-
Články
- Vzdělávání
- Časopisy
Top články
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Volná místa
Doporučené pozice
Reklama- Praxe
The Regulatory Network of Natural Competence and Transformation of
The human pathogen Vibrio cholerae is an aquatic bacterium frequently encountered in rivers, lakes, estuaries, and coastal regions. Within these environmental reservoirs, the bacterium is often found associated with zooplankton and more specifically with their chitinous exoskeleton. Upon growth on such chitinous surfaces, V. cholerae initiates a developmental program termed “natural competence for genetic transformation.” Natural competence for transformation is a mode of horizontal gene transfer in bacteria and contributes to the maintenance and evolution of bacterial genomes. In this study, we investigated competence gene expression within this organism at the single cell level. We provide evidence that under homogeneous inducing conditions the majority of the cells express competence genes. A more heterogeneous expression pattern was observable on chitin surfaces. We hypothesize that this was the case due to the heterogeneity around the chitin surface, which might vary extensively with respect to chitin degradation products and autoinducers; these molecules contribute to competence induction based on carbon catabolite repression and quorum-sensing pathways, respectively. Therefore, we investigated the contribution of these two signaling pathways to natural competence in detail using natural transformation assays, transcriptional reporter fusions, quantitative RT–PCR, and immunological detection of protein levels using Western blot analysis. The results illustrate that all tested competence genes are dependent on the transformation regulator TfoX. Furthermore, intracellular cAMP levels play a major role in natural transformation. Finally, we demonstrate that only a minority of genes involved in natural transformation are regulated in a quorum-sensing-dependent manner and that these genes determine the fate of the surrounding DNA. We conclude with a model of the regulatory circuit of chitin-induced natural competence in V. cholerae.
Published in the journal: . PLoS Genet 8(6): e32767. doi:10.1371/journal.pgen.1002778
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1002778Summary
The human pathogen Vibrio cholerae is an aquatic bacterium frequently encountered in rivers, lakes, estuaries, and coastal regions. Within these environmental reservoirs, the bacterium is often found associated with zooplankton and more specifically with their chitinous exoskeleton. Upon growth on such chitinous surfaces, V. cholerae initiates a developmental program termed “natural competence for genetic transformation.” Natural competence for transformation is a mode of horizontal gene transfer in bacteria and contributes to the maintenance and evolution of bacterial genomes. In this study, we investigated competence gene expression within this organism at the single cell level. We provide evidence that under homogeneous inducing conditions the majority of the cells express competence genes. A more heterogeneous expression pattern was observable on chitin surfaces. We hypothesize that this was the case due to the heterogeneity around the chitin surface, which might vary extensively with respect to chitin degradation products and autoinducers; these molecules contribute to competence induction based on carbon catabolite repression and quorum-sensing pathways, respectively. Therefore, we investigated the contribution of these two signaling pathways to natural competence in detail using natural transformation assays, transcriptional reporter fusions, quantitative RT–PCR, and immunological detection of protein levels using Western blot analysis. The results illustrate that all tested competence genes are dependent on the transformation regulator TfoX. Furthermore, intracellular cAMP levels play a major role in natural transformation. Finally, we demonstrate that only a minority of genes involved in natural transformation are regulated in a quorum-sensing-dependent manner and that these genes determine the fate of the surrounding DNA. We conclude with a model of the regulatory circuit of chitin-induced natural competence in V. cholerae.
Introduction
The bacterium Vibrio cholerae is a facultative pathogen and the causative agent of the disease cholera. Cholera is far from being extinct and is, in fact, considered a re-emerging disease [1]. The destructive capacity of cholera is demonstrated by its current outbreak in Haiti. According to a recent health bulletin issued by the Ministère de la Santé Publique et de la Population (MSPP) of Haiti and the PAHO, 515'699 cholera cases have been reported from Haiti up to November 30th 2011 with more than 6'942 deaths. This epidemic highlights the fact that new modeling approaches are required to allow for the prediction of cholera outbreaks in time and space [2]–[5]. However, it also initiated discussions on the origin of the V. cholerae strain, which appears more closely related to south Asian strains than to Latin American and U.S. Gulf Coast isolates [6]. This study by Chin et al. [6] once more reminded us of the differentiation power that whole genome sequencing provides. Indeed an earlier study by Rita Colwell and collaborators compared 23 V. cholerae strains isolated over the past 98 years using whole genome sequencing [7]. These authors concluded that “V. cholerae undergoes extensive genetic recombination via lateral gene transfer”. It is therefore of major importance to understand the mechanisms underlying horizontal gene transfer (HGT).
Natural competence for transformation, as one of the three modes of HGT in bacteria, describes the physiological state that allows a bacterium to take up free DNA from the environment. If the internalized DNA is recombined into the chromosome, the bacterium is considered naturally transformed. V. cholerae commonly occurs in aquatic ecosystems, its true habitat, where it intimately associates with zooplankton and their chitinous exoskeleton [8]–[10]. In this context, it has been shown that chitin, the polymer used as the building block of planktonic exoskeletons, induces natural competence for transformation of V. cholerae [11]. Thus, HGT is tightly linked to the environmental niche of V. cholerae and potentially also to the niche of many other Vibrio species. In fact, three other species of the genus Vibrio, V. fischeri, V. vulnificus and V. parahaemolyticus, are naturally transformable in a chitin-dependent manner [12]–[14].
Transforming DNA can be used to repair damaged genes and, therefore, contributes to genome maintenance or to the acquisition of new alleles/genes, which lead to genetic diversity and evolution. Indeed, experimental laboratory microcosm experiments that simulate aquatic environments have succeeded in recapitulating a V. cholerae O1-to-O139 serogroup conversion by means of natural transformation [15]. This result provides a potential explanation for the devastating occurrence of the O139 serogroup variant of V. cholerae. Today this strain is almost undetectable in endemic regions even though researchers have feared its occurrence as the onset of a new and, therefore 8th, cholera pandemic [16]. However, an important lesson can be learned from the emergence of this new strain: by means of horizontal gene transfer (HGT), Vibrio species may exchange genetic material and become more virulent to mankind.
Chitin-induced natural competence and transformation is poorly understood in spite of its importance. Based on a few suggestive experiments on how natural competence could be regulated in V. cholerae, the authors of a previous study proposed a model that involved at least three regulatory pathways [11]: 1) induction by chitin, 2) catabolite repression, and 3) quorum-sensing (QS). We and others followed up on this study and provided further evidence for an involvement of these three pathways [17]–[23]. However, all of these studies have only looked at a population-wide level. This could lead to a lack of information on how competence is regulated within a single cell. This is exemplified in one of the best-studied naturally competent bacterial species, Bacillus subtilis, for which it is known that “a majority of the bacteria being insusceptible and a minority being highly susceptible to transformation” [24]. David Dubnau and collaborators explained why only 10–20% of cells within a B. subtilis population enter the competence state, and demonstrated that such bistability is caused by intrinsic noise in competence gene expression [25], [26]. Here we show, for the first time, that under homogeneous competence–inducing conditions V. cholerae displays a homogeneous expression pattern as the vast majority of cell within a population scored positive for expression of competence genes.
Taking this important finding into consideration, we then moved on to establish an inducible competence system for V. cholerae, which is based on low levels of TfoX production and not on tfoX overexpression as previously done. To date all studies on natural competence and transformation in V. cholerae have only looked at single genes involved in the competence program, at single pathways, and at varying inter-experimental conditions (e.g., chitin surface transformation phenotypes compared to artificial competence induction with plasmids in rich medium etc.) [17], [20]–[23]. This inducible and chromosomally encoded competence system allowed us to look at different aspects of the regulatory network of natural competence under standardized conditions. Based on these new data, we propose a model of how the regulatory network of natural competence functions in V. cholerae.
The goals of our study were to 1) investigate whether natural competence is induced in a whole population under natural or optimized conditions, 2) establish a homogeneous competence-inducing system to investigate the contribution of separate and interconnected regulatory pathways to competence induction, and 3) test whether different competence genes are subject to the same regulatory circuits. We achieved these aims by using transcriptional reporter fusion constructs of representative competence genes. We combined these fluorescent reporter fusions with the following detection methods: epifluorescence microscopy and flow cytometry, which allowed us to visualize the expression of fluorescent reporter genes at the level of single cells and to quantify gene expression accordingly; and fluorescent plate reading, which we used to investigate a plethora of regulatory mutants and regulated genes based on population average fluorescent value measurements.
Results/Discussion
Visualization of competence gene expression upon chitin surface colonization
To better understand whether natural competence of V. cholerae is a developmental program followed by (almost) all members of a population or rather a state, which only a subpopulation acquires, we investigated gene expression at the single cell level. Therefore, we transcriptionally fused the promoter regions of competence genes to those genes encoding fluorescent proteins (FPs). Our choice of FPs was GFP-mut3* [27] and DsRed.T3[DNT] [28], [29] as both of them have been optimized for fluorescence intensity and can be visualized within the same cell (i.e., the excitation/emission spectra are adequately separated). We initially focused on two promoter regions: the upstream region of the pilA-D operon [30], hereafter referred to as the pilA promoter, and the region upstream of comEA. Both of these genes, pilA and comEA, are upregulated on chitin ([11], [31]; Blokesch and Schoolnik, unpublished) and essential for natural transformation to occur [11]. PilA encodes a major pilin, which, as a part of a hypothetical type IV-like pilus [30], is most likely involved in the DNA uptake process. The same involvement in DNA uptake holds true for ComEA, which shows homology to ComEA of Bacillus subtilis [32] as it also contains a helix-hairpin-helix (HhH) motif (pfam12836). HhH motifs have been previously described as short DNA-binding domains that bind DNA in a non-sequence-specific manner [33]. How exactly the DNA uptake, including the involvement of the type IV-like pilus and ComEA, functions is so far unknown for V. cholerae and for other naturally competent bacteria.
With these reporter fusions in hand, we moved on to visualize competence gene expression. We first tested the expression in V. cholerae strains after allowing them to colonize chitin beads. Chitin beads mimic the natural environment of V. cholerae in which the bacteria are often found associated with the chitinous exoskeletons of zooplankton [10]. In contrast to other competence-inducing chitin surfaces, such as crab shell fragments or chitin flakes [11], [19], chitin beads are amenable to light microscopy. As shown in Figure 1, no significant green or red fluorescence was detectable by epifluorescence microscopy for V. cholerae cells grown on chitin beads if the bacteria carried the promoter-less FP reporter plasmid (vector control; panel I). In contrast, bright fluorescence signals were visible when the FP genes were driven by either of the two promoters belonging to pilA or comEA (panel II and reciprocal with respect to the FP-fusions in panel III). From the obtained images, it was apparent that not all of the bacteria within the population were fluorescent and, therefore, expressing the competence genes at detectable levels (Figure 1). Based on the finding that not all bacteria appeared as fluorescent we constructed another reporter as positive control, which consisted of the promoter preceeding the housekeeping gene gyrA (encoding gyrase) transcriptionally fused to gfp. This fusion was cloned onto the same plasmid as the PcomEA-dsRed fusion (see Material and Methods). We tested this reporter strain in our chitin bead colonization assay (Figure 1, panel IV). In contrast to dsRed being driven by the comEA promoter the gyrA promoter led to detectable gfp expression in a significantly larger fraction of cells. The same expression pattern as for gyrA was observable for three additional transcriptional reporter fusions containing the promoter region of recA, clpX, and ftsH, respectively (Figure S1). We included these reporter strains in our study as the expression of the housekeeping gene gyrA is most likely controlled by DNA supercoiling as it was demonstrated for E. coli [34] and/or might be cell cycle-dependent as shown for Caulobacter crescentus [35]. The rational for choosing recA, clpX, and ftsH as additional positive controls was based on the fact that other researchers have already used these genes to normalize quantitative RT-PCR expression data. Furthermore, expression of none of these genes was significantly changed in microarray expression studies using different V. cholerae mutant strains ([11], [36] and Blokesch and Schoolnik, unpublished) or using different growth conditions (e.g. comparing rabbit ileal loop grown cells versus exponential in vitro cultures of V. cholerae [37]).
