Interplay between Synaptonemal Complex, Homologous Recombination, and Centromeres during Mammalian Meiosis
The intimate synapsis of homologous chromosome pairs (homologs) by synaptonemal complexes (SCs) is an essential feature of meiosis. In many organisms, synapsis and homologous recombination are interdependent: recombination promotes SC formation and SCs are required for crossing-over. Moreover, several studies indicate that initiation of SC assembly occurs at sites where crossovers will subsequently form. However, recent analyses in budding yeast and fruit fly imply a special role for centromeres in the initiation of SC formation. In addition, in budding yeast, persistent SC–dependent centromere-association facilitates the disjunction of chromosomes that have failed to become connected by crossovers. Here, we examine the interplay between SCs, recombination, and centromeres in a mammal. In mouse spermatocytes, centromeres do not serve as SC initiation sites and are invariably the last regions to synapse. However, centromeres are refractory to de-synapsis during diplonema and remain associated by short SC fragments. Since SC–dependent centromere association is lost before diakinesis, a direct role in homolog segregation seems unlikely. However, post–SC disassembly, we find evidence of inter-centromeric connections that could play a more direct role in promoting homolog biorientation and disjunction. A second class of persistent SC fragments is shown to be crossover-dependent. Super-resolution structured-illumination microscopy (SIM) reveals that these structures initially connect separate homolog axes and progressively diminish as chiasmata form. Thus, DNA crossing-over (which occurs during pachynema) and axis remodeling appear to be temporally distinct aspects of chiasma formation. SIM analysis of the synapsis and crossover-defective mutant Sycp1−/− implies that SCs prevent unregulated fusion of homolog axes. We propose that SC fragments retained during diplonema stabilize nascent bivalents and help orchestrate local chromosome reorganization that promotes centromere and chiasma function.
Published in the journal:
. PLoS Genet 8(6): e32767. doi:10.1371/journal.pgen.1002790
Category:
Research Article
doi:
https://doi.org/10.1371/journal.pgen.1002790
Summary
The intimate synapsis of homologous chromosome pairs (homologs) by synaptonemal complexes (SCs) is an essential feature of meiosis. In many organisms, synapsis and homologous recombination are interdependent: recombination promotes SC formation and SCs are required for crossing-over. Moreover, several studies indicate that initiation of SC assembly occurs at sites where crossovers will subsequently form. However, recent analyses in budding yeast and fruit fly imply a special role for centromeres in the initiation of SC formation. In addition, in budding yeast, persistent SC–dependent centromere-association facilitates the disjunction of chromosomes that have failed to become connected by crossovers. Here, we examine the interplay between SCs, recombination, and centromeres in a mammal. In mouse spermatocytes, centromeres do not serve as SC initiation sites and are invariably the last regions to synapse. However, centromeres are refractory to de-synapsis during diplonema and remain associated by short SC fragments. Since SC–dependent centromere association is lost before diakinesis, a direct role in homolog segregation seems unlikely. However, post–SC disassembly, we find evidence of inter-centromeric connections that could play a more direct role in promoting homolog biorientation and disjunction. A second class of persistent SC fragments is shown to be crossover-dependent. Super-resolution structured-illumination microscopy (SIM) reveals that these structures initially connect separate homolog axes and progressively diminish as chiasmata form. Thus, DNA crossing-over (which occurs during pachynema) and axis remodeling appear to be temporally distinct aspects of chiasma formation. SIM analysis of the synapsis and crossover-defective mutant Sycp1−/− implies that SCs prevent unregulated fusion of homolog axes. We propose that SC fragments retained during diplonema stabilize nascent bivalents and help orchestrate local chromosome reorganization that promotes centromere and chiasma function.
Introduction
The formation of gametes typically involves halving of the cellular chromosome complement from diploid to haploid. This is achieved via two consecutive rounds of chromosome segregation during the process of meiosis [1]. Prior to the first meiotic division, replicated chromosomes associate into homologous pairs and become connected along their lengths by synaptonemal complexes (SCs) [2], [3]. SCs are proteinaceous structures with a zipper-like morphology [4]–[8]. The tripartite SC structure comprises two lateral elements, inferred to be elaborations of cohesin-based homolog axes, and a central element consisting of transverse filaments that interconnect the two lateral elements [7]–[10]. SC components show tendencies for self-assembly into ordered arrays and SC formation is believed to occur via polymerization from specific nucleation sites where the homolog axes have been brought into close proximity [6], [11].
In many organisms, including plants, fungi and mammals, the template-dependent DNA-repair process called homologous recombination is coopted during meiosis to facilitate homolog pairing and synapsis [12]. In these cases, SC formation often nucleates at points where recombination brings the homolog axes together [11]. However, organisms such as Drosophila and C. elegans do not require recombination for homolog pairing and SC formation and instead have evolved dedicated chromosome pairing sites [3], [13].
In addition to promoting chromosome pairing and synapsis, recombination plays a critical function in directing the disjunction of homologs at the first meiotic division. Specifically, crossover recombination in conjunction with sister-chromatid cohesion results in structures called chiasmata that tether homolog pairs and thereby facilitate their stable biorientation on the spindle [14]–[16]. The interdependence of recombination and SCs is further highlighted by the fact that synapsis promotes crossing-over, at least in part by recruiting crossover-specific recombination factors [17], [18]. Furthermore, studies in a number of organisms imply a functional relationship between SC nucleation sites and crossovers (reviewed in [11]). Specifically, SC formation often initiates at sites where crossovers will subsequently form.
Recent studies in Saccharomyces cerevisiae and Drosophila suggest that centromeres play special roles in meiotic chromosome pairing and the initiation of SC formation [19], [20]. During early prophase in S. cerevisiae, centromeres undergo homology-independent “coupling”, which depends on the SC central element component, Zip1 [21], [22]. Centromere coupling is proposed to be a driving force for two-by-two chromosome association that facilitates recombination-dependent homolog pairing [22], [23]. However, analysis of recombination patterns in the zip1 mutant support an alternative proposal that coupling helps to suppress centromere-proximal crossing-over, which is associated with chromosome nondisjunction [24] [25]–[27]. Following initial coupling, centromeres appear to act as nucleation sites for SC polymerization, although it is clear that recombination sites within the chromosome arms are also utilized [11], [28]. Consistent with a role for centromeres in nucleating SC formation, in a mutant situation where SCs are assembled independently of recombination, centromeres appear to be the exclusive sites of initiation [29]. This same study indicated that Zip3, a putative E3-ligase for conjugation of the small protein modifier SUMO, negatively regulates SC initiation between centromeres. A role for centromeres in SC formation is further supported by recent studies in Drosophila females, which showed that centromeres undergo SC-dependent clustering and function as SC initiation sites [19], [30], [31].
Although crossing-over is a highly efficient process, achiasmate homologs do occasionally arise. In budding yeast, Zip1-mediated centromere coupling plays an additional late role to facilitate the disjunction of achiasmate chromosomes [21], [32]. In conjunction with the spindle assembly checkpoint, this process promotes the accurate disjunction of a single pair of achiasmate chromosomes in about 90% of meioses (random segregation predicts disjunction in only 50% of cells) [32], [33]. Efficient achiasmate segregation is also observed in Drosophila, although a role for SC components has not been demonstrated [34].
In some mammals, SC components are also inferred to promote the disjunction of achiasmate chromosomes, specifically the sex chromosomes [35]–[37]. Typically, mammalian X and Y chromosomes synapse at short regions of homology, termed pseudoautosomal regions, where crossing-over occurs to form X-Y chiasmata [38]–[40]. However, in the Elegant Fat-tailed Mouse Opossum (a marsupial) and the Mongolian gerbil (a eutherian mammal), X-Y chiasmata are not formed, but persistent structures composed of SC proteins appear to tether the X and Y to facilitate their disjunction during anaphase [35]–[37]. Finally, in some insects, SCs are retained until anaphase I and appear to completely supersede the function of chiasmata in directing disjunction [6].
In this study we analyze the interplay between SCs, recombination and centromeres in the mouse. Immunocytological analysis of prophase spermatocytes from wild-type and mutant mice indicates that centromeres do not undergo early-stage coupling and SC assembly never initiates from centromeres. However, centromeres remain associated throughout much of the diplotene stage, connected by short SC fragments. While this general, SC-dependent centromere association appears to be lost prior to diakinesis, we detect a distinct class of inter-centromeric bridges at this stage. These structures could play a more direct role in biorienting homologs on the spindle and raise the possibility of an achiasmate segregation system in mouse.
A second distinct class of retained SC fragment is also observed during diplotene and shown to be crossover-dependent. Structured illumination microscopy reveals that these structures mark sites of developing chiasmata and are lost as the homolog axes fuse. Analysis of the SC-defective Sycp1 mutant suggests a novel role for the SC central element in preventing inappropriate interactions between homolog axes. We discuss the idea that SC fragments retained during diplonema function to locally stabilize homolog associations and coordinate important morphological and compositional changes in preparation for chromosome segregation.
