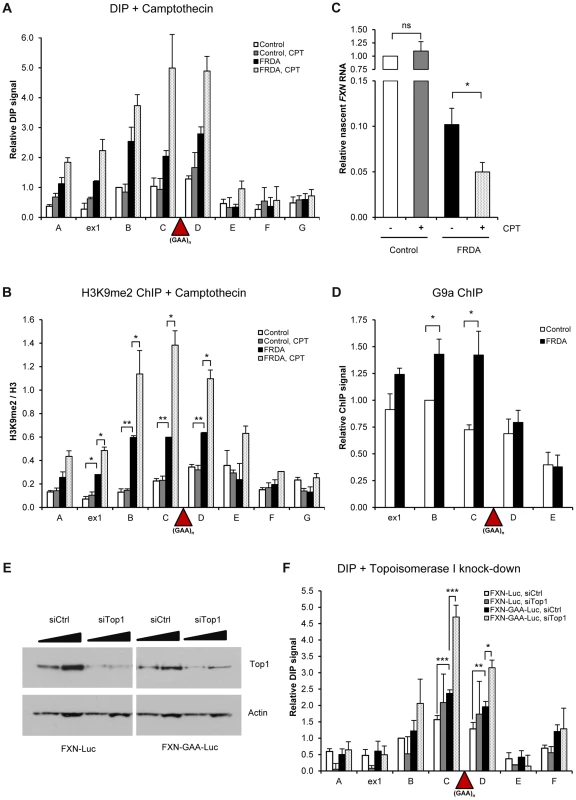
A. DIP analysis on FXN gene in control and FRDA cells, treated with 10 µM camptothecin (CPT) for 6 hours. B. H3K9me2 ChIP on FXN gene in control and FRDA cells, treated with 10 µM camptothecin (CPT) for 6 hours. H3K9me2 levels were normalized to the total H3 levels. C. RT-qPCR analysis of FXN nascent RNA in control and FRDA cells, treated with 10 µM camptothecin for 6 hours. Values are relative to untreated control cells and normalized to γ-actin nascent RNA. D. G9a ChIP on FXN gene in control and FRDA cells. G9a levels are normalised relative to amplicon B in control cells. E. Western blot analysis of 20 and 40 µg of protein extracts obtained from FXN-Luc and FXN-GAA-Luc cells, treated with control and Top1 siRNAs. Western blot was probed with anti-Top1 and anti-actin antibody. F. DIP analysis on FXN gene in FXN-Luc and FXN-GAA-Luc HEK293 cells, treated with control and Top1 siRNAs. Bars in A–D and F are average values +/− SEM (n>3). Interestingly, increase in the level of R-loops coincided with increase in the amount of repressive H3K9me2 mark over FXN regions B–D surrounding the expansion in FRDA cells (Figure 5B and S5B). This also resulted in down-regulation of nascent FXN RNA in FRDA cells, but not in control cells, as observed in three independent control and FRDA cell lines (Figure 5C and S5C–D). These data suggest that R-loops at expanded repeats directly trigger FXN transcriptional repression and promote formation of repressive H3K9me2 marks. To understand the molecular mechanism of this process, we investigated the binding of G9a histone methyltransferase, which deposits H3K9me2 marks on histones, to FXN gene. Interestingly, we observed that G9a is enriched over the expanded region in FRDA cells (Figure 5D). This suggests that R-loops may recruit G9a to the expanded repeat regions, thereby promoting the formation of repressive H3K9me2 marks.
R-loops are formed over (CGG)n expanded repeats of FMR1 gene
To test if R-loop formation is a general feature of trinucleotide expansion diseases, we also examined the FMR1 gene. In Fragile X syndrome patients, the FMR1 allele containing a (CGG)n>200 expansion in the 5′UTR is fully methylated and transcriptionally silenced [27]. Therefore to investigate the potential role of R-loops in FXS, FMR1 transcription was reactivated by treatment with the DNA methylation inhibitor 5-aza-2′-deoxycytidine (5-azadC). This resulted in expression of FMR1 mRNA in FXS cells to 25% of control cells, as previously reported [28]. However, FMR1 expression was unchanged in control cells (Figure 6B). Using DIP, we detected low R-loop signal in control and untreated FXS cells (Figure 6C). Following 5-azadC treatment, we observed ∼4-fold increase in R-loops over the exon 1 region upstream of the expansion in FXS cells, while no significant changes were detected in control cells. The specificity of the DIP signal was confirmed by RNase H treatment (Figure 6D). These data suggest that R-loops are transcriptionally-dependent and localise to the expanded (CGG) repeat region. Since inhibition of DNA methylation only partially reactivates expanded FMR1 allele in FXS cells, it is possible that R-loops at expanded (CGG) repeats prevent full reactivation.