Fig. 1. Visualization of competence gene expression on chitin surfaces. 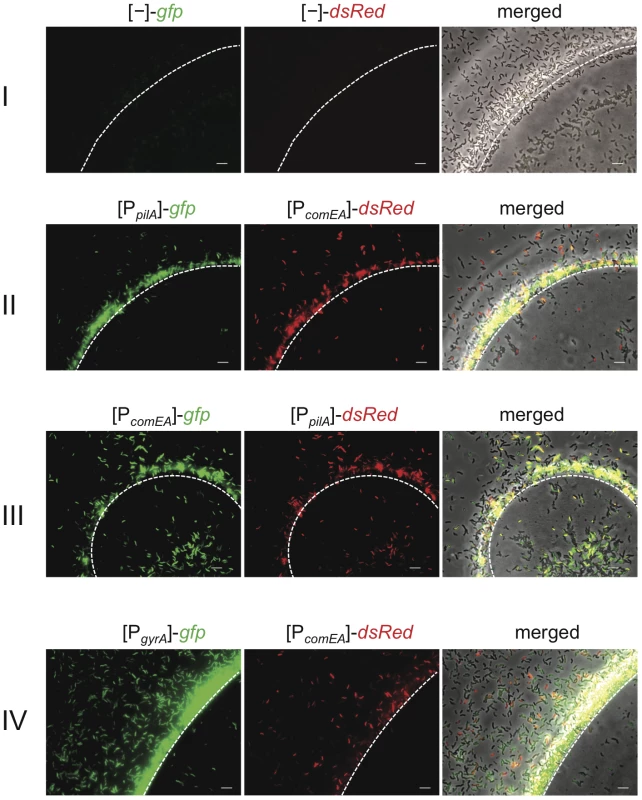
V. cholerae cells were grown on chitin beads as previously described [23]. The white dashed line notes the edge of the chitin surface. The bacteria carried diverse transcriptional FP reporter fusions. I: vector control containing promoter-less gfp and dsRed; II: plasmid containing gfp driven by the pilA promoter and dsRed downstream of the comEA promoter region; III: swapped reporter genes in comparison to II; IV: plasmid containing gfp driven by the gyrA promoter and dsRed driven by the comEA promoter. Bacteria were grown statically for 48 h before pictures were taken. The order of the images here and in the following figures is (from left to right): green channel (GFP), red channel (dsRed), and merged image composed of a phase contrast image overlaid by both fluorescence channel images. Scale bar = 5 µm. We considered three different reasons for the finding that competence genes are only expressed at detectable levels in a fraction of the population: 1) competence gene expression in V. cholerae is a bistable phenomenon, which is similar to B. subtilis [25]; 2) the environment around the chitinous surface is heterogeneous and, thus, does not lead to competence induction in all cells; and 3) the fluorescence signal in cells that appear as non-induced for pilA and comEA expression is too weak to be detected with our epifluorescence microscopy settings.
To follow up on these three possibilities, we aimed at differentiating between the existence of an intrinsic bistable switch versus the idea of a heterogeneous expression pattern due to heterogeneous conditions and to concomitantly judge whether the seemingly uninduced cells observed on the chitinous surfaces (Figure 1) were the result of experimental limitations in our system.
Population-wide expression of competence genes under homogeneous conditions
First, we wanted to test if we would observe a bistable competence gene expression pattern under homogeneous growth conditions. Therefore, we changed the chitin substrate from chitin beads (an insoluble GlcNAc polymer) to soluble hexa-N-acetylchitohexaose (from here on referred to as GlcNAc6). This chitin oligomer has been used before to induce natural competence in V. cholerae [11], [23]. As a control, we grew the same V. cholerae reporter strains under identical minimal medium conditions (defined artificial seawater medium, DASW; [11]) but changed the main carbon source from GlcNAc6 to the non-competence inducing chitin monomer N-acetylglucosamine (GlcNAc). All strains were grown to the same optical density before being either visualized by epifluorescence microscopy (Figure S2) or quantified with respect to their fluorescence intensity using flow cytometry (Figure 2). As shown in Figure S2A and S2B, we did not detect significant fluorescent signals for the promoterless reporter control using microscopy, which is in accordance with the low levels of fluorescent signals measured by flow cytometry (Figure 2A). Thus, the fluorescence intensity of these bacteria (panel A, flow cytometry graphs) was considered background. For the strains grown with GlcNAc, FP expression driven by the comEA promoter was not detectable using microscopy (Figure S2C, S2E, S2G including the respective image analysis) and quantified as extremely low fluorescent signals using flow cytometry (Figure 2B–2D upper row). However, weak pilA promoter-driven gfp expression in the presence of GlcNAc was observed after an extended exposure time (Figure S2C). We confirmed this basal pilA promoter-driven gfp expression in the presence of GlcNAc by flow cytometry (mean FU = 7.2×102; Figure 2B, upper row). Swapping the FP reporter gene behind the pilA promoter from gfp to dsRed resulted in undetectable red fluorescence using our epifluorescence microscopy settings (Figure S2E); however, an increased pilA promoter-driven expression of dsRed compared to the promoterless reporter plasmid control (Figure 2A) was detectable by flow cytometry (Figure 2C, upper row), confirming the low level of pilA expression under non-competence inducing conditions.
Fig. 2. Competence genes are expressed by the majority of cells under homogeneous competence-inducing conditions. 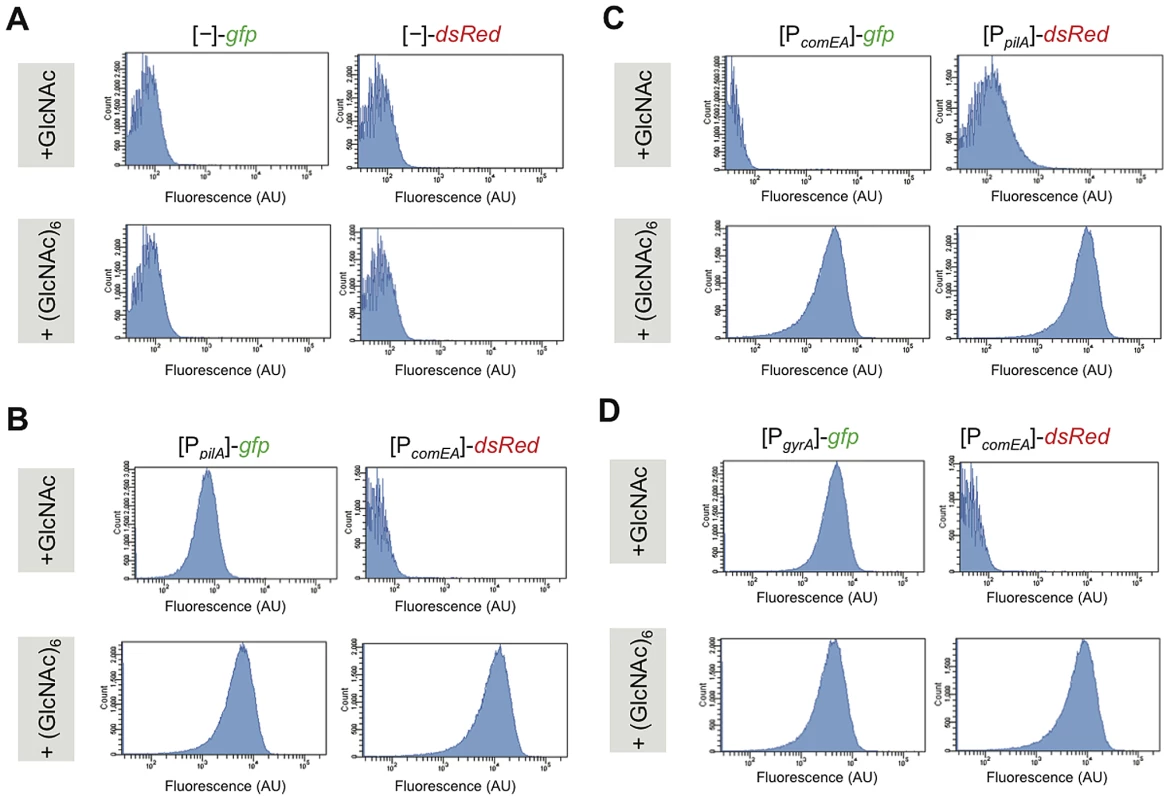
V. cholerae strains were grown aerobically in defined artificial seawater medium with the addition of either N-acetylglucosamine (GlcNAc) or hexa-N-acetylchitohexaose (GlcNAc)6 as sole carbon source. The latter is known as a potent inducer of natural competence/transformation [11]. Competence gene expression was quantified for fluorescence intensities using flow cytometry. Reporter fusions are indicated above each panel. Panel A: promoter-less gfp and dsRed reporter; panel B: [PpilA]-gfp/[PcomEA]-dsRed; and panel C: [PcomEA]-gfp and [PpilA]-dsRed. Panel D: [PgyrA]-gfp/[PcomEA]-dsRed. The flow cytometry graphs indicate the number of cell counts on the y-axis and the fluorescence signal intensity (as arbitrary units, AU) on the x-axis. In cells grown under competence-inducing conditions (e.g. in the presence of GlcNAc6), the reporter strains displayed strong pilA and comEA promoter-driven fluorescent signals (Figure S2D, S2F, S2H). This was the case for the majority of the cells and only a minority appeared as non-fluorescent under these conditions. There was a distribution of fluorescence intensities as depicted in the flow cytometry graphs (Figure 2B–2D, lower row) but the distribution was not bimodal. To further investigate whether the non-fluorescent-appearing cells in the microscopy images (Figure S2) were meaningful with respect to competence expression, we again investigated the behavior of the housekeeping gene reporter strains under these homogenous competence-inducing conditions (for gyrA see Figure 2D and Figure S2G and S2H; or for recA, clpX, and ftsH see Figure S3). The same expression pattern as for the competence genes was observed for these strains with a minority of cells not displaying any detectable fluorescence using our epifluorescence microscopy and image display settings. Thus, and also based on the image analysis (Figure S2) and on the flow cytometry measurements (Figure 2), this minority of non-fluorescent-appearing bacteria probably corresponds to cells, which fluoresce at lower levels (mostly with a good correlation between both FPs; Figure S2).