Results
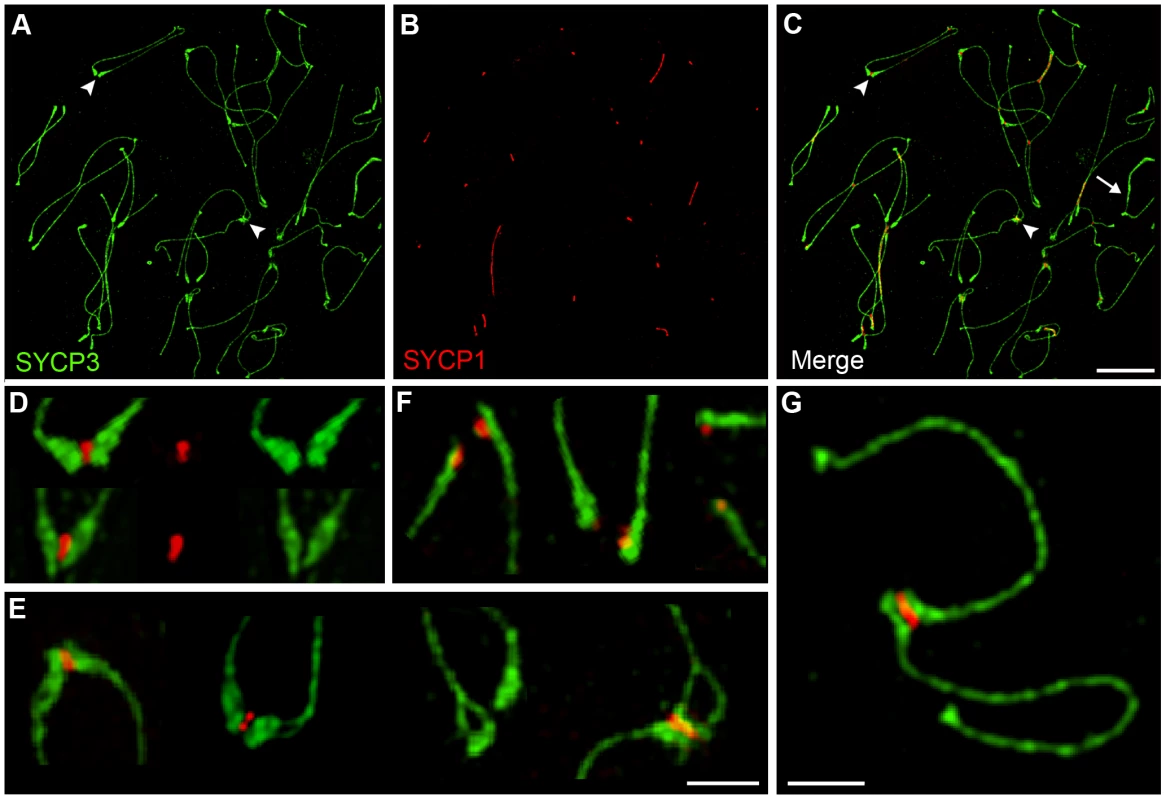
The formation and disassembly of SCs, and the pairing status of centromeres were assessed throughout meiotic prophase using immunofluorescence staining of surface-spread chromosomes from mouse spermatocytes (Figure 1; Materials and Methods). Centromeres were detected using CREST antiserum, which recognizes the constitutively centromere-associated proteins, CENP-A, -B, and -C [41]. We stringently defined “associated” centromeres as pairs of CREST foci that were ≤0.6 µm apart (regardless of whether or not they were associated with SC; Figure 1A). A cutoff of 0.6 µm was chosen because this was the maximum distance measured between synapsed CREST foci in pachytene nuclei. We also quantified frequencies of “synapsed” centromeres, which were defined as associated CREST signals that also colocalized with SC central-element protein, SYCP1 (the mammalian ortholog of budding yeast Zip1 [42]; Figure 1A).
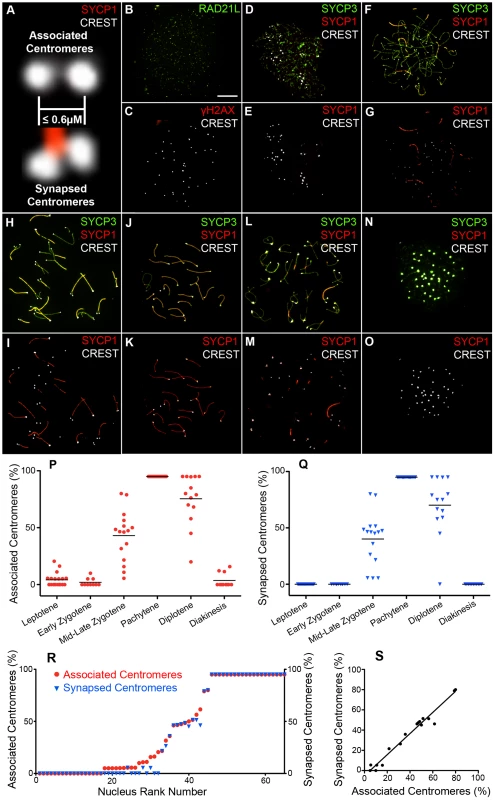
To identify nuclei in very early prophase (pre-leptonema), slides were co-stained for the cohesin marker, RAD21L (a meiosis-specific kleisin [43]–[45]) and the DSB marker, γH2AX (a histone H2A variant that is rapidly phosphorylated at sites of DSB formation [46]). Pre-leptonema nuclei were defined as being RAD21L positive and γH2AX negative (Figure 1B and 1C). Other prophase stages were defined using standard cytological criteria by immunostaining for the homolog axis component, SYCP3, and SC central element protein, SYCP1. During leptonema, short stretches of SYCP3 staining mark the developing homolog axes (Figure 1D and 1E). SYCP3 axes elaborate into contiguous structures throughout zygonema and homologs progressively synapse (Figure 1F–1I). Cells reach pachynema when all the autosomes are fully synapsed, as shown by contiguous SYCP1 staining (Figure 1J and 1K). Progressive desynapsis of homologs occurs during diplonema to reveal homolog axes connected by nascent chiasmata (Figure 1L and 1M). Subsequently SYCP3 axes breakdown and cells enter diakinesis (Figure 1N and 1O). Diplotene and diakinesis stages are also marked by a pronounced enrichment of SYCP3 staining at the centromeric ends of the telocentric mouse chromosomes [10], [47].
Centromere Association during Meiotic Prophase
Figure 1 shows representative nuclei from each stage of meiotic prophase. Quantification of centromere association and centromere synapsis throughout these stages is presented in panels 1P and 1Q, respectively. This analysis reveals several key features of prophase centromere behavior in mouse. Analogous observations are made by Bisig et al. in the accompanying study [48].
Centromeres are not coupled prior to zygonema
In pre-leptotene and leptotene stage nuclei, low levels of centromere association are observed (<21%), but centromeres are never associated with SYCP1 (Figure 1B–1E). In fact, unlike budding yeast Zip1, no chromosome-associated SYCP1 staining is detected in spread nuclei at these stages. Thus, a metastable pre-DSB centromere-association process, analogous to the Zip1-dependent centromere-coupling described in budding yeast, does not occur in mouse.
Synapsis does not initiate at centromeres
During early zygonema, initial stretches of SC detected by SYCP1 staining are not associated with centromeres (0 out of 92 SYCP1 stretches; 10 nuclei analyzed), which is reflected in the very low levels of centromere association and the absence of synapsed centromeres in these nuclei (Figure 1F, 1G and 1Q), i.e. SC formation does not initiate at centromere regions. This again contrasts budding yeast in which synapsis frequently initiates at or close to centromeres [28].
Centromeres are the last regions to synapse
Analysis of late-zygonema nuclei reveals that centromeres are generally the last chromosomal regions to synapse (Figure 1H and 1I). The fact that mouse chromosomes are telocentric raises the possibility that late synapsis of centromeres is a consequence of their sub-terminal location. However, we observed that the non-centromeric ends of homologs nearly always synapsed before the centromeric ends did. In 89.2% of homologs with synapsis at only one end, the synapsed end was the non-centromeric end (107/120 chromosomes from 16 nuclei). Therefore, late synapsis appears to be an inherent feature of centromeres and is not a consequence of their terminal location in mouse. Consistent with this inference, late synapsis of centromeres has been noted in a number of organisms with metacentric chromosomes [11], [49]–[52], including humans [53].
Centromere association correlates with centromere synapsis
During zygonema, levels of centromere association and centromere synapsis are closely correlated (Figure 1P, 1Q, 1R and 1S). Overall, 90.4% of associated centromere pairs detected in zygotene nuclei are also synapsed (123/136, 19 nuclei). However, low levels of associated but not synapsed centromeres are detected. These ostensibly synapsis-independent centromere associations could be bona fide interactions, but may also result from a low level of fortuitous colocalizations, or be driven by closely adjacent synapsis that hasn't yet converged on the CREST signals. Overall, these data imply that the close juxtaposition of homologous centromeres during mouse meiosis is driven primarily by advancing synapsis that initiated at a distal site.
The centromeres of sex chromosomes do not pair in pachynema
Stable pairing and synapsis of mouse sex chromosomes occurs between the pseudoautosomal regions, short (≤1 Mb) stretches of homology located at the non-centromeric ends of the X and Y [54]. Given the observations above, that SC formation does not initiate at centromeres and centromere association appears to be dependent on synapsis (that must have initiated from an adjacent site), we predicted that the sex chromosome centromeres should never be paired. Indeed, this is the case: in 20 pachynema nuclei, in which PAR synapsis was observed, the centromeres of the X and Y chromosomes were never associated.
Centromeres and nascent chiasmata are the last sites to desynapse
As previously described [6], [49], [50], [52], centromeres and chiasma sites behave distinctly from other chromosomal regions as homologs desynapse during diplonema (Figure 1L and 1M). First, centromeres are typically some of the last regions to separate, with high levels of association remaining until the end of diplonema (Figure 1P, 1Q). Second, the last remnants of SYCP1 staining are invariably localized with associated centromeres as well as with nascent chiasmata (discussed below). Notably, even for bivalents in which desynapsis appears to have initiated adjacent to the centromeres, a focus of SYCP1 often remains colocalized with the paired CREST signals. Thus, centromeres and crossover sites are relatively refractory to desynapsis.