Fig. 6. R-loops are formed over (CGG)n expanded repeats of FMR1 gene. 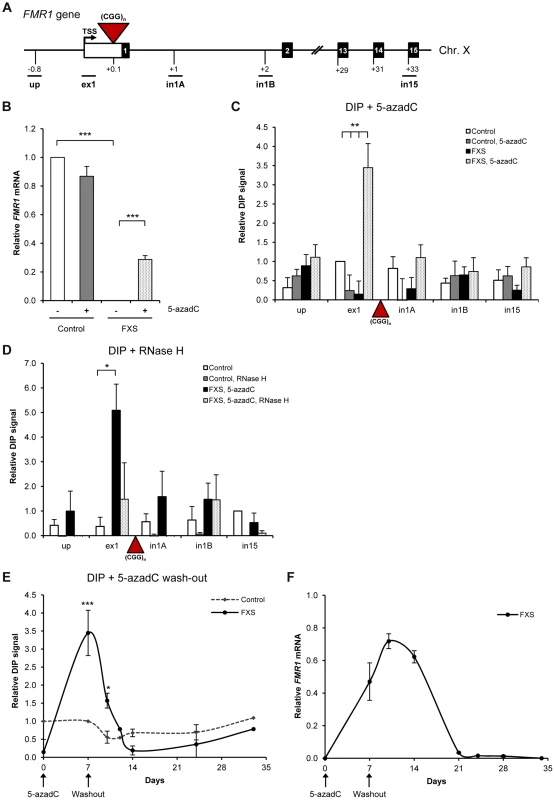
A. Diagram of FMR1 gene. Black boxes are exons, white box is 5′ UTR and lines are introns. Red triangle is (CGG)n expansion. qPCR amplicons are shown below the diagram. TSS is the transcriptional start site. Numbers indicate the distances to TSS in kilobases. B. RT-qPCR analysis of FMR1 mRNA in control and FXS cells, treated with 1 µM 5-azadC for 7 days, normalized to GAPDH. C. DIP analysis on endogenous FMR1 gene in control and FXS cells, treated with 1 µM 5-azadC for 7 days. Values are relative to ex1 region in control untreated cells. D. FMR1 R-loops are sensitive to RNase H digestion, following the treatment with 25 U of RNase H for 6 hours at 37°C prior to immuno-precipitation. Values are relative to in15 region in control untreated cells. E. R-loop kinetics on exon 1 of FMR1 gene in control and FXS cells during the process of transcriptional re-activation with 1 µM 5-azadC (7 days) followed by 5-azadC wash out with drug-free media (28 days). Values are relative to ex1 region in control untreated cells on day 7. F. RT-qPCR analysis of FMR1 mRNA in control and FXS cells, treated with 1 µM 5-azadC (7 days) followed by 5-azadC wash out with drug-free media (28 days). The level of FMR1 mRNA in control cells is taken as 1. Bars in B–D are average values +/− SEM (n>3). To further characterize the relationship between R-loops and FMR1 expression, we carried out kinetic experiments. In particular, we studied R-loop and FMR1 mRNA levels during the process of transcriptional re-activation with 5-azadC treatment (7 days) followed by 5-azadC wash out with drug-free media for 28 days (Figure 6E, F). We observed that the R-loop levels over the exon 1 of FMR1 gene stayed at the background during activation and wash-out period in control cells (dotted line in Figure 6E). In FXS cells, the R-loops were at their peak during the re-activation procedure with 5-azadC on day 7. After removal of 5-azadC, R-loop levels gradually diminished and completely disappeared after 7 days (day 14 of the full experiment). This pattern of R-loop dynamics correlated with FMR1 expression profile (Figure 6F), suggesting that R-loops are associated with FMR1 gene regulation.
Discussion
We demonstrate that R-loops are formed over endogenous expanded (GAA) and (CGG) repeats in vivo, associated with FRDA and FXS disorders, respectively (Figure 1, 6). We show that these R-loops interfere with nascent Pol II transcription on FXN gene (Figure 2E). We also demonstrate that R-loops can trigger gene silencing irrespectively of their genomic location (Figure 3). R-loops over expanded repeats are very stable in human cells (Figure 2B), possibly due to failure of their complete turn-over by endogenous enzymes, which may contribute to FRDA pathology. Interestingly, expansion-associated R-loops can be resolved by over-expressed exogenous RNase H1, which leads to transcription up-regulation of FXN expression in vivo (Figure 3).
Previous work has demonstrated that co-transcriptionally formed RNA/DNA hybrids mediate transcription elongation impairment in vitro and in yeast S.cerevisiae [29], [30], suggesting that R-loops may provide roadblocks for RNA polymerases. R-loops over expanded repeats may form a structural block, directly interfering with Pol II transcription elongation. Similar to R-loops at the 3′ends of human genes [16], expansion-associated R-loops could promote RNA Pol II termination, resulting in reduction of active Pol II downstream of the expansion, as detected in this study.
Recently it was suggested that repressive chromatin H3K9me2 modification was not directly responsible for the FXN transcriptional repression [21]. In line with this, reversal of repressive DNA methylation on FMR1 gene was not sufficient to fully restore FMR1 expression [28]. We now show that a decrease in the level of the repressive H3K9me2 chromatin mark does not result in decrease of R-loops on the expanded allele or up-regulation of FXN RNA (Figure 4). These data indicate that R-loop formation is an early event in the process of FXN transcriptional gene silencing, which happens prior to the appearance of heterochromatin marks. We also show that increasing R-loop levels lead to an increase in repressive chromatin marks and subsequent repression of FXN gene expression (Figure 5). Furthermore, we observed that recruitment of G9a methyltransferase is enhanced on expanded FXN allele (Figure 5), providing an interesting possibility that R-loops may directly recruit this enzyme to promote H3K9me2 histone mark deposition. Altogether our results suggest that R-loops act as the primary trigger for repression of expanded FXN and FMR1 alleles which may in turn act to promote heterochromatin formation. Consistent with our data, recently it has been demonstrated that promoter-bound trinucleotide repeat-containing mRNA induces epigenetic silencing in Fragile X syndrome [31]. Indeed R-loops have been implicated in the formation and maintenance of heterochromatin at centromeres in S. pombe [32]. In line with this, H3K9me2 modification is also enriched at the R-loop-containing pause region of β-actin gene (Figure S6), essential for Pol II transcriptional termination [16]. This suggests that R-loops may promote H3K9me2 modification at the 3′end of this gene, similar to expanded repeats of FXN gene. We propose that R-loops may be a common feature of many trinucleotide expansion diseases, contributing to their pathology in vivo (Figure 7). The ability of R-loops to trigger transcriptional silencing in trinucleotide expansion diseases makes them an attractive target for future therapeutic approaches to treat these devastating diseases.