An issue that we had to consider was the fact that our FP reporter constructs were plasmid-encoded. Indeed, plasmid copy numbers can change in V. cholerae according to growth rate [38]. However, in our experiments we mainly looked at different strains but under similar growth conditions (and the biological replicates were highly reproducible; Figure S4). Furthermore, a recent study by Silander et al. provided evidence that plasmid-based systems are useful to study gene expression in bacteria; they observed that both the mean and the variation of expression correlated well between both settings [39].
A chromosomally encoded inducible competence system allows quantification of gene expression
Based on the data described above, we concluded that bistability of competence gene expression is unlikely in a population of V. cholerae cells under homogeneous competence-inducing conditions. Therefore, it seemed feasible to further investigate the regulatory circuit at a population-wide level given that homogeneous, competence-inducing growth conditions were provided. Unfortunately, GlcNAc6, a competence-inducer, has recently been discontinued by the Seikagaku Corporation, and multiple and large-scale experiments using this commercially available compound are extremely costly. Furthermore, shorter GlcNAc oligomers such as GlcNAc2 often result in large variations with respect to transformation frequencies (M. Blokesch, unpublished), which is most likely due to GlcNAc monomer impurities within the preparation that exert catabolite repression on natural transformation [23]. Therefore, we thought of establishing a chitin-independent, competence-inducing system. One possibility was to artificially overexpress the major regulator of transformation, TfoX, from a plasmid as previously performed [11], [17], [22], [23]. However, we observed major disadvantages using this method. First, the morphology of some of the cells changed towards a filamentous form after competence induction due to the plasmid-maintaining antibiotic ampicillin (Blokesch, unpublished). Second, TfoX overexpression from a multi-copy plasmid resulted in the induction of heat shock proteins and chaperones [11], which might be indicative of stress conditions. Indeed, the toxicity of TfoX overexpression has been previously described for Escherichia coli and Haemophilus influenzae [40], [41]. Third, we wanted to avoid working with V. cholerae cells containing two different plasmids within the same cell (tfoX and FP reporter fusions-carrying). Therefore, we constructed a chromosomally encoded competence induction system, which is based on inducible low-level TfoX production. The system was composed of tfoX under the control of an arabinose-inducible promoter (PBAD) and the gene encoding AraC, which act as a repressor or initiator of gene expression in the absence or presence of L-arabinose, respectively [42]. Both of these elements were cloned into a mini-Tn7 transposon [43], which integrates into the large chromosome of V. cholerae (later referred to as TntfoX). We transferred this transposon by triparental mating into the V. cholerae wild type strain A1552 and tested the respective strain for natural transformability in LB medium (Figure 3). By adding a low amount of L-arabinose (0.02%), we obtained transformation frequencies that were two orders of magnitude higher than what has been described for overproduced tfoX in V. cholerae [11] (Figure 3A). Furthermore, the transformation frequency (2.1×10−4) was in the same range as the frequencies we usually obtain using optimized chitin-inducing conditions (3.1×10−4; [19]). We were unable to detect any naturally transformed colony-forming units in the inducer-free control culture (shown for the wild type in Figure 3A but likewise tested in all other strains). We also examined the abundance of TfoX at the protein level. To do so we grew the V. cholerae strain A1552-TntfoX in LB medium in the absence or presence of different L-arabinose concentrations followed by western blot analysis of cellular proteins using antibodies against TfoX (Figure S5). In parallel we grew a strain containing inducible tfoX on a plasmid similar to the tfoX-overexpression system described earlier [11]. As shown in Figure S5 we observed a major difference in TfoX protein levels comparing the previous [11] and current experimental setup as indicated by the two arrows. This reassured us that this system was not heavily overproducing TfoX and was therefore adequate for further analysis to establish the genetic interactions downstream of TfoX.
Fig. 3. Artificial induction of natural transformation by expression of the competence regulatory gene tfoX in cis. 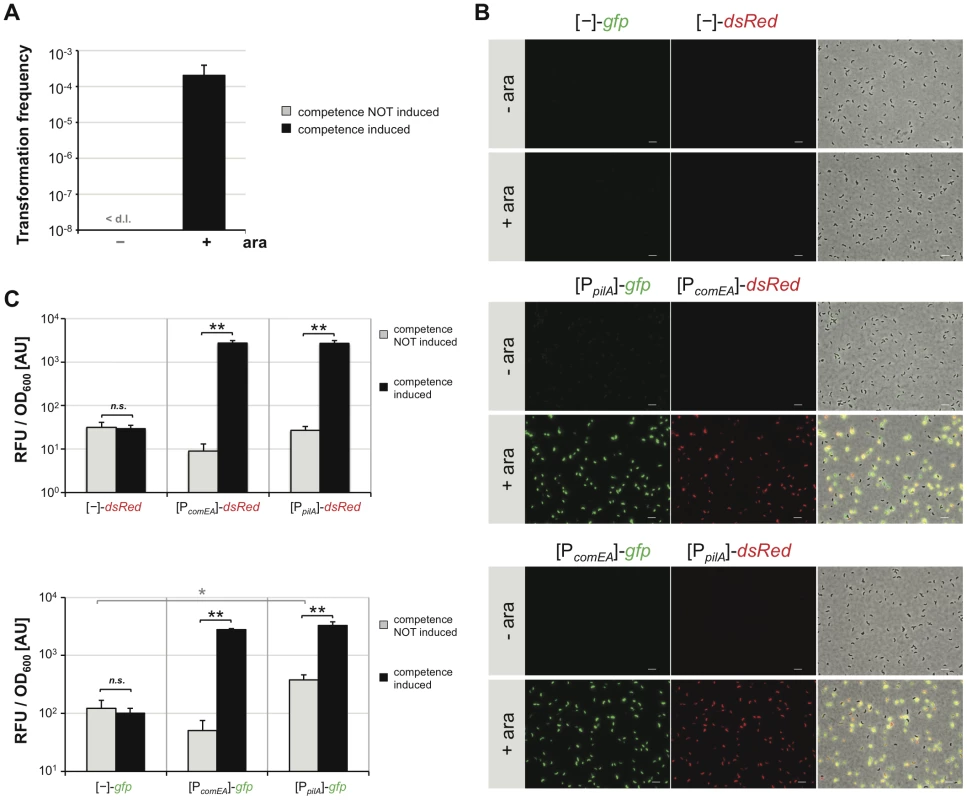
V. cholerae cells were grown in rich medium in the absence (−) or presence (+) of the artificial inducer arabinose (0.02%). Cells were either tested for natural transformability (panel A) or competence gene promoter activity based on FP reporters (panels B and C). Panel A: Transformation frequencies are given on the y-axis for competence-uninduced (−) and competence-induced (+) bacteria. <d.l. = below detection limit. Panel B and C: V. cholerae cells harboring the different transcriptional reporter fusions were grown without or with competence induction. Bacteria were either visualized by epifluorescence microscopy (panel B; image arrangements as in Figure 1; Scale bar = 5 µm) or measured with respect to relative fluorescence units (RFU) and optical density at 600 nm (panel C). Panel C: RFU per OD600 values are given on the y-axis. All experiments in Figure 3 were repeated at least three independent times. Error bars reflect standard deviations. Statistics were applied using the Student's t test. * P<0.05, ** P<0.01, n.s. = not significant. We first wanted to visualize competence gene expression in this chitin-independent system. We transferred the respective FP reporter fusion constructs into a wild type V. cholerae strain carrying the chromosomally encoded tfoX construct (A1552-TntfoX) and visualized FP gene expression by epifluorescence microscopy (Figure 3B). As can be appreciated from the images in Figure 3B (middle and lower part), the pilA - and comEA promoter-driven expression pattern of the FP reporters under such chitin-independent, competence-inducing conditions mirrored what we observed under the GlcNAc6-mediated induction of competence (Figure S2). As described above for the chitin-dependent experiment, only a minority of cells did not display any detectable fluorescence using this microscopy technique, which was also the case for gyrA promoter-driven FP reporter expression (data not shown). The fluorescent signal was below the detection limit in cells grown in the absence of inducer (Figure 3B, -ara) or in cells harboring the promoter-less plasmid as a control (Figure 3B, upper two rows).
The fluorescent signal was quantified using a 96-well plate reader (Figure 3C). A statistically significant increase in fluorescence intensity was observed upon induction of competence for all promoter-driven FP reporter fusion constructs (Figure 3C, middle and right columns). No significant difference in fluorescence signals between competence-uninduced and competence-induced conditions was observed for the promoter-less FP reporter control (Figure 3C). In agreement with the chitin data described above, we detected a statistically significant increase in pilA-driven gfp expression compared to the promoter-less construct even in the absence of inducer. Therefore, we conclude that this basal expression of pilA is TfoX-independent.
Induction of natural competence is dependent on the cAMP level within cells
We then moved on to investigate the regulatory network of natural competence in V. cholerae in further detail. The first assay aimed at testing whether carbon catabolite repression (CCR) plays a role in this chitin-independent setup. Carbon catabolite repression occurs if preferred phosphoenolpyruvate∶carbohydrate phosphotransferase system (PTS)-transported sugars are abundant. However, in their absence the PTS systems are unsaturated and indirectly lead to the activation of the enzyme adenylate cyclase, which subsequently synthesizes cAMP within the cell (for review on CCR see [44]). We were mostly interested in V. cholerae strains that are impaired in this synthesis or in the degradation of cAMP. The concentration of cAMP within cells is accomplished by interplay between adenylate cyclase (CyaA) and cAMP-degrading phosphodiesterases (CpdA). A recent study by Kim et al. demonstrated the importance of CpdA in balancing the intracellular cAMP level in Vibrio vulnificus [45]. As the cpdA gene of V. cholerae is located at the same chromosomal locus (as analyzed using SynTView, a synteny viewer developed by the Genomic and Genetic Department of Institute Pasteur, Paris) and cpdA/CpdA display 64%/68% identity (76%/82% similarity) at the DNA and protein levels, respectively, the functionality of the protein is most likely identical in both organisms. To disrupt the equilibrium between cAMP production and degradation, the cpdA gene in V. cholerae was deleted (Table 1). Although the production of cAMP does not change in this mutant, cAMP degradation should be impaired, resulting in the accumulation of cAMP within the cell. We tested this strain in a chitin surface colonization assay [23] and observed a hyper-colonization phenotype consistent with increased intracellular cAMP levels (data not shown). More importantly, we transferred the TntfoX transposon into this V. cholerae strain as well as a strain lacking adenylate cyclase (ΔcyaA) and tested both strains with respect to natural transformability (Table 2) and pilA/comEA promoter-driven FP gene expression (Figure S6). The results confirmed part of what we had previously demonstrated on chitin surfaces [23], namely that adenylate cyclase is essential for natural transformation even under rich culture medium conditions. However, in this study we extended this knowledge by showing that a statistically significant increase in natural transformability occurred in the newly constructed cpdA mutant compared to the wild type parental strain (Table 2). With respect to competence gene induction, pilA and comEA promoter-driven FP gene expression was abolished in the absence of cAMP (Figure S6). This was in contrast to the fluorescent signal measured for the cpdA mutant, that is, both the pilA and the comEA promoter efficiently drove FP gene expression in this genetic background upon competence induction (Figure S6). These data highlight the necessity of cAMP for competence gene expression even when competence induction is uncoupled from chitin surface colonization and chitin degradation (e.g., from metabolism of carbon sources).