Centromere association is lost by late diplonema
As the level of homolog desynapsis reaches ≥80%, centromere association is lost. By diakinesis, as axial SYCP3 structures diminish and centromere-associated SYCP3 further accumulates, homolog centromeres are infrequently associated and SYCP1 is undetectable (Figure 1N, 1O, 1P and 1Q).
Synapsis Does Not Initiate at Centromeres in the Absence of Recombination or the Mammalian Zip3 Ortholog, Rnf212
In budding yeast spo11 mutants, which fail to initiate recombination, persistent Zip1-mediated centromere coupling is observed [22]. Moreover, in the absence of recombination, centromeres become the primary sites of SC initiation in budding yeast [29]. Therefore, we analyzed the relationship between centromeres and SC in Spo11−/− knock-out mice (Figure S1). Neither early centromere-associations nor preferential initiation of SC formation from centromeres were observed in Spo11−/− spermatocytes.
Budding yeast Zip3 is a RING-domain protein and putative E3-ligase for the ubiquitin-like modifier, SUMO [55], [56] (also see [57]). While yeast zip3 mutants show a general synapsis defect, the vast majority of SCs that do form are initiated between centromeres, leading to the proposal that Zip3 specifically inhibits SC initiation between centromeres [28], [29]. We recently constructed a knock-out mutation of the mouse Zip3 homolog, Rnf212 (A.R., H.Q., J.K.C. and N.H., unpublished data), allowing us to test the idea that RNF212 has an analogous inhibitory function in mammals [58]. Analysis of initial SC stretches in zygotene-stage Rnf212−/− spermatocytes shows that, as in wild type, SC formation does not initiate between centromeres (Figure S2). Thus, absence of the Zip3 homolog, RNF212, does not permit SC to initiate between centromeres in mammals.
SC Remnants during Diplonema Contain SC Central Element Components, SYCE1, SYCE2, SYCP1, and TEX12
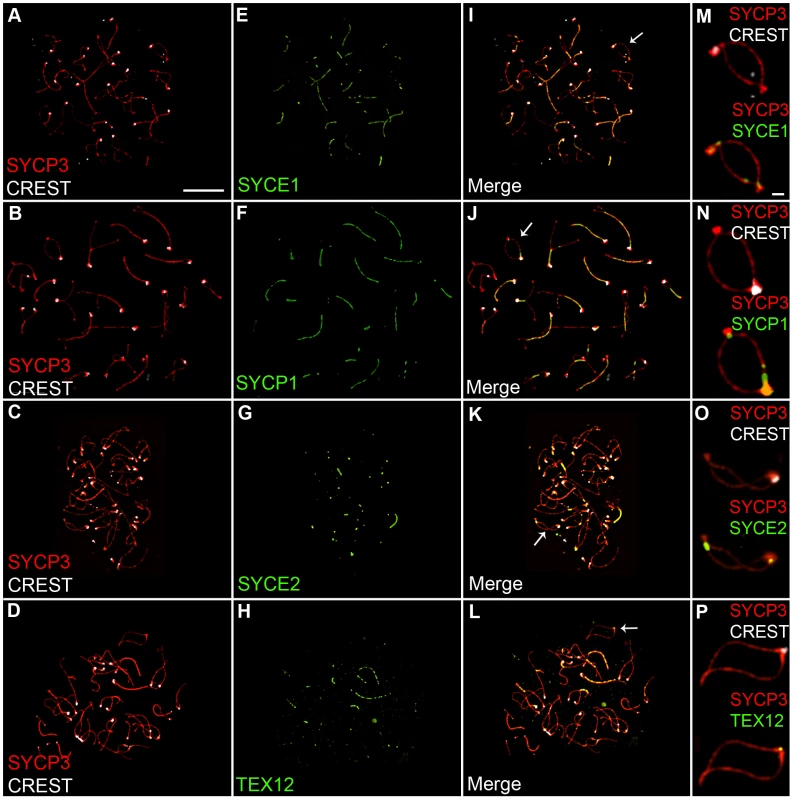
The SYCP1 staining associated with centromeres and nascent chiasmata during diplonema could reflect the retention of fragments of normal SC, or modified structures peculiar to these sites. To begin to distinguish these possibilities, we examined the localization of three SC central element proteins, SYCE1, SYCE2 and TEX12, in addition to SYCP1 [59]–[61]. As shown in Figure 2, all four central element components localize to sites of paired centromeres and chiasmata during diplonema, consistent with the idea that normal SC fragments are retained at these sites.
SC Remnants Associated with Centromeres and Chiasmata Are Distinct and Independent
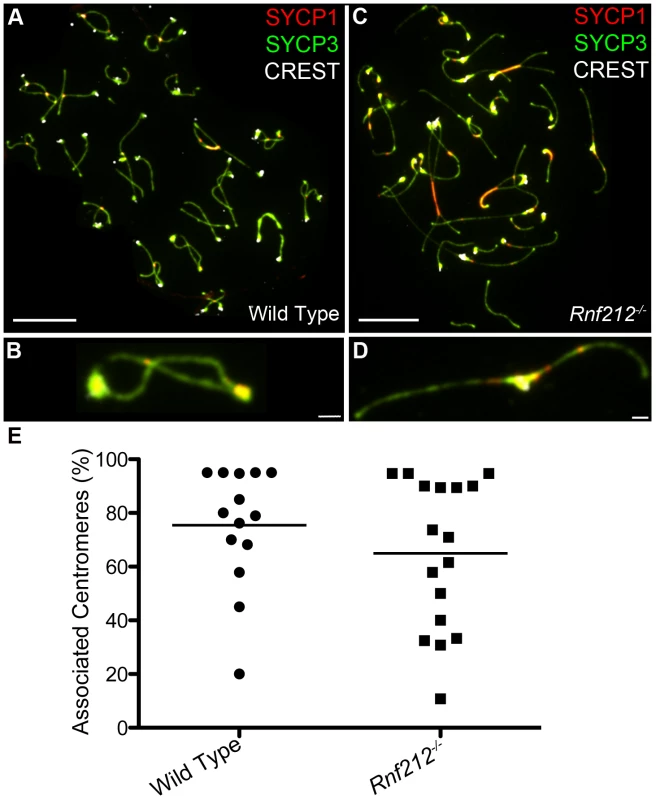
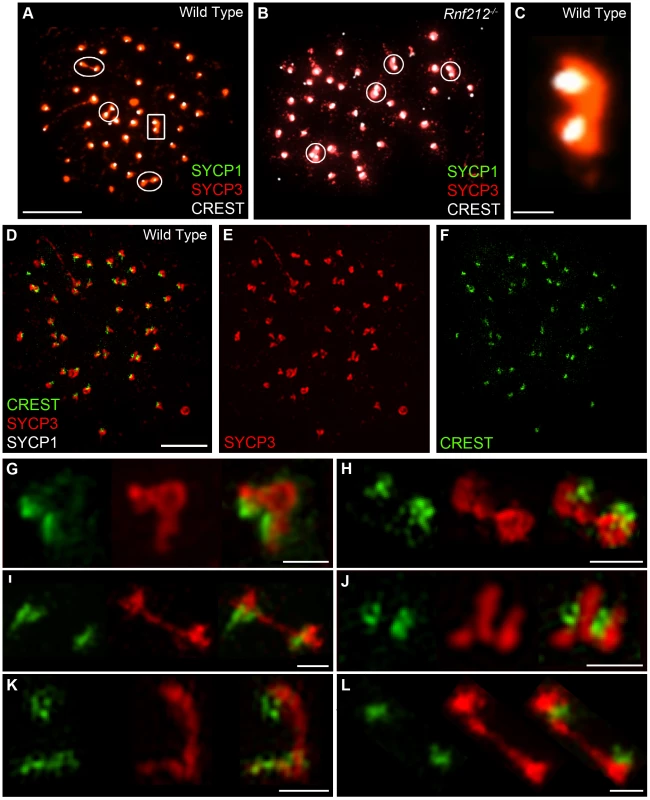
Previous studies have correlated non-centromeric SC fragments at diplonema with crossover sites (identified as silver-staining recombination nodules [6], [50], [52], [62]) but, to our knowledge, dependency of these structures on crossing-over has not been directly demonstrated. On the other hand, we predict that persistent centromere-associated SC fragments should occur independently of crossing-over. These inferences were tested by analyzing Rnf212−/− mutant mice, which have normal homolog synapsis, but show a ≥95% reduction of crossing-over (A.R., H.Q., J.K.C. and N.H., unpublished data) (Figure 3). As diplonema progresses and desynapsis ensues in Rnf212−/− spermatocytes, chromosome arms completely dissociate, but SYCP1-associated centromeres frequently remain connected (Figure 3C–3E). Persistent interstitial SC fragments were not observed in Rnf212−/− nuclei, indicating dependence on crossing over.
SC Remnants Are Fragments of Tripartite Synaptonemal Complex
During diplonema, it is not unusual to detect foci of central element components that remain associated with separated homolog axes (e.g. Figure 2). Thus, the association of central element components with centromeres and crossover sites might not represent true tripartite SC, but merely SC remnants associated with only one homolog axis. These two possibilities were discriminated using structured illumination microscopy (SIM), which has sufficient resolving power to distinguish the two SYCP3-stained SC lateral elements from the SYCP1-stained central element (Figure 4) [63], [64]. SIM imaging reveals that centromere and crossover associated SYCP1 is sandwiched between the two homolog axes, as expected for true tripartite SC (Figure 4A–4D).