Fig. 7. Model for the role of R-loops in mediating FXN and FMR1 gene silencing. 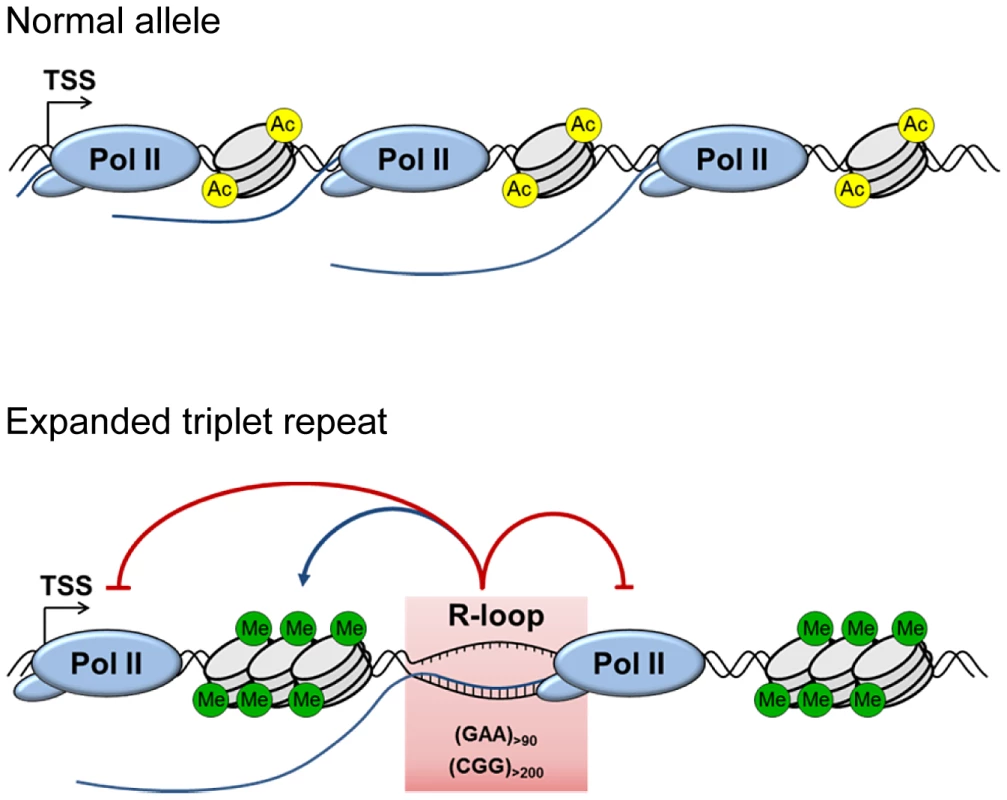
Background R-loop level on wild type allele allows efficient transcriptional elongation and gene expression. Transcribed (GAA)n and (CGG)n expanded repeats form R-loops resulting in decreased initiation and elongation of RNA Pol II. This leads to downregulation of FXN and FMR1 expression, associated with formation of repressive DNA and chromatin marks. In addition to uncovering the molecular mechanisms underlying FRDA and Fragile X pathologies, our work also provides interesting implications for R-loop biology. Taken into consideration this work and the work of others in the field, depending on their genomic location, R-loops may have different functions (reviewed in [11]). Therefore, stable R-loops formed over expanded triplet repeats (this study) may be different from R-loops at the 3′-ends of genes [16], [33] and R-loops formed over CpG island promoters [15], [33]. At promoters, R-loops play a protective role against epigenetic silencing. By contrast, R-loops over FXN expanded repeats correlate with a reduction in transcription elongation and the enrichment of repressive chromatin marks. This suggests that R-loops may be ‘sensed’ differently, depending on their genomic location and sequence composition. Understanding the molecular mechanisms that cells use to distinguish ‘harmful’ from ‘useful’ R-loops is an important biological question in the study of human diseases.