Tab. 1. Bacterial strains and plasmids. 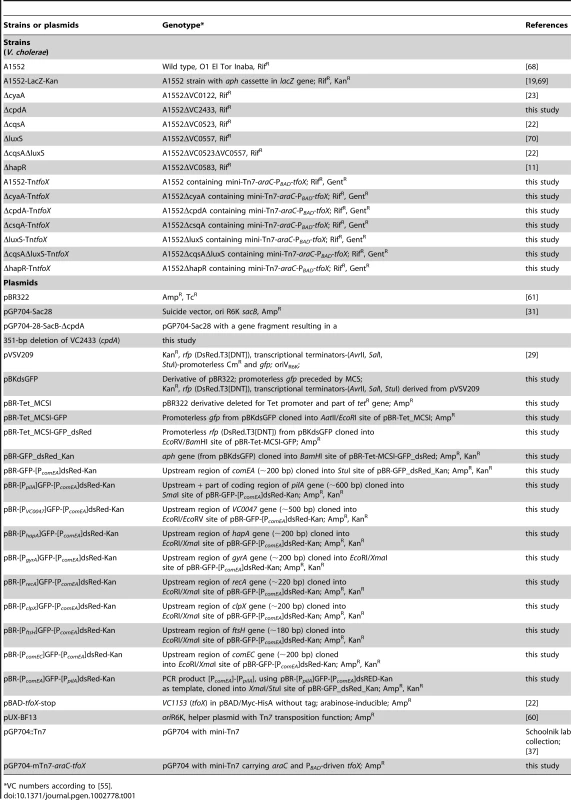
VC numbers according to [55]. Tab. 2. Natural transformation is dependent on cAMP, which is produced by adenylate cyclase (CyaA) and degraded by cAMP-degrading phosphodiesterase (CpdA). 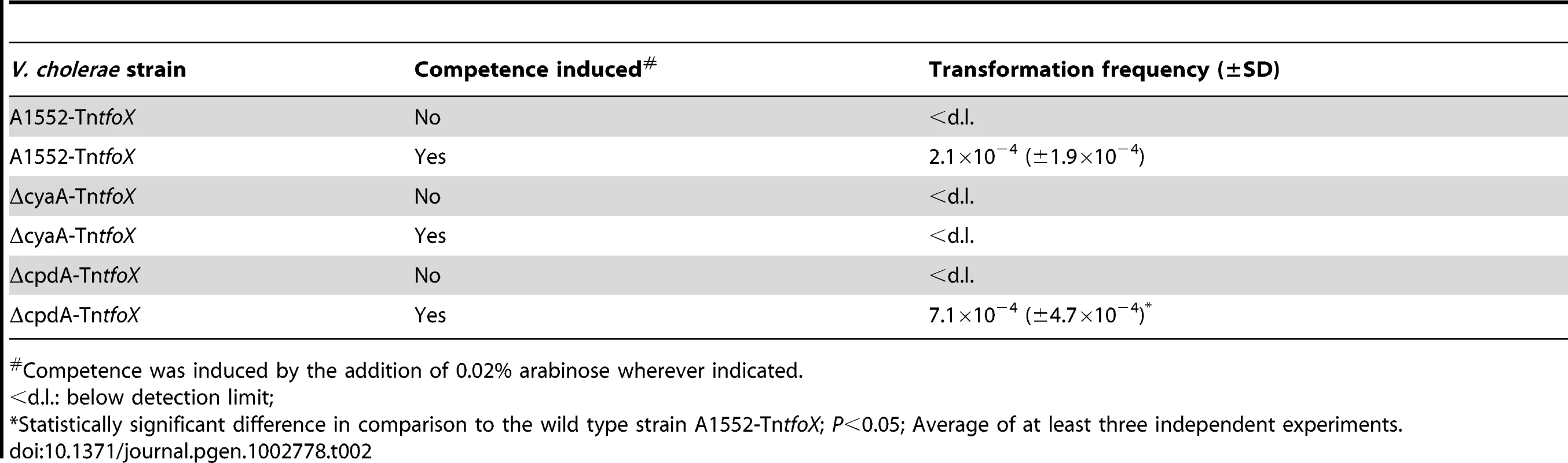
Competence was induced by the addition of 0.02% arabinose wherever indicated. Quorum-sensing only regulates a subset of competence genes
The next question we wanted to address was with respect to QS and the involved autoinducer molecules. We recently showed that the species-specific cholera autoinducer 1 (CAI-1; [46], [47]) plays a major role in natural competence for transformation and suggested that CAI-1 could be considered a competence pheromone [22]. We showed that the absence of the non-species-specific autoinducer 2 (AI-2; [48]) had no statistically significant effect on natural transformation on chitin surfaces, whereas strains devoid of CAI-1 synthesis were rarely transformable, and, even then, only at very low transformation frequencies [22]. However, as described above, chitin surfaces appear to be a rather heterogeneous environment, and cells might not all encounter the same autoinducer concentration in time and space, making chitin surface experiments difficult to conclusively judge the involvement of QS in the regulatory circuit of V. cholerae. Therefore, we investigated the role of QS in natural competence and transformation using our homogeneous competence-inducing system (Figure 4). We constructed TntfoX-containing V. cholerae deletion strains, which were devoid of either or both of the autoinducer-synthesizing enzymes CqsA and LuxS, or which lacked the gene encoding the major regulator of QS, HapR. These strains were grown in LB medium in the presence or absence of the competence-inducer arabinose and scored for natural transformability or competence gene promoter-driven FP expression (Figure 4). In the absence of inducer, the transformation frequency was consistently below the level of detection in all strains (Figure 4A). In the absence of the AI-2-producing enzyme LuxS, only a minor and statistically not significant decrease in transformation frequency upon competence induction was detectable when compared to the wild type parental strain (Figure 4A). More importantly, the dependency on CAI-1 was even enhanced when compared to our previous study on chitin surfaces, in that a deletion in the gene cqsA completely abolished natural transformation. This was also the case in strains devoid of both autoinducer synthases (CqsA and LuxS) and the major regulator of QS, HapR (Figure 4A). We argue that few occasionally detectable transformants in a CAI-1 negative mutant in our previous study on chitin flake surfaces [22] were the result of the heterogeneity of the chitin surface environment in which at least three components are not evenly distributed: autoinducers, transforming DNA and nuclease. However, under homogeneous conditions as tested here, a full dependency on CAI-1 is apparent.
Fig. 4. Quorum sensing only regulates a subset of competence genes. 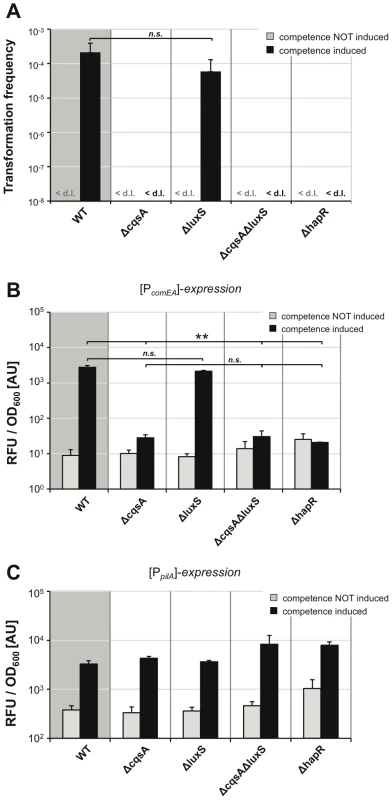
The tfoX-expression construct was transferred onto the chromosome of mutant V. cholerae strains, which were defective in the quorum-sensing circuit. Strains were grown in LB medium with or without 0.02% arabinose and tested for natural transformability (panel A), comEA (panel B), and pilA promoter-driven FP expression (panel C) as described for Figure 3. Experiments were repeated at least three times. Statistically significant differences were calculated using the Student's t test. * P<0.05, ** P<0.01, n.s. = not significant. <d.l. = below detection limit. We then visualized and quantified competence gene expression in these strains using the above-described FP report fusions (Figure 4B, 4C). The data for comEA promoter-driven FP expression (Figure 4B) mirrored the data of the transformation assay (Figure 4A). Under non-competence-inducing conditions, only the background fluorescence was measurable (in the range of the vector control shown in Figure 3). There was no statistically significant difference between the fluorescence signal detected in the wild type strain and the signal derived from the luxS-deficient strain upon competence-inducing conditions. A highly significant reduction of comEA promoter-driven FP gene expression was detected in the cqsA, cqsA/luxS and hapR negative strains. We also measured for the first time pilA promoter-driven FP expression in the different QS mutants and thus in the presence or absence of the two autoinducers. We observed that the expression pattern looked completely different from the comEA data; though the fluorescent signal increased upon competence induction, the fluorescence units were in the same range for all strains tested (Figure 4C). Therefore, we conclude that, in contrast to comEA, pilA is not subject to QS-dependent regulation.
Taken together, we provide evidence that CAI-1 is essential for comEA expression and natural transformation under homogeneous competence-inducing conditions. These data are in slight contrast to another study where a gradual decrease in comEA expression and natural transformation from a wild type V. cholerae strain towards an AI-2 - and CAI-1-deficient strain, respectively, was displayed [21]. The authors of this study concluded that not only CAI-1 but also AI-2 contributes to natural transformation. We believe that the discrepancy between studies (the study described here, [21] and [22]) could reflect the different V. cholerae O1 El Tor strains used in both studies (A1552 here and in [22], versus C6707str in [21]). This hypothesis is in excellent agreement with a recent finding by Fong and Yildiz [36], who showed that the cAMP-CRP-mediated negative regulation of the biofilm regulatory gene vpsR only occurred in three out of four tested V. cholerae strains, namely strains A1552, N16961 and MO10. In contrast to this result, V. cholerae strain C6706 displayed no such regulation [36], highlighting the fact that different regulatory circuits exist in these V. cholerae strains.
The amount of HapR within cells dictates comEA expression and nuclease repression
The involvement of QS in natural transformation has previously been demonstrated by elucidating a role for HapR in the repression of a gene encoding the extracellular nuclease Dns [17]. This conclusion was mainly based on comparing a wild type V. cholerae strain to a ΔhapR mutant with respect to dns gene expression or nuclease activity. However, a direct correlation between HapR protein levels within the cell, dns repression, and comEA induction has never been demonstrated. We addressed this missing information by performing western blot analysis of non-competence-induced and competence-induced cells to detect the HapR protein within different QS mutant strains (Figure 5A). Whereas the amount of HapR did not differ between tfoX-induced and tfoX-uninduced cells significant differences between the tested strains were observable (Figure 5A). That is, whereas the HapR level was only slightly reduced in an AI-2-deficient strain (ΔluxS), HapR was almost undetectable in a strain lacking CAI-1 (ΔcqsA) (Figure 5A). Better detection was only possible upon overexposure of the film and such an overexposure did not reveal any HapR protein in the dual autoinducer mutant strain (ΔcqsAΔluxS) or the hapR negative control (Figure S7). This HapR protein pattern directly reflects the comEA-promoter driven FP expression data quantified in Figure 4B and also indicates that CAI-1 is the stronger autoinducer compared to AI-2 in V. cholerae strain A1552 (consistent with what was shown for strain C6707 using a heterologous read-out [49]). We also tested the expression of another QS-dependent but competence-independent gene, hapA, by transcriptionally fusing the hapA promoter to gfp. The hapA gene encodes a hemagglutinin protease (HA protease) and is positively regulated by HapR [50]. When we compared the comEA expression data (Figure 4B) to those of hapA (Figure 5B) a very similar expression pattern was observable under competence inducing conditions. We hypothesize that the low amount of HapR present in the cqsA mutant (Figure 5A and Figure S7) is not sufficient to activate expression of either comEA or hapA. A potential reason for this might be that HapR displays only a weak affinity for these promoters, which is consistent with a LuxR promoter affinity model described for Vibrio harveyi [51].