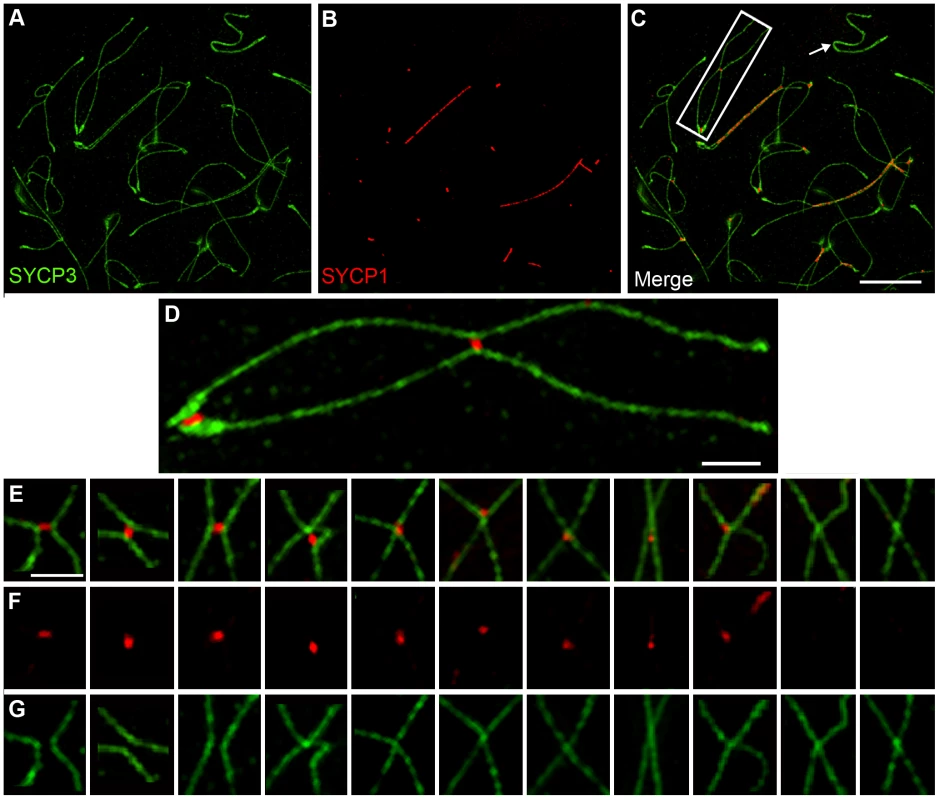
Crossover-Associated SC Fragments Precede Axis Remodeling at Chiasmata
SIM analysis of late diplotene nuclei indicates that crossover-associated SC fragments can be very short, comprising on average only 0.24 µm of SYCP1 (Figure 4), less than 3% of the length of an average late pachytene SC (8.6 µm). Moreover, in 41% of these structures (29/71 from 6 nuclei), the SYCP3-staining homolog axes remain clearly separate implying that they have yet to be exchanged to form chiasmata (Figure 4E–4G). Notably, in mouse, DNA exchange to form crossovers has been shown occur during pachynema [65], [66]. Thus, DNA crossing-over and axis-remodeling appear to be temporally distinct aspects of chiasma formation.
Intriguingly, the 31% of nascent chiasmata sites in which SYCP3 axes converge and begin to fuse (22/71) are associated with smaller and less intense SYCP1 foci, which are typically localized to one side of the presumed axis-exchange point (Figure 4E–4G). Finally, 28% of nascent chiasmata (20/71) comprised SYCP3 fusion-points without associated SYCP1.
SC Central Element Prevents Unregulated Fusion of Homolog Axes at Recombination Sites and Chromosome Ends
The analysis above suggests that SC fragments are retained at crossover sites to regulate the exchange of homolog axes. To further explore this idea, we performed SIM analysis of diplotene-like nuclei from the Sycp1−/− mouse, which fails to form SC central element (Figure 5A, 5B). Although synapsis fails in Sycp1−/− meiocytes, the early steps of recombination occur normally, homologs pair and axes closely associate at sites of recombination (so called, “axial associations”) [17]. In nuclei with late-stage chromosome morphology, previously defined to be in diplonema [17], axial associations appeared as a mixture of separate and conjoined/fused SYCP3 axes (Figure 5C), similar to the nascent chiasmata observed in wild-type diplotene nuclei. However, chiasma-like structures in Sycp1−/− nuclei are distinct from those in wild type. First, they are more numerous, averaging 1.8 (±1.1 SD, n = 89) per homolog pair compared to 0.96 (±0.63 SD, n = 74) in wild type (P<0.0001, Mann-Whitney test). Second, 87.3% (137/157) of chiasma-like structures in Sycp1−/− cells have conjoined or fused axes compared to 59.1% in wild type (42/71; P<0.0002, z-test). This observation suggests that nascent chiasma sites with separate SYCP3 axes are stabilized by SC central elements. Moreover, given that crossovers are almost completely abolished by Sycp1 mutation [17] (as shown by the absence of both crossover-specific MLH1 foci and chiasmata), the observed chiasma-like structures are forming independently of interhomolog crossing over.
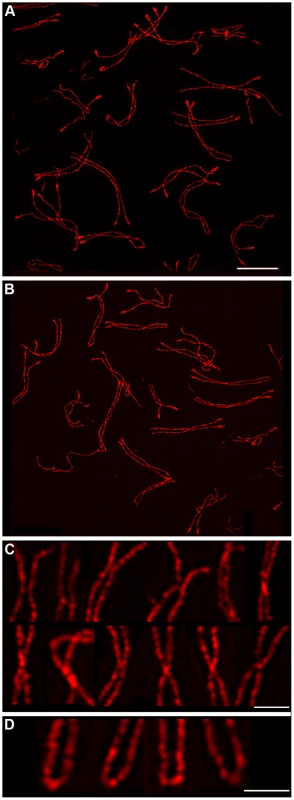
In addition, in Scyp1−/− diplotene cells, we repeatedly observed chromosomes in which the non-centromeric ends of the homolog axes had fused to form contiguous terminal loops (Figure 5D); 15.2% of homolog pairs (14/89) had such structures. Analogous terminal loops were never observed in wild-type diplotene-stage nuclei imaged by SIM. Both the chiasma-like structures and terminal fusions detected in Sycp1−/− spermatocytes suggest a tendency for unregulated interactions between homolog axes in the absence of SC central element.
Features of Diplotene Centromeres Revealed by Structured Illumination Microscopy
Several characteristics of SC-associated centromeres in diplonema were also refined by SIM analysis (Figure 6). First, the well-characterized accumulations of SYCP3 at the centromeric termini [47] form paddle-like structures that can be more than three times broader than the homolog axes (Figure 6A and 6C, highlighted by an arrowhead, and 6D). Second, 7% (12/172) of these terminal SYCP3 structures show a dual morphology suggestive of sister-chromatid individualization; in fact, clear examples of associated centromeres with split sister-axes were observed (Figure 6A and 6C, highlighted by an arrowhead, and 6E). Third, dissociated or even widely separated centromeres can retain SYCP1 staining (Figure 6F). This observation raises the possibility that final desynapsis of centromeres may not occur by simple dissociation of central element proteins from the axes, but by separation of central element transverse filaments (comprising SYCP1 and other proteins) that connect homolog axes via a head-to-head configuration of overlapping homodimers [6]. Alternatively, centromeric connections may have become sufficiently weak that they are mechanically disrupted during the spreading procedure. Fourth, we observed several examples of ostensibly achiasmate homologs, without an internal SYCP3 connection, that remain connected by synapsed centromeres (4/74 homolog pairs; Figure 6G). Whether these homologs are truly achiasmate and their ultimate segregation fate remains unclear, but their detection is consistent with our observation that persistent centromere synapsis occurs independently of crossing-over (above). The possibility that persistent centromere synapsis facilitates the segregation of achiasmate chromosomes is also raised by this observation. Finally, the centromeric ends of the X and Y chromosomes do not show the dramatic SYCP3 accumulation and morphological changes seen for autosomes at this stage. Instead, a general accumulation of SYCP3 signal along the lengths of the X-Y pair is observed (Figure 6C; highlight by and arrow).
Centromere Association during Diplonema Is Mediated by the SC Central Element
The analysis above implies that both the association of centromeres in pachynema and their continued connection during diplonema are mediated by SC central element. To test these inferences, we examined centromere association in spermatocytes from the synapsis-defective Sycp1−/− mutant (Figure 7). For individual pachytene-like Sycp1−/− spermatocyte nuclei, we determined the frequency of associated centromeres as well as the extent of homolog coalignment or “pseudo-synapsis”, which was defined as the fraction of homolog axes that were separated by ≤0.8 µm (Figure 7A–7D; 0.8 µm was determined to be the maximum distance measured between regions of coaligned axes in Sycp1−/− nuclei). This analysis indicates that centromere association is not absolutely dependent on the SC central element (Figure 7D). In fact, we observed several examples in which centromeric ends of homologous SYCP3 axes appear fused with one another to form a contiguous loop, even though adjacent regions are clearly separated (e.g. Figure 7B and 7C). These terminal fusions are distinct from those observed in diplotene-like nuclei, which involve the non-centromeric chromosome ends (above and Figure 5D). Overall, however, centromere pairing in Sycp1−/− spermatocytes never reaches wild-type levels and centromere regions remain the last to pair. Even in nuclei with >70% pseudo-synapsis, ≤50% centromere pairing is observed.