Materials and Methods
Cell culture and drug treatments
Epstein-Barr virus (EBV)-transformed lymphoblastoid cell lines from healthy (GM15851, GM14926, GM06895), FRDA (GM15850, GM16243, GM16209) and FXS (GM03200) patients were obtained from Coriell Institute for Medical Research. Frataxin (GAA) repeat sizes were 650/1030 (GM15850), 800/800 (GM16209) and 670/1170 (GM16243). (CGG) repeat size in FMR1 gene was 530 (GM03200). All experiments were performed with early passages of the cell lines. Lymphoblastoid cell lines were grown in RPMI 1640 medium supplemented with 15% fetal bovine serum (FBS), 100 U/ml penicillin and 100 µg/ml streptomycin at 37°C in 5% CO2. 1 µM 5-azadC (Sigma, A3656) was added to the media for 7 days. 4 µM BIX-01294 (Sigma, B9311) was added to the media for the total of 72 hours. The media was replenished every 24 hours by replacement of half of the conditioned media with fresh media and drug. 10 µM camptothecin (Sigma, C9911) was added for 6 hours. Actinomycin D (Sigma, A9415) was added to a final concentration of 5 µg/ml to the media for 6–48 hours. FXN-Luc and FXN-GAA-Luc HEK293 cells were described in [23] and cultured in DMEM medium supplemented with 10% FBS, 100 U/ml penicillin/streptomycin, 100 µg/ml hygromycin B (Life Technologies). For 5-azadC wash-out experiments, cells were treated with 1 µM 5-azadC for 7 days. On day 7, cell were washed twice with fresh RPMI 1640 medium and cultured in the absence of 5-azadC during the indicated time.
ChIP analysis
ChIP analysis on endogenous genes was carried out as previously described [34], [35]. 5 µg of the following antibodies were used: Pol II antibody (Santa Cruz, H-224), H3 (AbCam, ab1791), H3K9me2 (AbCam, ab1220), G9a (AbCam, ab40542). The immuno-precipitated DNA was used as template for real-time quantitative PCR performed using a Rotor-Gene RG-3000 machine (Corbett Research). The PCR mixture contained QuantiTect SYBR green PCR master mix (Qiagen), 2 µl of the template DNA and corresponding primers from Table S1. Cycling parameters were 95°C for 15 min, followed by 45 cycles of 94°C for 20 s, 58°–62°C for 20 s, and 72°C for 20 s. Fluorescence intensities were plotted against the number of cycles by using an algorithm provided by the manufacturer. Amount of immuno-precipitated protein at a particular gene region was calculated as ‘% of Input’ after subtracting the background signal, as determined by the ‘no antibody’ control.
Expansion PCR in HEK293 cells
20 ng of genomic DNA was used as template in a 25 µl PCR reaction, containing 2.5 U Biotaq DNA polymerase, 3 mM MgCl2, 0.4 mM dNTPs, 0.4 µM GAA104F primer, 0.4 µM GAA629R primer in 1× NH4 reaction buffer. Products were amplified using protocol from [36] with minor modifications. In particular, 5 min at 94°C were followed by 10 cycles of 94°C for 20 s, 65°C for 30 s, 72°C for 5 min. This was followed by 20 cycles of 94°C for 20 s, 65°C for 30 s and 72°C for 5 min, with the 72°C step becoming 20 s longer in each cycle. After a final step at 72°C for 10 min, PCR products were resolved on a 1% agarose gel.
DIP analysis
DNA immuno-precipitation (DIP) analysis on endogenous genes was performed with antibody, recognising RNA/DNA hybrids, purified from S9.6 hybridoma cell lines [37], as described in [16]. In particular, lymphoblastoid and HEK293 cells were split one day before DIP. 10×106 cells were harvested, washed in PBS and incubated in cell lysis buffer (85 mM KCl, 5 mM PIPES pH 8.0, 0.5% NP-40) for 10 min on ice. Nuclei were collected by centrifugation and then incubated in nuclei lysis buffer (50 mM TRIS pH 8.0, 5 mM EDTA, 1% SDS) on ice. Proteins were digested by incubation with proteinase K (Roche) for 3 h at 55°C. Proteins and cell debris were removed by centrifugation after addition of KOAc to the final concentration of 1 M. Genomic DNA containing R-loops was then precipitated by addition of isopropanol. After washing the DNA pellet with 70% EtOH, genomic DNA was resuspended in 400 µl IP dilution buffer (16.7 mM TRIS pH 8.0, 1.2 mM EDTA, 167 mM NaCl, 0.01% SDS, 1.1% Triton X-100) and used for sonication (Diagenode Bioruptor). Bioruptor (Diagenode) settings were 3 min sonication, at the Medium setting 30 sec on/30 sec off interval and the average size of the fragments was ∼500 nt. Sonicated genomic DNA was then pre-cleared with 50 µl protein A agarose beads (Millipore) in 3 ml IP dilution buffer including protease inhibitors (0.5 mM PMSF, 0.8 µg/ml pepstatin A, 1 µg/ml leupeptin) for 1 h at 4°C. 10 µl of S9.6 antibody was added to DNA corresponding to 10×106 cells. Immuno-precipitation was carried out over night at 4°C. Subsequent washes and elution steps are identical to the procedure as described for ChIP.
The immuno-precipitated, non-precipitated, and input DNAs were used as templates for qPCR. DIP RNase H-sensitivity analysis was carried out following the genomic isolation and prior to immuno-precipitation step with the addition of 25 U of RNase H (NEB, M0297S). 100 µl nuclease digestion reaction contained 1× reaction buffer, and it was performed for 6 hours at 37°C. Amount of immuno-precipitated RNA/DNA hybrid at a particular gene region was calculated as ‘% of Input’ after subtracting the background signal, as determined by the ‘no antibody’ control. In the case of FXN gene, all the values are relative to the FXN B amplicon in control cells.