Fig. 5. Correlation among HapR protein levels, hapA gene expression, and the nuclease Dns. 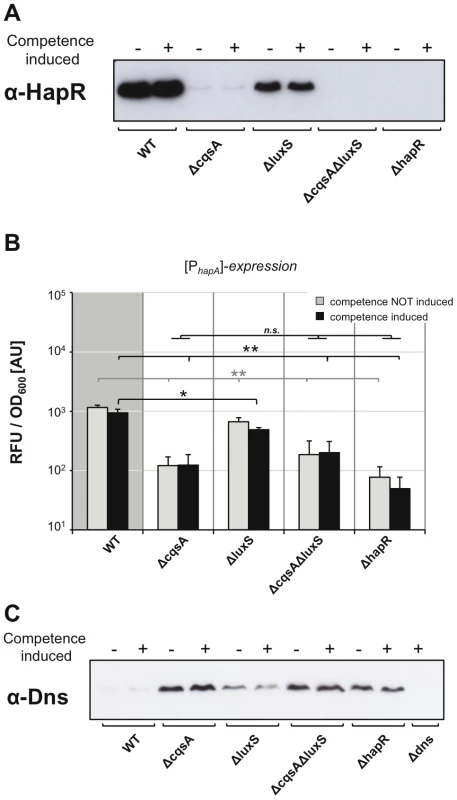
Panel A and C: Proteins of the indicated strains, each containing artificially inducible tfoX on the chromosome, were separated by SDS-PAGE. After blotting, the relative abundance of proteins HapR (panel A) or Dns (panel C) were determined by detection with protein-specific antibodies. For each sample 6 µg (panel A) and 12 µg total protein (panel C), respectively, were applied per lane. Strains were tested under non-competence-inducing and competence-inducing conditions as indicated above each image. Panel B: HapR-dependent expression of hapA promoter-driven gene expression was quantified as described in Figure 3 for comEA/pilA. The growth conditions were as described for Figure 4. Averages of three independent experiments are indicated. Error bars indicate standard deviations. Statistically significant differences were calculated using the Student's t test. * P<0.05, ** P<0.01, n.s. = not significant. We also tested the impact of the HapR level on the protein amount of the nuclease Dns (Figure 5C) and observed an inverse correlation: Dns repression only occurred in those strains in which we detected high levels of HapR protein (e.g. WT and ΔluxS in Figure 5A). This is in good agreement with the absence of any transformants in a cqsA mutant (Figure 3A), as the abundance of the nuclease in this strain would avoid uptake of intact DNA. This is the first time that a direct correlation between HapR protein levels, nuclease levels and comEA expression has been shown, which is the critical link between QS and natural competence/transformation.
Investigation of other tfoX and QS–regulated competence genes
Finally, we were curious to determine whether this chitin-independent, competence-inducing system would allow us to investigate other genes that are potentially involved in natural transformation. We initially focused on two potential promoter regions belonging to genes VC0047 and comEC. The first promoter precedes VC0047, which is part of a four-gene operon (VC0047-50). The gene VC0048 in this cluster encodes DprA, which is essential for natural transformation of V. cholerae [22]. The function of DprA in Streptococcus pneumoniae and probably also in V. cholerae is to protect the incoming single-stranded DNA from degradation and to convey the DNA to RecA-mediated recombination [52]. As shown in Figure 6A, a tfoX-dependent expression pattern was detectable in our VC0047 promoter-driven gfp reporter strain.
Fig. 6. TfoX drives expression of QS–dependent and QS–independent competence genes. 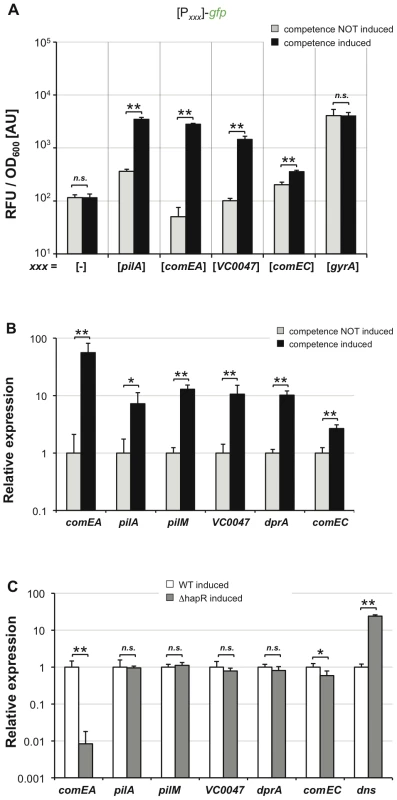
V. cholerae wild type strain containing artificially inducible tfoX in cis was tested for the expression of different competence genes. Panel A: Different transcriptional FP reporter fusions were tested for TfoX-dependent induction. These fusions were composed of the potential promoter region of the respective competence gene(s) (x-axis) fused to gfp. The housekeeping gene gyrA was tested as control. Relative fluorescence per OD600 unit is given on the y-axis. Panel B and C: qRT-PCR data comparing the relative expression of the indicated genes in a wild type strain under competence non-inducing and competence-inducing conditions (panel B). In panel C both the wild type strain and the hapR mutant were tested for competence gene expression under tfoX-expressing conditions. All panels depict averages of at least three independent experiments and error bars indicate standard deviations. Statistically significant differences were determined using Student's t tests. * P<0.05, ** P<0.01, n.s. = not significant. Another gene that we have previously shown to be essential for transformation of V. cholerae is comEC ([22]; annotated as inner membrane transporter). As so far nothing was known about its regulation we sought to investigate whether comEC is regulated in a TfoX-dependent manner. Using a comEC promoter-driven gfp reporter strain we were able to measure a slight but statistically significant increase in comEC upon competence induction (Figure 6A). This is the first time that comEC has been shown to belong to the chitin-/TfoX-regulon in V. cholerae and as such being co-regulated with other competence genes.
We were also interested in the regulation of the pilM-Q operon as pilQ is also required for natural transformation [11], [22]. Thus, we fused the pilM promoter region to gfp and determined FP expression under competence inducing conditions. Unfortunately, the signal intensity was too low to allow us to unambiguously judge pilM promoter-driven expression. To overcome this obstacle we established quantitative RT-PCR in our laboratory to further monitor competence gene expression using our chitin-independent system. We first compared competence-uninduced to competence-induced cells with respect to expression of comEA, pilA, pilM, VC0047, dprA and comEC. As indicated in Figure 6B all of these genes were significantly induced upon competence-induction. This again confirmed the TfoX-dependent regulation of comEC shown in Figure 6A, which was missed in previous chitin/TfoX-dependent expression studies [11], [31]. We suggest that this was the case, as the change in expression did not pass the significance filter in these microarray expression studies. Indeed, the fold-difference for comEC expression upon competence induction was only 2.7 (Figure 6B). Interestingly, this 2.7-fold increase in comEC expression is in the same range as what we observed using the comEC FP reporter construct (1.8-fold change; Figure 6A), though the latter system was plasmid-based.
Finally, we wanted to test whether any of these other competence genes is also regulated in a QS-dependent manner. We therefore tested and compared the expression of these genes under competence-inducing conditions in a wild type and hapR negative strain, respectively (Figure 6C). Apart from comEA and dns, only comEC turned out as also HapR-dependent (Figure 6C, P = 0.0157). This is in nice agreement with our regulatory model in which the fate of the surrounding DNA is determined by QS (Figure 7 and conclusion below).