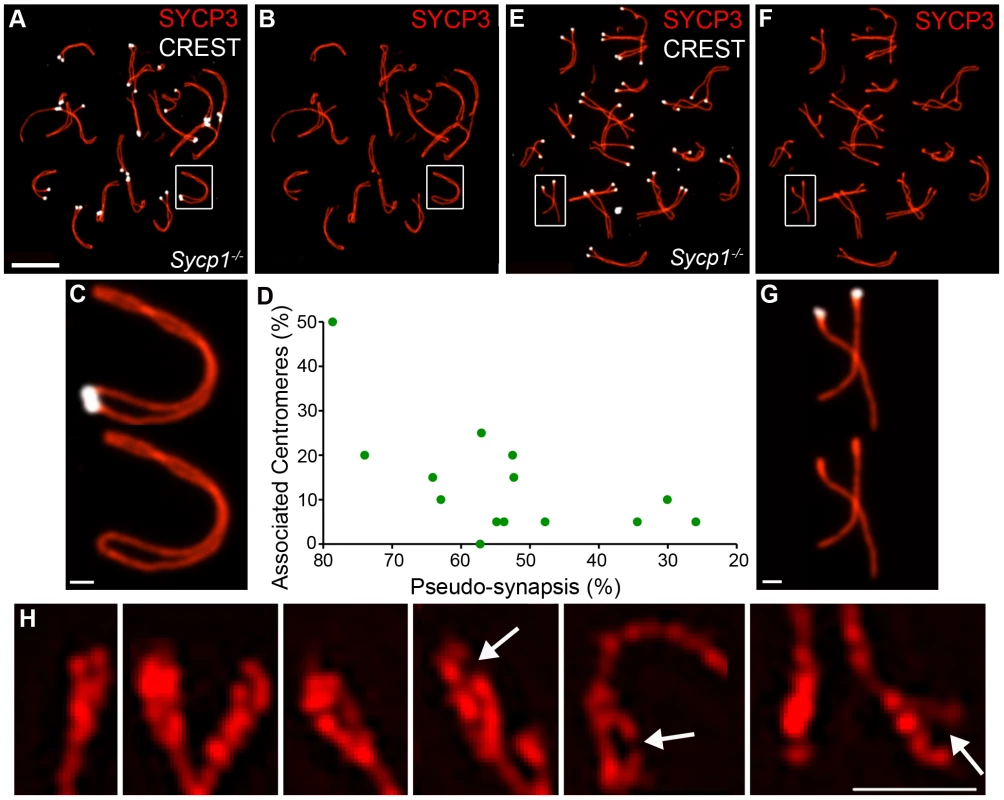
In contrast to the pachytene-like nuclei analyzed above, centromeres are not associated in diplotene-like Sycp1−/− nuclei even though homologs remain stably tethered by one or more axial association (Figure 7E–7G; 0/89 homolog pairs analyzed by SIM had associated centromeres). Thus, persistent centromere association during diplonema is SYCP1 dependent.
SIM Analysis of Centromeric SYCP3 Structures from Sycp1 Mutant Spermatocytes
A number of possible functions can be imagined for centromeric SC fragments during diplonema. For example, continued synapsis of centromeres could resist or maybe even promote the splitting of sister-axes that is detected at this stage (described above; Figure 6E); or it could promote the reorganization of centromere regions, such as the accumulation of SYCP3, modification of cohesion and assembly of kinetochore components; or persistent centromere synapsis could indirectly facilitate homolog biorientation, for example by helping establish connections between centromeric heterochromatin similar to those described in Drosophila [67].
To begin to explore these possibilities, SYCP3-stained diplotene-like nuclei from Sycp1−/− spermatocytes were imaged by SIM (Figure 7H; also see Figure 5). Thickening of the SYCP3-stained centromeric termini was still clearly apparent in Sycp1−/− mutants (Figure 7H). However, duality and splitting, indicative of sister-chromatid individualization, was exaggerated in Sycp1−/− cells (Figure 7H), being observed at 19.2% (34/177) of centromeric termini compared to 7% (12/172) of wild-type ends (P = 0.0007, z-test). Moreover, Sycp1−/− centromeric termini had a more fragile and fractured appearance (Figure 7H, arrows highlight gaps or fractures); 13.6% (24/177) of centromeric ends had clear gaps or breaks, a morphology that was never observed in wild-type cells. Thus, diplotene-stage centromeric SYCP3 structures appear to be stabilized by continued synapsis.
Evidence for Persistent Centromere Linkages after SYCP1 Dissociation
SYCP1 staining is lost and homologous centromeres desynapse in late diplonema (above). Therefore, unlike Zip1-mediated coupling in budding yeast, persistent centromeres synapsis seems unlikely to play a direct role in promoting the stable biorientation of homologs on the meiosis I spindle, which doesn't assemble until diakinesis when the nuclear membrane breaks down.
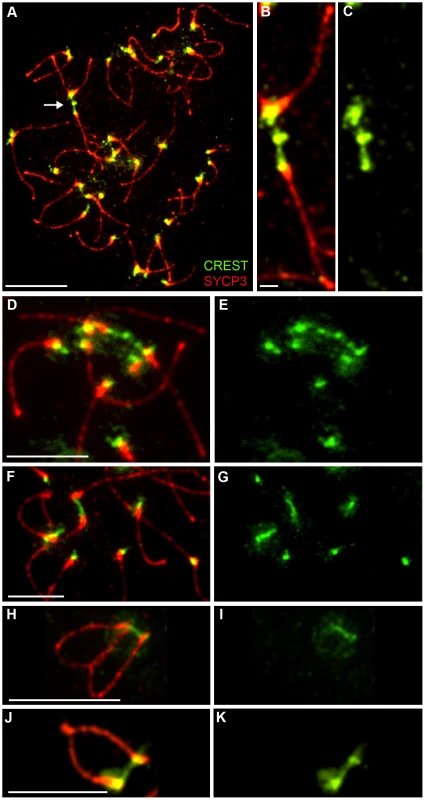
Although homologous centromeres desynapse during late diplonema, we noted that they often appear closely associated and oriented towards one another, even in the absence of chiasmata (in Rnf212−/− spermatocytes). Moreover, we routinely detected inter-centromeric CREST-staining structures at this stage (Figure 8) giving the impression of interconnecting chromatin bridges. However, distinct DAPI-staining bridges cannot be discerned at this stage because the chromatin is very diffuse (not shown).
As described previously, axial SYCP3 mostly disappears from diakinesis/metaphase-I chromosome axes to leave only faint interchromatid foci that define the chiasmata (e.g. Figure 1N) [47]. In contrast, centromeric SYCP3 becomes more abundant and remains closely associated with CREST-staining kinetochores [47], [68], [69]. Intriguingly, in diakinesis/metaphase I nuclei, we regularly detected closely apposed CREST signals associated with apparently contiguous, bi-lobed SYCP3 structures, or structures that are connected by thin SYCP3-staining strands (Figure 9A–9C). The close apposition of the centromeres in these structures could, in theory, be caused by proximal chiasmata (although crossover-specific MLH1 foci are rarely found close to centromeres). However, analysis of crossover-defective Rnf212−/− mutant nuclei indicates that they arise independently of chiasmata (Figure 9B). On average, around two SYCP3-linked centromere pairs were observed in both wild-type and Rnf212−/− spermatocytes (1.9±1.8 SD, n = 13 and 2.2±1.4 SD, n = 18).
SIM analysis of diakinesis/metaphase-I nuclei supports the inference that these linked centromere-pairs are associated with contiguous SYCP3-staining structures (Figure 9D–9L). In one example, discontinuous CREST staining also appears to bridge between the two homolog-kinetochores (Figure 9G). Taken together, our observations support the possibility that the centromere regions of homologs can remain interconnected long after they have desynapsed during late diplonema.
Discussion
Centromeres Do Not Drive Homolog Synapsis in Mammals
In contrast to budding yeast and Drosophila, mouse centromeres appear to be refractory to SC formation and are the last sites to synapse. The same conclusions are drawn by Bisig et al. in the accompanying study [48]. The possibility that this is a general feature of mammalian meiosis is supported by the study of Hassold and colleagues, which showed that centromeres of human spermatocyte chromosomes constitute a barrier to the polymerization of SC [11], [53]. In fact, the early synapsis of budding yeast and Drosophila centromeres may be more the exception than the rule, as late pericentric synapsis is typical of many organisms including fungi, plants and mammals [11], [49]–[53].
We conclude that centromeres do not drive SC formation in mammals, but are in fact refractory to synapsis. This is true despite the fact that during zygonema, mouse centromeres cluster into a single large chromocenter, which might have been expected to facilitate centromere synapsis [70]. Indeed, recent studies in Drosophila females show that synapsis initiates within the chromocenter, indicating a fundamental difference with mammals [19], [30], [31]. We suggest that mammalian centromeres are synapsed only after SC polymerization switches from the initial homology-dependent, recombination-driven phase to the well-characterized (but poorly understood) homology-indifferent “synaptic adjustment” mode [6], [71].
How and why mammalian centromeres resist synapsis remains unclear. Random (non-homologous) association of centromeric major satellite DNA within chromocenters could oppose forces that attempt to drive homologous pairing and synapsis of centromeres. Also, suppression of recombination initiation close to centromeres could limit not only local SC initiation, but also the extension of SC polymerization from adjacent sites. Another possibility is that chromatin and/or axial structures associated with centromeres are modified in ways that impede early synapsis. Notably, Roig et al. [51] showed that centromere synapsis is unusually dependent on the AAA+ ATPase, TRIP13, which facilitates the removal of HORMA-domain proteins from synapsing homolog axes [72].