RNA analysis
Total RNA was harvested using TRIZOL reagent (Invitrogen) followed by DNase I treatment (Roche). 1–2 µg of total RNA was reverse-transcribed using SuperScript Reverse Transcriptase III (Invitrogen) with random hexamers (Invitrogen), oligodT primer (for polyA+ RNA) or gene-specific reverse primer (Table S1). The qPCR primers for amplification of polyA+ RNAs were the following: β-actin (ex5F/ex6R), GAPDH (F/R3), γ-actin (γ-actin spliced F/R), FXN (ex3F/ex4R), FMR1 (ex14 F/ex 15R). For analysis of nascent RNA in Figures 2A and 4C FXN primer FXN D was used, while in Figure 3C FXN primer B was used. For quantitative real-time PCR, 2 µl of cDNA was analyzed using a Rotor-Gene RG-3000 real-time PCR machine (Corbett Research) with QuantiTect SYBR green (Qiagen). For analysis of nascent FXN-Luc RNA in HEK293 cells, lucR primer was used for reverse transcription, and in4F and ex5R were used for qPCR.
RNAi and protein analysis
The RNAi was carried out as described [38]. Control siRNA duplex was 5′-UAGCGACUAAACACAUCAA -3′ (Thermo Scientific siGENOME Non-Targeting siRNA #1D-001210-01-20), RNase H1 siRNA duplex was s48357 (Ambion). mRNA target sequence for Topoisomerase I siRNA duplex was 5′-GGACUCCAUCAGAUACUAU -3′. Total protein extracts were harvested using RIPA buffer. 20 and 40 µg of total protein extracts were resolved on SDS-PAGE and detected by Western blotting. Western blots were probed with Topoisomerase I (AbCam, ab109374), actin (Sigma, A2066), RNase H1 (AbCam, ab83179) antibodies.
RNase H1 over-expression
FXN-Luc and FXN-GAA-Luc cells were freshly split into 10 cm dishes and transfected on the following day with 10 µg Flag or RNase H1-Flag plasmids using TransFectin reagent (BioRad), following the manufacturer's instructions. Cells were harvested 48 hours after transfection. RNaseH1-Flag plasmid was cloned by replacing the GFP tag in the RNaseH1-GFP plasmid, provided by Prof.R.J. Crouch, with the Flag tag using RNaseH1-FLAG(F) and RNaseH1-FLAG(R) primers.
Br-UTP nuclear run-on analysis
The Br-UTP NRO analysis was carried as described in [16]. The equivalent of 8×106 nuclei from lymphoblastoid cells were used for each Br-UTP NRO reaction.
Statistical analysis
Unless otherwise stated, the figures present the average values of at least three independent experiments +/ − SEM. Asterisks (*) indicate statistical significance (* p<0.05; ** p<0.01; *** p<0.001), based on unpaired, two-tailed distribution Student's t test.
Supporting Information
Zdroje
1. Lopez CastelA, ClearyJD, PearsonCE (2010) Repeat instability as the basis for human diseases and as a potential target for therapy. Nat Rev Mol Cell Biol 11 : 165–170.
2. CampuzanoV, MonterminiL, MoltoMD, PianeseL, CosseeM, et al. (1996) Friedreich's ataxia: autosomal recessive disease caused by an intronic GAA triplet repeat expansion. Science 271 : 1423–1427.
3. MarmolinoD (2011) Friedreich's ataxia: past, present and future. Brain Res Rev 67 : 311–330.
4. KumariD, BiacsiRE, UsdinK (2011) Repeat expansion affects both transcription initiation and elongation in friedreich ataxia cells. J Biol Chem 286 : 4209–4215.
5. HermanD, JenssenK, BurnettR, SoragniE, PerlmanSL, et al. (2006) Histone deacetylase inhibitors reverse gene silencing in Friedreich's ataxia. Nat Chem Biol 2 : 551–558.
6. KimE, NapieralaM, DentSY (2011) Hyperexpansion of GAA repeats affects post-initiation steps of FXN transcription in Friedreich's ataxia. Nucleic Acids Res 39 : 8366–8377.
7. GreeneE, MahishiL, EntezamA, KumariD, UsdinK (2007) Repeat-induced epigenetic changes in intron 1 of the frataxin gene and its consequences in Friedreich ataxia. Nucleic Acids Res 35 : 3383–3390.
8. KumariD, UsdinK (2012) Is Friedreich ataxia an epigenetic disorder? Clin Epigenetics 4 : 2.
9. ChanPK, TorresR, YandimC, LawPP, KhadayateS, et al. (2013) Heterochromatinization induced by GAA-repeat hyperexpansion in Friedreich's ataxia can be reduced upon HDAC inhibition by vitamin B3. Hum Mol Genet 22 : 2662–2675.
10. WellsRD (2008) DNA triplexes and Friedreich ataxia. FASEB J 22 : 1625–1634.
11. AguileraA, Garcia-MuseT (2012) R loops: from transcription byproducts to threats to genome stability. Mol Cell 46 : 115–124.
12. HuangFT, YuK, HsiehCL, LieberMR (2006) Downstream boundary of chromosomal R-loops at murine switch regions: implications for the mechanism of class switch recombination. Proc Natl Acad Sci U S A 103 : 5030–5035.