Fig. 7. Model of the regulatory network of natural competence and transformation of V. cholerae. 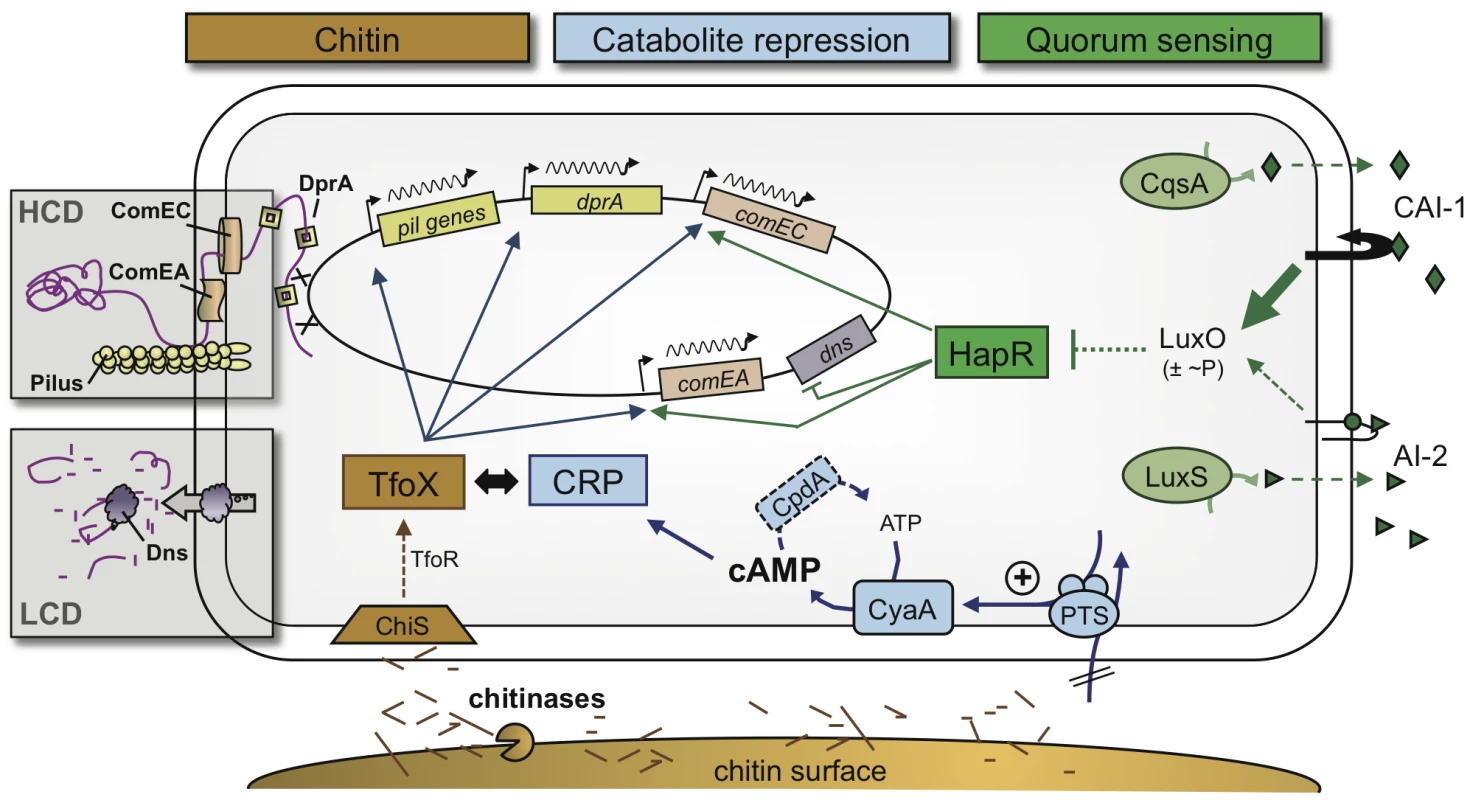
At least three extracellular and intracellular signaling molecules must be present to allow natural transformation to occur in V. cholerae. 1) Chitin degradation products such as chitin oligomers, which lead to the induction of the sRNA TfoR and the main regulator of transformation TfoX (chitin pathway shown in brown). 2) The secondary messenger cAMP, which has to accumulate within cells (CCR pathway shown in blue). 3) Extracellular autoinducers, with an emphasis on the stronger autoinducer CAI-1, which feed into the quorum-sensing circuit (shown in green). Whereas chitin- and TfoX-dependent induction and the requirement for cAMP and CRP are universal for all, so far investigated, competence genes, the QS-dependent circuit regulates only a subset of those, such as comEA and comEC. Therefore, QS acts as a switch in gene expression and is responsible for the final fate of the surrounding DNA (boxed areas). At a low cell density (LCD), the DNA (shown in purple) is degraded by the nuclease Dns. As a consequence, the cells are non-transformable. At a high cell density (HCD) and, therefore, high abundance of the autoinducer CAI-1, the nuclease gene dns is transcriptionally repressed, whereas comEA and comEC are activated. ComEA as well as ComEC then contribute to the DNA uptake process, probably due to their ability to shuffle the DNA through the periplasmic space and the inner membrane, respectively. Concluding remarks and proposed model of the regulatory network
In this study, we analyzed the regulatory network of chitin-induced natural competence and transformation in V. cholerae in its full complexity (Figure 7). Our results suggest that under homogeneous conditions bistability is unlikely for V. cholerae. However, the conditions might be less homogeneous in time and space around biotic surfaces (e.g., different concentrations of autoinducers and PTS sugars, which interfere with natural competence via QS and CCR, respectively). Such “environmental heterogeneity” might foster a non-synchronized response by the chitin-associated bacteria. We hypothesize that due to such a heterogeneity competence gene expression appeared absent in a subpopulation of bacteria grown on chitin beads, whereas housekeeping genes were almost uniformly expressed throughout the population (Figure 1 and Figure S1).
Based on the results obtained in this study and combined with the knowledge from earlier studies by us and others [11], [17], [22], [23], we developed a model for the regulatory network of natural competence and transformation (Figure 7). The model predicts the interplay between three pathways for the initiation of competence: chitin sensing followed by TfoX activation, CCR, and QS. The first pathway, the dependency on a chitin surface or on chitin oligomers (e.g., GlcNAc2–6) for competence induction, was discovered as these compounds lead to an upregulation of potential competence genes. Within these competence genes were the so-called pil genes, which encode for a type IV pilus that is potentially involved in the DNA uptake process [22], [31] (Figure 7). Chitin also led to an induction of the gene encoding the main regulator of natural transformation, TfoX [11], [31]. This finding is supported by recent studies that show that chitin oligomers (GlcNAc>2) lead to an increase of tfoX transcription but also to its enhanced translation [18]. The latter effect could be explained after the discovery of a chitin-induced small RNA TfoR, which activates translation of TfoX mRNA and, therefore, contributes to the induction of natural competence [20]. Attempts from our group to look at transcriptional reporter fusions between tfoX and gfp were unsuccessful, which was most likely due to the low activity of the tfoX promoter (M. Lo Scrudato, M. Grasser, M. Blokesch, unpublished). The flow of information in this part of the regulatory circuit (e.g., chitin sensing) is therefore as follows (Figure 7): the presence of chitin is sensed by the chitin sensor ChiS, due to chitinase-released GlcNAc di-/oligomers [31], [53], and the signal is then transferred via TfoR towards the production of TfoX.
As this chitin-dependent pathway is well established, we excluded it in the second part of this study and designed a chitin-independent, competence-inducing system, which is based on artifical tfoX expression (though not overproduction). With this chitin-independent system, we obtained transformants at comparable frequencies to our optimized chitin-induced transformation protocol [19] and at 10 - to 10,000-fold higher frequencies than recent studies by other groups [18], [20], [21]. As the induction occurs under homogeneous conditions in this system, it allowed us to us to better investigate the two other pathways involved in natural competence regulation, CCR and QS.
Based on experimental data showing that glucose interferes with chitin-induced transformation, Meibom et al. suggested that catabolite repression might be involved in the competence phenotype [11]. This hypothesis was recently extended as we showed that competing PTS-dependent carbon sources indeed repress natural transformation [23]. Such sugars are known to play a role in the intracellular accumulation of the secondary messenger cAMP, which, together with the cAMP receptor protein CRP, contributes to chitin surface colonization, chitin degradation and natural competence [23]. In this current study, we circumvented the problem that CCR mutants are often impaired for colonization and growth on chitin as a sole carbon source [23] by uncoupling natural competence-induction from chitin. This allowed us to better understand the dependency on cAMP for competence gene expression. Indeed we observed a change in natural transformability upon creating an imbalance in the intracellular cAMP pool, either by inhibiting cAMP production or, alternatively, by avoiding cAMP degradation. The latter effect was accomplished by deleting the gene encoding the cAMP phosphodiesterase, an enzyme that has never been studied in V. cholerae before. We also confirmed that the TfoX-induced expression of pilA and comEA requires cAMP, which is consistent with the idea proposed, but not yet unequivocally demonstrated, for Haemophilus influenzae that TfoX and cAMP-CRP act in concert to induce competence genes [40], [41], [54] (Figure 7).
The third pathway that participates in natural competence induction is quorum-sensing (QS). The involvement of QS in competence initiation was initially speculated based on different facts. First, the first sequenced strain of V. cholerae N16961 [55] was non-transformable [11]. In this strain, the indigenous hapR gene contains a frameshift mutation that renders it non-functional [56]. This deficiency could be overcome by providing a functional copy of hapR back in cis [11]. The second line of evidence for the involvement of QS in natural competence and transformation came from the fact that cells were more efficiently transformable after longer growth on chitin surfaces, which is equivalent to higher cell densities [11]. However, such a finding could also be explained by elevated intracellular cAMP levels after extended growth on chitin. The involvement of QS in natural transformation was more directly demonstrated by elucidating a role for HapR in the repression of a gene encoding the extracellular nuclease Dns [17]. The finding that HapR “acts as a negative regulator for dns transcription” was recently also confirmed by others [57]. The study by Blokesch and Schoolnik unambiguously demonstrated that the HapR-induced repression of the nuclease is the main, but not the only, contribution of QS to natural transformability, and the authors proposed that HapR also acts as a positive regulator of comEA [17]. This speculation was based on earlier microarray expression data, which showed that comEA expression upon growth on chitin was significantly reduced in the absence of HapR [11]. A QS-dependent regulation of comEA has recently been demonstrated [21] but differed from the data presented here as discussed above. In the current study, we extended this analysis by studying natural transformation and competence gene expression using the same competence-inducing conditions. Furthermore, we simultaneously monitored the expression of two competence genes, namely comEA and pilA, and compared their expression levels in different QS mutants. Based on the data provided, we conclude that comEA but not pilA is regulated by QS (Figure 7). We also showed for the first time a direct link between intracellular HapR protein concentrations, nuclease repression and comEA induction (Figure 4 and Figure 5).
Finally, we tested other competence genes, namely pilM, VC0047, dprA, and comEC, with respect to their expression levels; for these genes less or no information concerning their regulation was known before this study. We conclusively showed that all tested competence genes were dependent on induction by TfoX but only a part of them was also co-regulated in a QS-dependent manner (Figure 6). In fact only three genes showed a significantly different expression under competence inducing conditions in a wild type strain compared to a hapR mutant, namely dns, comEA and comEC (Figure 7). The encoded proteins are all directly involved in the fate of the surrounding DNA: whereas Dns degrades free DNA at low cell density, ComEA and ComEC are required for the DNA uptake process once high cell density is reached on the chitin surface (Figure 7). Further studies will follow to provide a better insight into the DNA uptake process itself.
Materials and Methods
Bacterial strains and plasmids
Bacterial strains and plasmids used in this study are listed in Table 1. E. coli strains DH5α [58] and One Shot PIR1 or PIR2 (Invitrogen) were used as hosts for cloning purposes. E. coli strain S17-1λpir [59] served as mating donor for plasmid transfers between E. coli and V. cholerae.
Media and growth conditions
Overnight cultures were grown in LB medium under aerobic conditions. Defined artificial seawater medium (DASW) [11] supplemented with vitamins (MEM, Gibco) and 0.1% casamino acids (Becton, Dickson and Company) was used for static growth of V. cholerae on chitin beads (New England Biolabs) or for growth under shaking conditions with N-acetylglucosamine (GlcNAc) or hexa-N-acetylchitohexaose (GlcNAc)6 (obtained from Seikagaku Corporation via Northstar BioProducts and LuBioScience, Lucerne, Switzerland) as sole carbon source. Thiosulfate Citrate Bile Salts Sucrose (TCBS) agar plates were prepared following the manufacturer's instructions (Fluka) and were used to counter-select E. coli strains after triparental mating with V. cholerae. Experiments for artificial tfoX expression were performed in LB medium with or without the addition of 0.02% arabinose. LB medium and LB agar plates were supplemented with antibiotics wherever required. The final concentrations of antibiotics were 75 µg/ml for kanamycin, 50 or 100 µg/ml for ampicillin and 50 µg/ml for gentamicin. V. cholerae cells were always grown at 30°C.
Construction of V. cholerae strains
The V. cholerae cpdA deletion strain was constructed using plasmid pGP704-28-SacB-ΔcpdA (Table 1) and the gene disruption method described previously [31]. The oligonucleotides used for the construction of the deletion plasmid are indicated in Table S1.
V. cholerae strains carrying artificially inducible tfoX on the chromosome (e.g., in cis) were created by triparental mating between the respective V. cholerae strain (Table 1), E. coli strain S17λpir/pUX-BF13 (providing the transposase function; [60]), or E. coli strain S17λpir/pGP704-mTn7-araC-tfoX. The latter plasmid consists of the suicide vector pGP704 as the backbone and the mini-Tn7 transposon [43] containing the gene cluster araC-PBAD-tfoX as cargo. This gene cluster was amplified by PCR from the plasmid pBAD-tfoX-stop [22] (primers indicated in Table S1).