SC initiation sites have been correlated with crossing-over in a number of organisms. However, it remains unclear whether crossover-designation triggers SC formation, or SC initiation sites trigger crossing-over [2], [6], [11], [73]. Under the latter scenario, absence of SC initiation between centromeres could function to suppress crossing-over within the multi-megabase, repetitive DNA elements that constitute mammalian centromeres [74]. This in turn will minimize the risk of chromosome rearrangements that could result from non-allelic centromere exchanges. Suppression of allelic crossing-over near centromeres will also help minimize the incidence of nondisjunction, which has been associated with such events in yeast, flies and humans [27], [75]. By contrast, the tiny size of budding yeast centromeres (125 bp) makes them highly unlikely to engage in non-allelic crossing-over. We also note that although Drosophila centromeres serve as SC initiation sites [30], [31], they may be protected from meiotic instability by the fact that SC initiation sites do not correlate with crossing-over in this organism. Moreover, homolog pairing and synapsis are not driven by recombination in Drosophila.
Crossover-Dependent SC Fragments May Stabilize Nascent Chiasmata and May Coordinate Axis Exchange and Bivalent Maturation
Our analysis implies that DNA crossing-over and axis remodeling (at least with respect to SYCP3) are temporally distinct aspects of chiasma formation. Analysis of Sycp1−/− spermatocytes suggests that this temporal separation may be mediated by the SC central element. Furthermore, the chiasma-like structures and terminal fusions observed in Sycp1−/− mutants suggest a novel role for SC central element in preventing the unregulated fusion and exchange of homolog axes.
After diplonema, chromatin condenses and sister-chromatids individualize to become located on opposite sides of their cohesin axis. As bivalents further condense, chromatids also bend sharply at sites of crossing-over [14]. The requisite local flexibility appears to be reflected by two morphological features of crossover sites: relaxation of sister-chromatid cohesion and reduced chromatin condensation [6]. We suggest that SC fragments could help implement these features by triggering local loss of cohesin and/or differential loading of condensin (the loading of which may be coupled to SC disassembly [76]).
A role for crossovers in bivalent remodeling has been clearly demonstrated in C. elegans. In this organism, crossover sites trigger asymmetric loss of SC components and, subsequently, cohesion from bivalent arms [77], [78]. This global remodeling of bivalents may be peculiar to organisms with holocentric chromosomes. In organisms with conventional centromeres, in which all arm cohesion is lost at anaphase I, we suggest that crossovers only trigger local changes in cohesion and chromatin condensation, as described above.
A Role for Diplotene SC Fragments in Centromere Function?
SIM analysis has revealed a tendency for local separation of sister-chromatid axes at synpased centromeres during diplonema. Kleckner et al. have proposed that cycles or chromatin expansion and contraction drive such transient individualization of sister-chromatids in order to facilitate chromosome remodeling and installation of components required for subsequent stages [79]. The enhanced splitting of centromeric SYCP3 structures seen in Sycp1−/− mutants supports the idea that SC fragments retained at diplonema are part of a supporting framework that constrains and targets local expansion to help coordinate remodeling at centromere regions.
The chromosomal passenger complex (CPC) regulates and orchestrates several key processes during chromosome segregation and cell division. These include sister-chromatid cohesion, kinetochore-microtubule attachments, spindle stability and cell division [80]. In addition, during meiosis the CPC regulates the timing of SC disassembly [76], [81], [82]. Cytological analyses of mouse spermatocytes have shown that CPC components, INCENP and Aurora-B, relocalize from centromeric heterochromatin to the inner centromere domain during diplonema, i.e. concurrent with the retention of SC at centromeres [83], [84]. In addition, INCENP associates with the SC central element [84]. These observations raise the intriguing possibility that centromere-associated SC fragments in diplonema facilitate CPC relocalization and initial stages of kinetochore maturation.
What Are the Signals for Local SC Retention?
Centromere pairing in budding yeast requires PP4-dependent dephosphorylation of the SC component, Zip1 (which is phosphorylated in response to DSB formation; [23]). In addition, the budding yeast SC central element component, Zip1, can bind SUMO, which is a prominent modification at centromeric heterochromatin and kinetochores [56], [85]. In Drosophila females, the CPC stabilizes SCs presumably by antagonizing kinases that promote SC disassembly (see above [76], [81]). Thus, the high concentration of CPC at spermatocyte centromeres could promote local resistance to SC disassembly. How crossovers signal local retention of SC remains mysterious. In rat, the crossover marker, CDK2, remains at crossover sites until diplonema and could signal SC retention [86]. However, in male mice, CDK2 does not obviously persist at crossover sites beyond pachynema [87].
Post-Synapsis Bridges Suggest Tethering of Homologous Centromeres
Persistent association of centromeres throughout diplonema appears to be a conserved feature of meiosis in many organisms, including budding yeast, Drosophila and mouse ([4], [21], [30], [32], [50], [52], [62] and this study). In budding yeast, late centromere coupling promotes the correct, bipolar (syntelic) attachment of chiasmate bivalents to the spindle and thereby limits engagement of the spindle assembly checkpoint to correct misalignments. Coupling also serves as a backup mechanism for the disjunction of occasional achiasmate chromosomes [21], [32], [33]. The role of late centromere synapsis in Drosophila remains unclear, but association of centromeric heterochromatin is important for achiasmate segregation in this organism [88]–[90].
It seems unlikely that the persistent centromere synapsis observed in mouse is directly analogous to centromere coupling in budding yeast. Notably, centromere synapsis does not persist beyond diplonema so that a direct role in homolog biorientation and achiasmate disjunction is not envisioned. However, coupling could theoretically function indirectly in these processes by promoting centromere association, orientation and/or the organization of kinetochores prior to nuclear envelope breakdown and spindle assembly.
The inter-centromeric CREST-staining bridges we detect in late-diplotene/early diakinesis cells are reminiscent of the heterochromatin threads that connect achiasmate (and perhaps chiasmate) chromosomes during meiosis in Drosophila females [67]. These structures are proposed to facilitate the congression of achiasmate chromosomes during prometaphase and promote their disjunction at anaphase I. The achiasmate X-Y disjunction systems found in some mammals appear to use specialized structures, derived from SC components, to tether the X and Y chromosomes [35]–[37]. The inter-centromeric CREST bridges and SYCP3 structures that we detect in diakinesis/metaphase-I spermatocytes might reflect the existence of related processes in mouse that can favor the biorientation of homologous centromeres and/or facilitate the disjunction of chromosomes that have failed to crossover.
Materials and Methods
Ethics Statement
All experiments conformed to relevant regulatory standards and were approved by the U.C Davis Institutional Animal Care and Use Committee.
Mice
All mice were congenic with the C57BL/6J background. The Sycp1 and Spo11 knock-out lines were previously described [17], [91]. Generation of the Rnf212 knock-out line will be described elsewhere (Reynolds et al., submitted). PCR genotyping of Rnf212 mice was performed using primers exon forward (5′-CGCTGGAATGAACGCAGGCGC-3′), exon reverse (5′-CAGGGGAGTGAAGCCACGGTC-3′), pH530 (5′-TCCATGGGCTTAAACCAGTGC-3′), and VM3 (5′-GCGCATGCTCCAGACTGCCTTG-3′). Primers, exon forward and exon reverse, generate a 290-bp fragment diagnostic of the Rnf212 wild-type allele; pH530 and VM3 detect the Rnf212 mutant allele as a 383-bp fragment. PCR conditions were 30 seconds at 94°C, 30 seconds at 60°C, and 1 minute at 72°C for 30 cycles.
Cytology
Testes were removed from 2–4 month old mice and processed for surface spreading as described [92]. Immunofluorescence staining was performed as described [93] using the following primary antibodies overnight at room temperature (dilutions in parentheses): rabbit anti-SYCP3 (sc-33195 Santa Cruz, 1∶300); mouse anti-SYCP3 (sc-74568 Santa Cruz, 1∶200); mouse anti-rat SYCP1 monoclonal antibody [94] (1∶400); CREST antiserum (generously provided by Shelby White, ARUP Laboratories; 1∶10000); mouse monoclonal anti-γH2AX (05-636 Millipore, 1∶500), rabbit anti-mouse RAD21L (a generously gift of K. Ishiguro and Y. Watanabe, University of Tokyo [45] (1∶200); guinea pig anti-SYCE1 (1∶2000), guinea pig anti-SYCE2 (1∶400) and guinea pig anti-TEX12 (1∶200) [95], [96]. Slides were subsequently incubated with the following goat secondary antibodies for 1 hour at 37°C: anti-rabbit 488 (A11070 Molecular Probes, diluted 1∶10000), anti-rabbit 568 (A11036 Molecular Probes, diluted 1∶2000), anti-human 488 (A11013 Molecular Probes, 1∶2000), anti-mouse 594 (A11020 Molecular Probes, 1∶10000), anti-human DyLight 649 (109-495-088 Jackson Labs, 1∶200), and anti-guinea pig fluorescein isothiocyanate (106-096-006 FITC, Jackson Labs, 1∶200). Coverslips were mounted with ProLong Gold antifade reagent (Molecular Probes).
Imaging
Immunolabeled chromosome spreads were imaged using a Zeiss AxioPlan II microscope with 63× Plan Apochromat 1.4 objective and EXFO X-Cite metal halide light source. Images were captured by a Hamamatsu ORCA-ER CCD camera. Image processing and measurements were performed using Volocity (Perkin Elmer) and Photoshop (Adobe) software packages. Any pair of CREST foci that was ≤0.6 µm apart was classified as associated; convergent SYCP1 staining defined synapsed centromeres. To account for overlapping CREST foci, total numbers of CREST foci were counted for all nuclei. In nearly all cases, overlapping pairs of CREST foci could be discerned as larger, more intense, bi-lobed staining structures. Only nuclei for which all centromeres could be accounted for were used to determine levels of centromere association/synapsis. SIM analysis was performed using a Nikon N-SIM super-resolution microscope system and NIS-Elements 2 image processing software.