13. YuK, ChedinF, HsiehCL, WilsonTE, LieberMR (2003) R-loops at immunoglobulin class switch regions in the chromosomes of stimulated B cells. Nat Immunol 4 : 442–451.
14. LiX, ManleyJL (2005) Inactivation of the SR protein splicing factor ASF/SF2 results in genomic instability. Cell 122 : 365–378.
15. GinnoPA, LottPL, ChristensenHC, KorfI, ChedinF (2012) R-loop formation is a distinctive characteristic of unmethylated human CpG island promoters. Mol Cell 45 : 814–825.
16. Skourti-StathakiK, ProudfootNJ, GromakN (2011) Human senataxin resolves RNA/DNA hybrids formed at transcriptional pause sites to promote Xrn2-dependent termination. Mol Cell 42 : 794–805.
17. GrabczykE, MancusoM, SammarcoMC (2007) A persistent RNA.DNA hybrid formed by transcription of the Friedreich ataxia triplet repeat in live bacteria, and by T7 RNAP in vitro. Nucleic Acids Res 35 : 5351–5359.
18. ReddyK, TamM, BowaterRP, BarberM, TomlinsonM, et al. (2011) Determinants of R-loop formation at convergent bidirectionally transcribed trinucleotide repeats. Nucleic Acids Res 39 : 1749–1762.
19. LinY, DentSY, WilsonJH, WellsRD, NapieralaM (2010) R loops stimulate genetic instability of CTG.CAG repeats. Proc Natl Acad Sci U S A 107 : 692–697.
20. LinY, WilsonJH (2012) Nucleotide excision repair, mismatch repair, and R-loops modulate convergent transcription-induced cell death and repeat instability. PLoS One 7: e46807.
21. PungaT, BuhlerM (2010) Long intronic GAA repeats causing Friedreich ataxia impede transcription elongation. EMBO Mol Med 2 : 120–129.
22. PhillipsDD, GarbocziDN, SinghK, HuZ, LepplaSH, et al. (2013) The sub-nanomolar binding of DNA-RNA hybrids by the single-chain Fv fragment of antibody S9.6. J Mol Recognit 26 : 376–381.
23. LufinoMM, SilvaAM, NemethAH, Alegre-AbarrateguiJ, RussellAJ, et al. (2013) A GAA repeat expansion reporter model of Friedreich's ataxia recapitulates the genomic context and allows rapid screening of therapeutic compounds. Hum Mol Genet 22 : 5173–5187.
24. CerritelliSM, CrouchRJ (2009) Ribonuclease H: the enzymes in eukaryotes. FEBS J 276 : 1494–1505.
25. El HageA, FrenchSL, BeyerAL, TollerveyD (2010) Loss of Topoisomerase I leads to R-loop-mediated transcriptional blocks during ribosomal RNA synthesis. Genes Dev 24 : 1546–1558.
26. SordetO, RedonCE, Guirouilh-BarbatJ, SmithS, SolierS, et al. (2009) Ataxia telangiectasia mutated activation by transcription - and topoisomerase I-induced DNA double-strand breaks. EMBO Rep 10 : 887–893.
27. SantoroMR, BraySM, WarrenST (2011) Molecular mechanisms of fragile X syndrome: a twenty-year perspective. Annu Rev Pathol 7 : 219–245.
28. CoffeeB, ZhangF, CemanS, WarrenST, ReinesD (2002) Histone modifications depict an aberrantly heterochromatinized FMR1 gene in fragile x syndrome. Am J Hum Genet 71 : 923–932.
29. HuertasP, AguileraA (2003) Cotranscriptionally formed DNA:RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol Cell 12 : 711–721.
30. TousC, AguileraA (2007) Impairment of transcription elongation by R-loops in vitro. Biochem Biophys Res Commun 360 : 428–432.
31. ColakD, ZaninovicN, CohenMS, RosenwaksZ, YangWY, et al. (2014) Promoter-bound trinucleotide repeat mRNA drives epigenetic silencing in fragile X syndrome. Science 343 : 1002–1005.
32. NakamaM, KawakamiK, KajitaniT, UranoT, MurakamiY (2012) DNA-RNA hybrid formation mediates RNAi-directed heterochromatin formation. Genes Cells 17 : 218–233.
33. GinnoPA, LimYW, LottPL, KorfI, ChedinF (2013) GC skew at the 5′ and 3′ ends of human genes links R-loop formation to epigenetic regulation and transcription termination. Genome Res 23 : 1590–1600.
34. KadenerS, FededaJP, RosbashM, KornblihttAR (2002) Regulation of alternative splicing by a transcriptional enhancer through RNA pol II elongation. Proc Natl Acad Sci U S A 99 : 8185–8190.
35. WestS, GromakN, ProudfootNJ (2004) Human 5′→3′ exonuclease Xrn2 promotes transcription termination at co-transcriptional cleavage sites. Nature 432 : 522–525.
36. HollowayTP, RowleySM, DelatyckiMB, SarseroJP (2011) Detection of interruptions in the GAA trinucleotide repeat expansion in the FXN gene of Friedreich ataxia. Biotechniques 50 : 182–186.
37. BoguslawskiSJ, SmithDE, MichalakMA, MickelsonKE, YehleCO, et al. (1986) Characterization of monoclonal antibody to DNA.RNA and its application to immunodetection of hybrids. J Immunol Methods 89 : 123–130.