Construction of transcriptional reporter fusions
The FP reporter constructs are all based on plasmid pBR322 [61]. Initially, this plasmid was modified by (partial) deletion of the tetracycline resistance cassette as well as the constitutive promoter PTet, resulting in plasmid pBR-Tet_MCSI (Table 1). A kanamycin resistance cassette (aph) as well as promoter-less versions of the genes gfp and dsRed (DsRed.T3[DNT]), both pointing in the opposite direction, were inserted into pBR-Tet_MCSI to yield plasmid pBR-GFP_dsRed_Kan. Details of all plasmids are included in Table 1. The primers used for construction of the plasmids are listed in Table S1 (synthesized by Microsynth, Switzerland). PCR mixtures, PCR programs, restriction enzyme digestions, primer phosphorylation, and ligations followed standard protocols recommended by the manufacturers of the enzymes (Roche, Switzerland and New England Biolabs via Bioconcept, Switzerland).
Growth of V. cholerae on chitin beads
V. cholerae strains were grown aerobically in LB medium until an OD600 of ∼0.3. Cells were harvested, washed in DASW medium and mixed with an equal volume of prewashed chitin beads (New England Biolabs) and three volumes of DASW (final volume 1 ml). The mixture was supplemented with vitamins, 0.1% casamino acids, and kanamycin (for plasmid maintenance). The bacteria were grown as standing cultures in 12-well plates. Bacteria were visualized by epifluorescence microscopy after 24 and 48 hours of growth, respectively.
Epifluorescence microscopy settings
Specifications of the epifluorescence microscope were as follows: Zeiss Axio Imager M2 microscope; Zeiss High Resolution Microscopy Camera AxioCam MRm; Illuminator HXP 120 as fluorescence light source (metal halide); objective used in this study: Plan-Apochromat 100×/1.40 Oil Ph3 M27 (WD = 0.17 mm). Filters relevant to this study were Zeiss Filter set 63 HE mRFP shift free; EX BP 572/25, BS FT 590, EM BP 629/62 and Zeiss filter set 38 Endow GFP shift free; EX BP 470/40, BS FT 495, EM BP 525/50. Image acquisition was done using the Zeiss AxioVision software. Images were rotated, cropped and uniformly enhanced with respect to contrast and brightness using Zeiss AxioVision and Adobe Photoshop CS3.
Microscopy image analysis was performed using the Matlab-based MicrobeTracker Suite [62] and according to the instructions given by the inventors (http://microbetracker.org/). Fluorescence intensities were normalized with respect to the area of the cell and the exposure time.
Induction of natural competence by hexa-N-acetylchitohexaose and flow cytometry
Chitin-dependent but surface-independent growth of V. cholerae was performed as previously described [23] using 2 mM of hexa-N-acetylchitohexaose GlcNAc6 (obtained from Seikagaku Corporation via Northstar BioProducts and LuBioScience, Lucerne, Switzerland) as sole carbon source. The same strains were grown in parallel with N-acetylglucosamine (GlcNAc; control). Bacteria were harvested at an OD600 of 0.8. Bacteria were either immediately visualized by epifluorescence microscopy or fixed in 2% paraformaldehyde for 30 min. Fixed samples were washed, diluted in PBS (1∶5), and analyzed by flow cytometry using a BD LSR II Flow cytometer. BD FACSDiva software was used for data acquisition. GFP signals were excited with a blue laser (488 nm) and detected with a 525/50 filter. DsRed.T3[DNT] was excited with a green laser (561 nm) and detected with a 585/15 filter. For each sample, 100,000 events were counted in total. Biologically independent experimental replicates (three for Figure 1 and Figure 2; two for Figure S1 and Figure S3) were performed within two weeks. One representative experiment is depicted in Figure 2 and the averages of the mean fluorescence intensities from three different biological replicates are shown in Figure S4.
Chitin-independent induction of competence
Strains used for chitin-independent competence-induction all carried inducible tfoX (araC-PBAD-tfoX) on a mini-Tn7 transposon [43] within the chromosome. Induction was accomplished by growth in LB supplemented with 0.02% arabinose.
For transformation assays, cells were grown until an OD600∼1.0. At that point, aliquots of 0.5 ml cultures were transferred to 1.5-ml tubes and supplemented with 2 µg/ml transforming DNA (gDNA of strain A1552-LacZ-Kan; [19]). Tubes were shaken horizontally for 5 h. Transformed cells and total colony forming units (CFU) were enumerated by a variation of a previously described method [63]. Briefly, the cultures underwent a serial dilution, and 5 µl of each dilution step was spotted in duplicate or triplicate on plain LB or LB containing 75 µg/ml kanamycin plates, respectively. Transformation frequencies were calculated as number of transformants divided by total number of CFUs. Each experiment was repeated at least three independent times, and the averages of all experiments are given in the figures (± standard deviations). Statistical analyses of transformation frequencies were performed on log-transformed data [64] using a two-tailed Student's t test.
Strains were grown in LB medium with or without 0.02% arabinose to investigate transcriptional FP reporter fusions. After 24 h, the cells were either visualized by epifluorescence microscopy or measured for relative fluorescence using a Tecan Infinite M200 plate reader. Parameters for detection of GFP were: excitation (Ex) at 485 nm (9 nm bandwidth) and emission (Em) at 515 nm (20 nm bandwidth). DsRed.T3[DNT] was detected using Ex 560 (9)/Em 587 (20) nm, as previously described for DsRed.T3 [28]. The samples were also measured with respect to their OD600, and results are given as relative fluorescence units (RFU) divided by OD600 values. The averages of three biological replicates are shown. Error bars indicate standard deviations. Statistically significant differences were analyzed using two-tailed Student's t tests.
Quantitative reverse transcription PCR (qRT-PCR)
V. cholerae strains were grown in 10 ml LB in the absence or presence of 0.02% arabinose until they reached an optical density of ∼1.7. At that time 5 ml of each culture was harvested and lysed in 1 ml Tri Reagent (Sigma). The samples were stored at −80°C. RNA isolation, DNase treatment, and reverse transcription using 1 µg of total RNA as template was done as previously described [23]. The obtained cDNA was diluted 40-fold and served as template in the qPCR. The primers used for the qPCR are indicated in Table S1. The qPCR mix was based on the Fast Start Essential DNA Green Master Mix (Roche, Switzerland), a ready-to use hot start reaction mix optimized for qPCR using the Light Cycler Nano system from Roche. The qPCR mix further contained 0.5 µM of each primer. The qPCR run using the Light Cycler Nano was performed according to these parameters: a denaturation step at 95°C for 10 min followed by 40 cycles of 95°C for 10 s, 60°C for 20 s, 72°C for 20 s. Each run was finished with a melting-curve ranging from 50°C to 95°C to validate specific amplification, which was also initially confirmed for each primer pair by standard PCR and visualization of PCR fragments in agarose gels. For each sample a reverse transcriptase-negative control was also performed while doing the reverse transcription and the respective samples were analyzed with at least three independent primer pairs to exclude residual DNA contaminations. A standard curve was prepared for each primer pair using purified genomic DNA of V. cholerae A1552 diluted in PCR grade water (from 1000 to 0.1 pg gDNA template). A negative control lacking any template was also tested for each primer pair. The expression values were normalized against expression of the housekeeping gene gyrA as previously described [65]. However, as discussed above the expression of gyrA might be dependent on DNA supercoiling and the cell cycle as shown for other bacteria [34], [35]. We therefore compared the relative expression of the four housekeeping genes gyrA, recA, clpX, ftsH as well as comEA in two different V. cholerae strains (WT and ΔhapR). The expression patterns were extremely similar no matter whether we normalized the expression data against gyrA expression (Figure S8A) or against recA expression (Figure S8B) as internal controls. The results were analyzed using the Light Cycler Nano software.
Preparation of cell lysates and determination of total protein concentration
For the preparation of cell lysates, bacteria were grown aerobically in LB medium in the absence or presence of 0.02% arabinose. Cells were harvested after reaching an OD600 of ∼1.5, resuspended in SDS-loading buffer (2×Laemmli buffer without β-mercaptoethanol and bromophenol blue), and boiled at 98°C for 15 min. Total protein concentration was quantified using the Pierce BCA Protein Assay kit (Thermo Scientific) before the addition of β-mercaptoethanol and bromophenol blue (7.5% and 0.01% final concentrations, respectively).
Generation of antibodies against HapR and Dns
Antibodies raised against the peptides derived from the proteins HapR, Dns, and TfoX were produced by Biomatik (Canada). The polyclonal antibody production service included suggestions for the design of two peptides per protein, peptide synthesis, conjugation of the peptides to the carrier proteins (keyhole limpet hemocyanin), and immunization of two rabbits per peptide mix. Polyclonal antibodies were affinity purified against the antigen and checked by ELISA. Each antibody was first validated for potential cross-reactions at the same size as the target protein using western blot analysis of the respective know-out strains.
Electrophoretic separations and Western blotting
Separation of proteins under denaturing conditions was conducted by SDS-PAGE using 15% acrylamide gels [66], [67]. The amount of total protein loaded per lane was 6 µg, 12 µg, and 50 µg for HapR, Dns, and TfoX detection, respectively. For western blot analysis, the proteins were transferred onto PVDF western blotting membranes (Roche), stained with amido black to verify transfer efficiency, incubated in blocking buffer, and reacted with primary antibodies directed against HapR (1∶5000), Dns (1∶1000), or TfoX (1∶2000). Detection of the primary antibody was performed using a secondary goat anti-rabbit IgG antibody conjugated to peroxidase (Sigma A9169; used at a 1∶20,000 dilution). Signals were revealed using Lumi-LightPLUS Western Blotting substrate (Roche, Switzerland) and were recorded by exposure to chemiluminescence-detecting films (Amersham Hyperfilm ECL, GE Healthcare).