Supporting Information
Zdroje
1. HunterN 2004 Lennarz WaLM Meiosis San Diego Elsevier Press
2. ZicklerD 2006 From early homologue recognition to synaptonemal complex formation. Chromosoma 115 158 174
3. BhallaNDernburgAF 2008 Prelude to a division. Annu Rev Cell Dev Biol 24 397 424
4. von WettsteinDRasmussenSWHolmPB 1984 The synaptonemal complex in genetic segregation. Annu Rev Genet 18 331 413
5. PageSLHawleyRS 2004 The genetics and molecular biology of the synaptonemal complex. Annu Rev Cell Dev Biol 20 525 558
6. ZicklerDKlecknerN 1999 Meiotic chromosomes: integrating structure and function. Annu Rev Genet 33 603 754
7. CostaYCookeHJ 2007 Dissecting the mammalian synaptonemal complex using targeted mutations. Chromosome Res 15 579 589
8. YangFWangPJ 2009 The Mammalian synaptonemal complex: a scaffold and beyond. Genome Dyn 5 69 80
9. PelttariJHojaMRYuanLLiuJGBrundellE 2001 A meiotic chromosomal core consisting of cohesin complex proteins recruits DNA recombination proteins and promotes synapsis in the absence of an axial element in mammalian meiotic cells. Mol Cell Biol 21 5667 5677
10. KouznetsovaANovakIJessbergerRHoogC 2005 SYCP2 and SYCP3 are required for cohesin core integrity at diplotene but not for centromere cohesion at the first meiotic division. J Cell Sci 118 2271 2278
11. HendersonKAKeeneyS 2005 Synaptonemal complex formation: where does it start? Bioessays 27 995 998
12. HunterN 2006 Meiotic Recombination. AguileraARothsteinR Molecular Genetics of Recombination Heidelberg Springer-Verlag 381 442
13. TsaiJHMcKeeBD 2011 Homologous pairing and the role of pairing centers in meiosis. J Cell Sci 124 1955 1963
14. JonesGH 1987 Chiasmata. MoensPB Meiosis Orlando Academic Press 213 244
15. SakunoTTanakaKHaufSWatanabeY 2011 Repositioning of aurora B promoted by chiasmata ensures sister chromatid mono-orientation in meiosis I. Dev Cell 21 534 545
16. HiroseYSuzukiROhbaTHinoharaYMatsuharaH 2011 Chiasmata promote monopolar attachment of sister chromatids and their co-segregation toward the proper pole during meiosis I. PLoS Genet 7 e1001329 doi:10.1371/journal.pgen.1001329
17. de VriesFAde BoerEvan den BoschMBaarendsWMOomsM 2005 Mouse Sycp1 functions in synaptonemal complex assembly, meiotic recombination, and XY body formation. Genes Dev 19 1376 1389
18. SymMEngebrechtJARoederGS 1993 ZIP1 is a synaptonemal complex protein required for meiotic chromosome synapsis. Cell 72 365 378
19. SubramanianVVHochwagenA 2011 Centromere clustering: where synapsis begins. Curr Biol 21 R920 922
20. StewartMNDawsonDS 2008 Changing partners: moving from non-homologous to homologous centromere pairing in meiosis. Trends Genet 24 564 573
21. KempBBoumilRMStewartMNDawsonDS 2004 A role for centromere pairing in meiotic chromosome segregation. Genes Dev 18 1946 1951
22. TsubouchiTRoederGS 2005 A synaptonemal complex protein promotes homology-independent centromere coupling. Science 308 870 873
23. FalkJEChanACHoffmannEHochwagenA 2010 A Mec1- and PP4-dependent checkpoint couples centromere pairing to meiotic recombination. Dev Cell 19 599 611
24. ChenSYTsubouchiTRockmillBSandlerJSRichardsDR 2008 Global analysis of the meiotic crossover landscape. Dev Cell 15 401 415
25. ObesoDDawsonDS 2010 Temporal characterization of homology-independent centromere coupling in meiotic prophase. PLoS ONE 5 e10336 doi:10.1371/journal.pone.0010336
26. HassoldTHallHHuntP 2007 The origin of human aneuploidy: where we have been, where we are going. Hum Mol Genet 16 Spec No. 2 R203 208
27. RockmillBVoelkel-MeimanKRoederGS 2006 Centromere-proximal crossovers are associated with precocious separation of sister chromatids during meiosis in Saccharomyces cerevisiae. Genetics 174 1745 1754
28. TsubouchiTMacqueenAJRoederGS 2008 Initiation of meiotic chromosome synapsis at centromeres in budding yeast. Genes Dev 22 3217 3226
29. MacqueenAJRoederGS 2009 Fpr3 and Zip3 ensure that initiation of meiotic recombination precedes chromosome synapsis in budding yeast. Curr Biol 19 1519 1526
30. TakeoSLakeCMMorais-de-SaESunkelCEHawleyRS 2011 Synaptonemal complex-dependent centromeric clustering and the initiation of synapsis in Drosophila oocytes. Curr Biol 21 1845 1851
31. TannetiNSLandyKJoyceEFMcKimKS 2011 A pathway for synapsis initiation during zygotene in Drosophila oocytes. Curr Biol 21 1852 1857
32. NewnhamLJordanPRockmillBRoederGSHoffmannE 2010 The synaptonemal complex protein, Zip1, promotes the segregation of nonexchange chromosomes at meiosis I. Proc Natl Acad Sci U S A 107 781 785
33. CheslockPSKempBJBoumilRMDawsonDS 2005 The roles of MAD1, MAD2 and MAD3 in meiotic progression and the segregation of nonexchange chromosomes. Nat Genet 37 756 760
34. HawleyRSTheurkaufWE 1993 Requiem for distributive segregation: achiasmate segregation in Drosophila females. Trends Genet 9 310 317
35. PageJBerriosSRufasJSParraMTSujaJA 2003 The pairing of X and Y chromosomes during meiotic prophase in the marsupial species Thylamys elegans is maintained by a dense plate developed from their axial elements. J Cell Sci 116 551 560
36. PageJVieraAParraMTde la FuenteRSujaJA 2006 Involvement of synaptonemal complex proteins in sex chromosome segregation during marsupial male meiosis. PLoS Genet 2 e136 doi:10.1371/journal.pgen.0020136
37. de la FuenteRParraMTVieraACalventeAGomezR 2007 Meiotic pairing and segregation of achiasmate sex chromosomes in eutherian mammals: the role of SYCP3 protein. PLoS Genet 3 e198 doi:10.1371/journal.pgen.0030198
38. SolariAJ 1974 The behavior of the XY pair in mammals. Int Rev Cytol 38 273 317
39. BurgoynePS 1982 Genetic homology and crossing over in the X and Y chromosomes of Mammals. Hum Genet 61 85 90
40. OttoSPPannellJRPeichelCLAshmanTLCharlesworthD 2011 About PAR: the distinct evolutionary dynamics of the pseudoautosomal region. Trends Genet 27 358 367
41. EarnshawWCRothfieldN 1985 Identification of a family of human centromere proteins using autoimmune sera from patients with scleroderma. Chromosoma 91 313 321
42. de BoerEHeytingC 2006 The diverse roles of transverse filaments of synaptonemal complexes in meiosis. Chromosoma 115 220 234
43. LeeJHiranoT 2011 RAD21L, a novel cohesin subunit implicated in linking homologous chromosomes in mammalian meiosis. J Cell Biol 192 263 276
44. HerranYGutierrez-CaballeroCSanchez-MartinMHernandezTVieraA 2011 The cohesin subunit RAD21L functions in meiotic synapsis and exhibits sexual dimorphism in fertility. Embo J 30 3091 3105
45. IshiguroKKimJFujiyama-NakamuraSKatoSWatanabeY 2011 A new meiosis-specific cohesin complex implicated in the cohesin code for homologous pairing. EMBO Rep 12 267 275
46. LukasJLukasCBartekJ 2011 More than just a focus: The chromatin response to DNA damage and its role in genome integrity maintenance. Nat Cell Biol 13 1161 1169
47. ParraMTVieraAGomezRPageJBenaventeR 2004 Involvement of the cohesin Rad21 and SCP3 in monopolar attachment of sister kinetochores during mouse meiosis I. J Cell Sci 117 1221 1234
48. BisigCGGuiraldelliMFKouznetsovaAScherthanAHHöögC 2012 Synaptonemal complex components persist at centromeres and are required for homologous centromere pairing in mouse spermatocytes. PLoS Genet 8 doi:10.1371/journal.pgen.1002701 e1002701
49. ZicklerDMoreauPJHuynhADSlezecAM 1992 Correlation between pairing initiation sites, recombination nodules and meiotic recombination in Sordaria macrospora. Genetics 132 135 148
50. ZicklerD 1977 Development of the synaptonemal complex and the “recombination nodules” during meiotic prophase in the seven bivalents of the fungus Sordaria macrospora Auersw. Chromosoma 61 289 316
51. RoigIDowdleJATothAde RooijDGJasinM 2010 Mouse TRIP13/PCH2 is required for recombination and normal higher-order chromosome structure during meiosis. PLoS Genet 6 e1001062 doi.org/10.1371/journal.pgen.1001062
52. HolmPBRasmussenSWZicklerDLuBCSageJ 1981 Chromosome pairing, recombination nodules and chiasma formation in the basidiomycete Coprinus cinereus. Carlsberg Res Commun 46 305 346
53. BrownPWJudisLChanERSchwartzSSeftelA 2005 Meiotic synapsis proceeds from a limited number of subtelomeric sites in the human male. Am J Hum Genet 77 556 566