38. WollertonMC, GoodingC, WagnerEJ, Garcia-BlancoMA, SmithCW (2004) Autoregulation of polypyrimidine tract binding protein by alternative splicing leading to nonsense-mediated decay. Mol Cell 13 : 91–100.
Štítky
Genetika Reprodukční medicína
Článek Ribosomal Protein Mutations Induce Autophagy through S6 Kinase Inhibition of the Insulin PathwayČlánek Recent Mitochondrial DNA Mutations Increase the Risk of Developing Common Late-Onset Human DiseasesČlánek G×G×E for Lifespan in : Mitochondrial, Nuclear, and Dietary Interactions that Modify LongevityČlánek PINK1-Parkin Pathway Activity Is Regulated by Degradation of PINK1 in the Mitochondrial MatrixČlánek Rapid Evolution of PARP Genes Suggests a Broad Role for ADP-Ribosylation in Host-Virus ConflictsČlánek The Impact of Population Demography and Selection on the Genetic Architecture of Complex TraitsČlánek Lifespan Extension by Methionine Restriction Requires Autophagy-Dependent Vacuolar AcidificationČlánek The Case for Junk DNA
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2014 Číslo 5- Růst a vývoj dětí narozených pomocí IVF
- Délka menstruačního cyklu jako marker ženské plodnosti
- Vztah užívání alkoholu a mužské fertility
- Šanci na úspěšný průběh těhotenství snižují nevhodné hladiny progesteronu vznikající při umělém oplodnění
- Vliv melatoninu a cirkadiálního rytmu na ženskou reprodukci
-
Všechny články tohoto čísla
- Genetic Interactions Involving Five or More Genes Contribute to a Complex Trait in Yeast
- A Mutation in the Gene in Dogs with Hereditary Footpad Hyperkeratosis (HFH)
- Loss of Function Mutation in the Palmitoyl-Transferase HHAT Leads to Syndromic 46,XY Disorder of Sex Development by Impeding Hedgehog Protein Palmitoylation and Signaling
- Heterogeneity in the Frequency and Characteristics of Homologous Recombination in Pneumococcal Evolution
- Genome-Wide Nucleosome Positioning Is Orchestrated by Genomic Regions Associated with DNase I Hypersensitivity in Rice
- Null Mutation in PGAP1 Impairing Gpi-Anchor Maturation in Patients with Intellectual Disability and Encephalopathy
- Single Nucleotide Variants in Transcription Factors Associate More Tightly with Phenotype than with Gene Expression
- Ribosomal Protein Mutations Induce Autophagy through S6 Kinase Inhibition of the Insulin Pathway
- Recent Mitochondrial DNA Mutations Increase the Risk of Developing Common Late-Onset Human Diseases
- Epistatically Interacting Substitutions Are Enriched during Adaptive Protein Evolution
- Meiotic Drive Impacts Expression and Evolution of X-Linked Genes in Stalk-Eyed Flies
- G×G×E for Lifespan in : Mitochondrial, Nuclear, and Dietary Interactions that Modify Longevity
- Population Genomic Analysis of Ancient and Modern Genomes Yields New Insights into the Genetic Ancestry of the Tyrolean Iceman and the Genetic Structure of Europe
- p53 Requires the Stress Sensor USF1 to Direct Appropriate Cell Fate Decision
- Whole Exome Re-Sequencing Implicates and Cilia Structure and Function in Resistance to Smoking Related Airflow Obstruction
- Allelic Expression of Deleterious Protein-Coding Variants across Human Tissues
- R-loops Associated with Triplet Repeat Expansions Promote Gene Silencing in Friedreich Ataxia and Fragile X Syndrome
- PINK1-Parkin Pathway Activity Is Regulated by Degradation of PINK1 in the Mitochondrial Matrix
- The Impairment of MAGMAS Function in Human Is Responsible for a Severe Skeletal Dysplasia
- Octopamine Neuromodulation Regulates Gr32a-Linked Aggression and Courtship Pathways in Males
- Mlh2 Is an Accessory Factor for DNA Mismatch Repair in
- Activating Transcription Factor 6 Is Necessary and Sufficient for Alcoholic Fatty Liver Disease in Zebrafish
- The Spatiotemporal Program of DNA Replication Is Associated with Specific Combinations of Chromatin Marks in Human Cells
- Rapid Evolution of PARP Genes Suggests a Broad Role for ADP-Ribosylation in Host-Virus Conflicts
- Genome-Wide Inference of Ancestral Recombination Graphs
- Mutations in Four Glycosyl Hydrolases Reveal a Highly Coordinated Pathway for Rhodopsin Biosynthesis and N-Glycan Trimming in
- SHP2 Regulates Chondrocyte Terminal Differentiation, Growth Plate Architecture and Skeletal Cell Fates
- The Impact of Population Demography and Selection on the Genetic Architecture of Complex Traits
- Retinoid-X-Receptors (α/β) in Melanocytes Modulate Innate Immune Responses and Differentially Regulate Cell Survival following UV Irradiation
- Genetic Dissection of the Female Head Transcriptome Reveals Widespread Allelic Heterogeneity
- Genome Sequencing and Comparative Genomics of the Broad Host-Range Pathogen AG8
- Copy Number Variation Is a Fundamental Aspect of the Placental Genome
- GOLPH3 Is Essential for Contractile Ring Formation and Rab11 Localization to the Cleavage Site during Cytokinesis in