54. PerryJPalmerSGabrielAAshworthA 2001 A short pseudoautosomal region in laboratory mice. Genome Res 11 1826 1832
55. AgarwalSRoederGS 2000 Zip3 provides a link between recombination enzymes and synaptonemal complex proteins. Cell 102 245 255
56. ChengCHLoYHLiangSSTiSCLinFM 2006 SUMO modifications control assembly of synaptonemal complex and polycomplex in meiosis of Saccharomyces cerevisiae. Genes Dev 20 2067 2081
57. PerryJKlecknerNBornerGV 2005 Bioinformatic analyses implicate the collaborating meiotic crossover/chiasma proteins Zip2, Zip3, and Spo22/Zip4 in ubiquitin labeling. Proc Natl Acad Sci U S A 102 17594 17599
58. KongAThorleifssonGStefanssonHMassonGHelgasonA 2008 Sequence variants in the RNF212 gene associate with genome-wide recombination rate. Science 319 1398 1401
59. Bolcun-FilasEHallESpeedRTaggartMGreyC 2009 Mutation of the mouse Syce1 gene disrupts synapsis and suggests a link between synaptonemal complex structural components and DNA repair. PLoS Genet 5 e1000393 doi:10.1371/journal.pgen.1000393
60. Bolcun-FilasECostaYSpeedRTaggartMBenaventeR 2007 SYCE2 is required for synaptonemal complex assembly, double strand break repair, and homologous recombination. J Cell Biol 176 741 747
61. HamerGWangHBolcun-FilasECookeHJBenaventeR 2008 Progression of meiotic recombination requires structural maturation of the central element of the synaptonemal complex. J Cell Sci 121 2445 2451
62. HolmPBRasmussenSW 1983 Human meiosis. VII. Chiasma formation in human spermatocytes. Carlsberg Res Coinrnitn 48 415 456
63. WangCJCarltonPMGolubovskayaINCandeWZ 2009 Interlock formation and coiling of meiotic chromosome axes during synapsis. Genetics 183 905 915
64. CarltonPM 2008 Three-dimensional structured illumination microscopy and its application to chromosome structure. Chromosome Res 16 351 365
65. GuillonHBaudatFGreyCLiskayRMde MassyB 2005 Crossover and noncrossover pathways in mouse meiosis. Mol Cell 20 563 573
66. GuillonHde MassyB 2002 An initiation site for meiotic crossing-over and gene conversion in the mouse. Nat Genet 32 296 299
67. HughesSEGillilandWDCotittaJLTakeoSCollinsKA 2009 Heterochromatic threads connect oscillating chromosomes during prometaphase I in Drosophila oocytes. PLoS Genet 5 e1000348 doi:10.1371/journal.pgen.1000348
68. MoensPBSpyropoulosB 1995 Immunocytology of chiasmata and chromosomal disjunction at mouse meiosis. Chromosoma 104 175 182
69. DobsonMJPearlmanREKaraiskakisASpyropoulosBMoensPB 1994 Synaptonemal complex proteins: occurrence, epitope mapping and chromosome disjunction. J Cell Sci 107 Pt 10 2749 2760
70. ScherthanHWeichSSchweglerHHeytingCHarleM 1996 Centromere and telomere movements during early meiotic prophase of mouse and man are associated with the onset of chromosome pairing. J Cell Biol 134 1109 1125
71. MosesMJDresserMEPoormanPA 1984 Composition and role of the synaptonemal complex. EvansCWDickinsonHG Controlling Events in Meiosis 245 270
72. WojtaszLDanielKRoigIBolcun-FilasEXuH 2009 Mouse HORMAD1 and HORMAD2, two conserved meiotic chromosomal proteins, are depleted from synapsed chromosome axes with the help of TRIP13 AAA-ATPase. PLoS Genet 5 e1000702 doi:10.1371/journal.pgen.1000702
73. AndersonLKStackSM 2005 Recombination nodules in plants. Cytogenet Genome Res 109 198 204
74. BlackBEJansenLEFoltzDRClevelandDW 2010 Centromere identity, function, and epigenetic propagation across cell divisions. Cold Spring Harb Symp Quant Biol 75 403 418
75. HassoldTHuntP 2001 To err (meiotically) is human: the genesis of human aneuploidy. Nat Rev Genet 2 280 291
76. ResnickTDDejKJXiangYHawleyRSAhnC 2009 Mutations in the chromosomal passenger complex and the condensin complex differentially affect synaptonemal complex disassembly and metaphase I configuration in Drosophila female meiosis. Genetics 181 875 887
77. NabeshimaKVilleneuveAMColaiacovoMP 2005 Crossing over is coupled to late meiotic prophase bivalent differentiation through asymmetric disassembly of the SC. J Cell Biol 168 683 689
78. Martinez-PerezESchvarzsteinMBarrosoCLightfootJDernburgAF 2008 Crossovers trigger a remodeling of meiotic chromosome axis composition that is linked to two-step loss of sister chromatid cohesion. Genes Dev 22 2886 2901
79. KlecknerNZicklerDJonesGHDekkerJPadmoreR 2004 A mechanical basis for chromosome function. Proc Natl Acad Sci U S A 101 12592 12597
80. RuchaudSCarmenaMEarnshawWC 2007 Chromosomal passengers: conducting cell division. Nat Rev Mol Cell Biol 8 798 812
81. MalmancheNOwenSGegickSSteffensenSTomkielJE 2007 Drosophila BubR1 is essential for meiotic sister-chromatid cohesion and maintenance of synaptonemal complex. Curr Biol 17 1489 1497
82. JordanPCopseyANewnhamLKolarELichtenM 2009 Ipl1/Aurora B kinase coordinates synaptonemal complex disassembly with cell cycle progression and crossover formation in budding yeast meiosis. Genes Dev 23 2237 2251
83. ParraMTGomezRVieraALlanoEPendasAM 2009 Sequential assembly of centromeric proteins in male mouse meiosis. PLoS Genet 5 e1000417 doi:10.1371/journal.pgen.1000417
84. ParraMTVieraAGomezRPageJCarmenaM 2003 Dynamic relocalization of the chromosomal passenger complex proteins inner centromere protein (INCENP) and aurora-B kinase during male mouse meiosis. J Cell Sci 116 961 974
85. La SalleSSunFZhangXDMatunisMJHandelMA 2008 Developmental control of sumoylation pathway proteins in mouse male germ cells. Dev Biol 321 227 237
86. EijpeMOffenbergHJessbergerRRevenkovaEHeytingC 2003 Meiotic cohesin REC8 marks the axial elements of rat synaptonemal complexes before cohesins SMC1beta and SMC3. J Cell Biol 160 657 670
87. AshleyTWalpitaDde RooijDG 2001 Localization of two mammalian cyclin dependent kinases during mammalian meiosis. J Cell Sci 114 685 693
88. DernburgAFSedatJWHawleyRS 1996 Direct evidence of a role for heterochromatin in meiotic chromosome segregation. Cell 86 135 146
89. KarpenGHLeMHLeH 1996 Centric heterochromatin and the efficiency of achiasmate disjunction in Drosophila female meiosis. Science 273 118 122
90. HawleyRSIrickHZitronAEHaddoxDALoheA 1992 There are two mechanisms of achiasmate segregation in Drosophila females, one of which requires heterochromatic homology. Dev Genet 13 440 467
91. BaudatFManovaKYuenJPJasinMKeeneyS 2000 Chromosome synapsis defects and sexually dimorphic meiotic progression in mice lacking Spo11. Mol Cell 6 989 998
92. HollowayJKBoothJEdelmannWMcGowanCHCohenPE 2008 MUS81 generates a subset of MLH1-MLH3-independent crossovers in mammalian meiosis. PLoS Genet 4 e1000186 doi:10.1371/journal.pgen.1000186
93. QiaoHLohmillerLAndersonL 2011 Cohesin proteins load sequentially during prophase I in tomato primary microsporocytes. Chromosome Research 19 193 207
94. MeuwissenRLOffenbergHHDietrichAJRiesewijkAvan IerselM 1992 A coiled-coil related protein specific for synapsed regions of meiotic prophase chromosomes. Embo J 11 5091 5100
95. CostaYSpeedROllingerRAlsheimerMSempleCA 2005 Two novel proteins recruited by synaptonemal complex protein 1 (SYCP1) are at the centre of meiosis. J Cell Sci 118 2755 2762
96. HamerGGellKKouznetsovaANovakIBenaventeR 2006 Characterization of a novel meiosis-specific protein within the central element of the synaptonemal complex. J Cell Sci 119 4025 4032
97. RomanienkoPJCamerini-OteroRD 2000 The mouse Spo11 gene is required for meiotic chromosome synapsis. Mol Cell 6 975 987
Štítky
Genetika Reprodukční medicínaČlánek vyšel v časopise
PLOS Genetics
2012 Číslo 6
- Primární hyperoxalurie – aktuální možnosti diagnostiky a léčby
- Mateřský haplotyp KIR ovlivňuje porodnost živých dětí po transferu dvou embryí v rámci fertilizace in vitro u pacientek s opakujícími se samovolnými potraty nebo poruchami implantace
- Intrauterinní inseminace a její úspěšnost
- Akutní intermitentní porfyrie
- Srdeční frekvence embrya může být faktorem užitečným v předpovídání výsledku IVF
Nejčtenější v tomto čísle
- Rumors of Its Disassembly Have Been Greatly Exaggerated: The Secret Life of the Synaptonemal Complex at the Centromeres
- The NSL Complex Regulates Housekeeping Genes in
- Tipping the Balance in the Powerhouse of the Cell to “Protect” Colorectal Cancer
- Interplay between Synaptonemal Complex, Homologous Recombination, and Centromeres during Mammalian Meiosis