- Hox Transcription Factors Access the RNA Polymerase II Machinery through Direct Homeodomain Binding to a Conserved Motif of Mediator Subunit Med19
- Drosha Promotes Splicing of a Pre-microRNA-like Alternative Exon
- Predicting the Minimal Translation Apparatus: Lessons from the Reductive Evolution of
- PAX6 Regulates Melanogenesis in the Retinal Pigmented Epithelium through Feed-Forward Regulatory Interactions with MITF
- Enhanced Interaction between Pseudokinase and Kinase Domains in Gcn2 stimulates eIF2α Phosphorylation in Starved Cells
- A HECT Ubiquitin-Protein Ligase as a Novel Candidate Gene for Altered Quinine and Quinidine Responses in
- dGTP Starvation in Provides New Insights into the Thymineless-Death Phenomenon
- Phosphorylation Modulates Clearance of Alpha-Synuclein Inclusions in a Yeast Model of Parkinson's Disease
- RPM-1 Uses Both Ubiquitin Ligase and Phosphatase-Based Mechanisms to Regulate DLK-1 during Neuronal Development
- More of a Good Thing or Less of a Bad Thing: Gene Copy Number Variation in Polyploid Cells of the Placenta
- More of a Good Thing or Less of a Bad Thing: Gene Copy Number Variation in Polyploid Cells of the Placenta
- Heritable Transmission of Stress Resistance by High Dietary Glucose in
- Revertant Mutation Releases Confined Lethal Mutation, Opening Pandora's Box: A Novel Genetic Pathogenesis
- Lifespan Extension by Methionine Restriction Requires Autophagy-Dependent Vacuolar Acidification
- A Genome-Wide Assessment of the Role of Untagged Copy Number Variants in Type 1 Diabetes
- Selectivity in Genetic Association with Sub-classified Migraine in Women
- A Lack of Parasitic Reduction in the Obligate Parasitic Green Alga
- The Proper Splicing of RNAi Factors Is Critical for Pericentric Heterochromatin Assembly in Fission Yeast
- Discovery and Functional Annotation of SIX6 Variants in Primary Open-Angle Glaucoma
- Six Homeoproteins and a linc-RNA at the Fast MYH Locus Lock Fast Myofiber Terminal Phenotype
- EDR1 Physically Interacts with MKK4/MKK5 and Negatively Regulates a MAP Kinase Cascade to Modulate Plant Innate Immunity
- Genes That Bias Mendelian Segregation
- The Case for Junk DNA
- An In Vivo EGF Receptor Localization Screen in Identifies the Ezrin Homolog ERM-1 as a Temporal Regulator of Signaling
- Mosaic Epigenetic Dysregulation of Ectodermal Cells in Autism Spectrum Disorder
- Hyperactivated Wnt Signaling Induces Synthetic Lethal Interaction with Rb Inactivation by Elevating TORC1 Activities
- Mutations in the Cholesterol Transporter Gene Are Associated with Excessive Hair Overgrowth
- Scribble Modulates the MAPK/Fra1 Pathway to Disrupt Luminal and Ductal Integrity and Suppress Tumour Formation in the Mammary Gland
- A Novel CH Transcription Factor that Regulates Expression Interdependently with GliZ in
- Phosphorylation of a WRKY Transcription Factor by MAPKs Is Required for Pollen Development and Function in
- Bayesian Test for Colocalisation between Pairs of Genetic Association Studies Using Summary Statistics
- Spermatid Cyst Polarization in Depends upon and the CPEB Family Translational Regulator
- Insights into the Genetic Structure and Diversity of 38 South Asian Indians from Deep Whole-Genome Sequencing
- Intron Retention in the 5′UTR of the Novel ZIF2 Transporter Enhances Translation to Promote Zinc Tolerance in
- A Dominant-Negative Mutation of Mouse Causes Glaucoma and Is Semi-lethal via LBD1-Mediated Dimerisation
- Biased, Non-equivalent Gene-Proximal and -Distal Binding Motifs of Orphan Nuclear Receptor TR4 in Primary Human Erythroid Cells
- Ras-Mediated Deregulation of the Circadian Clock in Cancer
- Retinoic Acid-Related Orphan Receptor γ (RORγ): A Novel Participant in the Diurnal Regulation of Hepatic Gluconeogenesis and Insulin Sensitivity
- Extensive Diversity of Prion Strains Is Defined by Differential Chaperone Interactions and Distinct Amyloidogenic Regions
- Fine Tuning of the UPR by the Ubiquitin Ligases Siah1/2
- Paternal Poly (ADP-ribose) Metabolism Modulates Retention of Inheritable Sperm Histones and Early Embryonic Gene Expression
- Allele-Specific Genome-wide Profiling in Human Primary Erythroblasts Reveal Replication Program Organization
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- PINK1-Parkin Pathway Activity Is Regulated by Degradation of PINK1 in the Mitochondrial Matrix
- Null Mutation in PGAP1 Impairing Gpi-Anchor Maturation in Patients with Intellectual Disability and Encephalopathy
- Phosphorylation of a WRKY Transcription Factor by MAPKs Is Required for Pollen Development and Function in
- p53 Requires the Stress Sensor USF1 to Direct Appropriate Cell Fate Decision
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MUDr. Irena Krčmová, CSc.
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