Supporting Information
Zdroje
1. MorensDMFolkersGKFauciAS 2004 The challenge of emerging and re-emerging infectious diseases. Nature 430 242 249
2. BertuzzoEMariLRighettoLGattoMCasagrandiR 2011 Prediction of the spatial evolution and effects of control measures for the unfolding Haiti cholera outbreak. Geophys Res Lett 38doi:10.1029/2011GL046823
3. AndrewsJRBasuS 2011 Transmission dynamics and control of cholera in Haiti: an epidemic model. Lancet 377 1248 1255
4. TuiteARTienJEisenbergMEarnDJMaJ 2011 Cholera Epidemic in Haiti, 2010: Using a Transmission Model to Explain Spatial Spread of Disease and Identify Optimal Control Interventions. Ann Intern Med 154 593 601
5. RinaldoABlokeschMBertuzzoEMariLRighettoL 2011 A transmission model of the 2010 cholera epidemic in haiti. Ann Intern Med 155 403 404
6. ChinCSSorensonJHarrisJBRobinsWPCharlesRC 2010 The origin of the Haitian cholera outbreak strain. N Engl J Med 364 33 42
7. ChunJGrimCJHasanNALeeJHChoiSY 2009 Comparative genomics reveals mechanism for short-term and long-term clonal transitions in pandemic Vibrio cholerae. Proc Natl Acad Sci USA 106 15442 15447
8. LippEKHuqAColwellRR 2002 Effects of global climate on infectious disease: the cholera model. Clin Microbiol Rev 15 757 770
9. ColwellRR 1996 Global climate and infectious disease: the cholera paradigm. Science 274 2025 2031
10. PruzzoCVezzulliLColwellRR 2008 Global impact of Vibrio cholerae interactions with chitin. Environ Microbiol 10 1400 1410
11. MeibomKLBlokeschMDolganovNAWuC-Y Schoolnik GK 2005 Chitin induces natural competence in Vibrio cholerae. Science 310 1824 1827
12. ChenYDaiJMorrisJGJrJohnsonJA 2010 Genetic analysis of the capsule polysaccharide (K antigen) and exopolysaccharide genes in pandemic Vibrio parahaemolyticus O3:K6. BMC Microbiol 10 274
13. GuligPATuckerMSThiavillePCJosephJLBrownRN 2009 USER friendly cloning coupled with chitin-based natural transformation enables rapid mutagenesis of Vibrio vulnificus. Appl Environ Microbiol 75 4936 4949
14. Pollack-BertiAWollenbergMSRubyEG 2010 Natural transformation of Vibrio fischeri requires tfoX and tfoY. Environ Microbiol 12 2302 2311
15. BlokeschM Schoolnik GK 2007 Serogroup Conversion of Vibrio cholerae in Aquatic Reservoirs. PLoS Pathog 3 e81 doi:10.1371/journal.ppat.0030081
16. FaruqueSMMekalanosJJ 2003 Pathogenicity islands and phages in Vibrio cholerae evolution. Trends Microbiol 11 505 510
17. BlokeschM Schoolnik GK 2008 The extracellular nuclease Dns and its role in natural transformation of Vibrio cholerae. J Bacteriol 190 7232 7240
18. YamamotoSMoritaMIzumiyaHWatanabeH 2010 Chitin disaccharide (GlcNAc)2 induces natural competence in Vibrio cholerae through transcriptional and translational activation of a positive regulatory gene tfoXVC. Gene 457 42 49
19. MarvigRLBlokeschM 2010 Natural transformation of Vibrio cholerae as a tool-optimizing the procedure. BMC Microbiol 10 155
20. YamamotoSIzumiyaHMitobeJMoritaMArakawaE 2011 Identification of a Chitin-Induced Small RNA That Regulates Translation of the tfoX Gene, Encoding a Positive Regulator of Natural Competence in Vibrio cholerae. J Bacteriol 193 1953 1965
21. AntonovaESHammerBK 2011 Quorum-sensing autoinducer molecules produced by members of a multispecies biofilm promote horizontal gene transfer to Vibrio cholerae. FEMS Microbiol Lett 322 68 76
22. SuckowGSeitzPBlokeschM 2011 Quorum Sensing Contributes to Natural Transformation of Vibrio cholerae in a Species-Specific Manner. J Bacteriol 193 4914 4924
23. BlokeschM 2012 Chitin colonization, chitin degradation and chitin-induced natural competence of Vibrio cholerae are subject to catabolite repression. Environ Microbiol doi:10.1111/j.1462-2920.2011.02689.x (article first published online: 6 JAN 2012)
24. NesterEWStockerBA 1963 Biosynthetic Latency in Early Stages of Deoxyribonucleic Acidtransformation in Bacillus Subtilis. J Bacteriol 86 785 796
25. MaamarHDubnauD 2005 Bistability in the Bacillus subtilis K-state (competence) system requires a positive feedback loop. Mol Microbiol 56 615 624
26. MaamarHRajADubnauD 2007 Noise in gene expression determines cell fate in Bacillus subtilis. Science 317 526 529
27. CormackBPValdiviaRHFalkowS 1996 FACS-optimized mutants of the green fluorescent protein (GFP). Gene 173 33 38
28. BevisBJGlickBS 2002 Rapidly maturing variants of the Discosoma red fluorescent protein (DsRed). Nat Biotechnol 20 83 87
29. DunnAKMillikanDSAdinDMBoseJLStabbEV 2006 New rfp - and pES213-derived tools for analyzing symbiotic Vibrio fischeri reveal patterns of infection and lux expression in situ. Appl Environ Microbiol 72 802 810
30. FullnerKJMekalanosJJ 1999 Genetic characterization of a new type IV-A pilus gene cluster found in both classical and El Tor biotypes of Vibrio cholerae. Infect Immun 67 1393 1404
31. MeibomKLLiXBNielsenATWuCYRosemanS 2004 The Vibrio cholerae chitin utilization program. Proc Natl Acad Sci USA 101 2524 2529
32. ProvvediRDubnauD 1999 ComEA is a DNA receptor for transformation of competent Bacillus subtilis. Mol Microbiol 31 271 280
33. DohertyAJSerpellLCPontingCP 1996 The helix-hairpin-helix DNA-binding motif: a structural basis for non-sequence-specific recognition of DNA. Nucleic Acids Res 24 2488 2497
34. MenzelRGellertM 1983 Regulation of the genes for E. coli DNA gyrase: homeostatic control of DNA supercoiling. Cell 34 105 113
35. LaubMTMcAdamsHHFeldblyumTFraserCMShapiroL 2000 Global analysis of the genetic network controlling a bacterial cell cycle. Science 290 2144 2148
36. FongJCYildizFH 2008 Interplay between cyclic AMP-cyclic AMP receptor protein and cyclic di-GMP signaling in Vibrio cholerae biofilm formation. J Bacteriol 190 6646 6659
37. NielsenATDolganovNARasmussenTOttoGMillerMC 2010 A Bistable Switch and Anatomical Site Control Vibrio cholerae Virulence Gene Expression in the Intestine. PLoS Pathog 6 doi:10.1371/journal.ppat.1001102 e1001102
38. SrivastavaPChattorajDK 2007 Selective chromosome amplification in Vibrio cholerae. Mol Microbiol 66 1016 1028
39. SilanderOKNikolicNZaslaverABrenAKikoinI 2012 A genome-wide analysis of promoter-mediated phenotypic noise in Escherichia coli. PLoS Genet 8 doi:10.1371/journal.pgen.1002443 e1002443
40. RedfieldRJCameronADQianQHindsJAliTR 2005 A novel CRP-dependent regulon controls expression of competence genes in Haemophilus influenzae. J Mol Biol 347 735 747
41. CameronADRedfieldRJ 2008 CRP binding and transcription activation at CRP-S sites. J Mol Biol 383 313 323
42. SchleifR 2010 AraC protein, regulation of the l-arabinose operon in Escherichia coli, and the light switch mechanism of AraC action. FEMS Microbiol Rev 34 779 796
43. LambertsenLSternbergCMolinS 2004 Mini-Tn7 transposons for site-specific tagging of bacteria with fluorescent proteins. Environ Microbiol 6 726 732
44. DeutscherJFranckeCPostmaPW 2006 How phosphotransferase system-related protein phosphorylation regulates carbohydrate metabolism in bacteria. Microbiol Mol Biol Rev 70 939 1031
45. KimHSKimSMLeeHJParkSJLeeKH 2009 Expression of the cpdA gene, encoding a 3′,5′-cyclic AMP (cAMP) phosphodiesterase, is positively regulated by the cAMP-cAMP receptor protein complex. J Bacteriol 191 922 930
46. HigginsDAPomianekMEKramlCMTaylorRKSemmelhackMF 2007 The major Vibrio cholerae autoinducer and its role in virulence factor production. Nature 450 883 886
47. NgWLBasslerBL 2009 Bacterial quorum-sensing network architectures. Annu Rev Genet 43 197 222
48. XavierKBBasslerBL 2005 Interference with AI-2-mediated bacterial cell-cell communication. Nature 437 750 753
49. MillerMBSkorupskiKLenzDHTaylorRKBasslerBL 2002 Parallel quorum sensing systems converge to regulate virulence in Vibrio cholerae. Cell 110 303 314
50. JoblingMGHolmesRK 1997 Characterization of hapR, a positive regulator of the Vibrio cholerae HA/protease gene hap, and its identification as a functional homologue of the Vibrio harveyi luxR gene. Mol Microbiol 26 1023 1034
51. WatersCMBasslerBL 2006 The Vibrio harveyi quorum-sensing system uses shared regulatory components to discriminate between multiple autoinducers. Genes Dev 20 2754 2767
52. Mortier-BarriereIVeltenMDupaignePMirouzeNPietrementO 2007 A key presynaptic role in transformation for a widespread bacterial protein: DprA conveys incoming ssDNA to RecA. Cell 130 824 836
53. LiXRosemanS 2004 The chitinolytic cascade in Vibrios is regulated by chitin oligosaccharides and a two-component chitin catabolic sensor/kinase. Proc Natl Acad Sci USA 101 627 631
54. MacfadyenLP 2000 Regulation of competence development in Haemophilus influenzae. J Theor Biol 207 349 359
55. HeidelbergJFEisenJANelsonWCClaytonRAGwinnML 2000 DNA sequence of both chromosomes of the cholera pathogen Vibrio cholerae. Nature 406 477 483
56. ZhuJMillerMBVanceREDziejmanMBasslerBL 2002 Quorum-sensing regulators control virulence gene expression in Vibrio cholerae. Proc Natl Acad Sci USA 99 3129 3134
57. SeperAFenglerVHRoierSWolinskiHKohlweinSD 2011 Extracellular nucleases and extracellular DNA play important roles in Vibrio cholerae biofilm formation. Mol Microbiol 82 1015 1037
58. Yanisch-PerronCVieiraJMessingJ 1985 Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33 103 119
59. SimonRPrieferUPühlerA 1983 A Broad Host Range Mobilization System for In Vivo Genetic Engineering: Transposon Mutagenesis in Gram Negative Bacteria. Nat Biotechnol 1 784 791
60. BaoYLiesDPFuHRobertsGP 1991 An improved Tn7-based system for the single-copy insertion of cloned genes into chromosomes of gram-negative bacteria. Gene 109 167 168
61. BolivarFRodriguezRLGreenePJBetlachMCHeynekerHL 1977 Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene 2 95 113
62. SliusarenkoOHeinritzJEmonetTJacobs-WagnerC 2011 High-throughput, subpixel precision analysis of bacterial morphogenesis and intracellular spatio-temporal dynamics. Mol Microbiol 80 612 627
63. MilesAAMisraSSIrwinJO 1938 The estimation of the bactericidal power of the blood. J Hyg 38 732 749
64. KeeneON 1995 The log transformation is special. Stat Med 14 811 819
65. LiuXBeyhanSLimBLiningtonRGYildizFH 2010 Identification and characterization of a phosphodiesterase that inversely regulates motility and biofilm formation in Vibrio cholerae. J Bacteriol 192 4541 4552
66. LaemmliUK 1970 Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227 680 685
67. SambrookJFritschEFManiatisT 2001 Molecular Cloning: A Laboratory Manual (Third Edition). FordNNolanCFergusonM New York Cold Spring Harbor Laboratory Press editors
68. YildizFH Schoolnik GK 1998 Role of rpoS in stress survival and virulence of Vibrio cholerae. J Bacteriol 180 773 784
69. De Souza SilvaOBlokeschM 2010 Genetic manipulation of Vibrio cholerae by combining natural transformation with FLP recombination. Plasmid 64 186 195
70. NielsenATDolganovNAOttoGMillerMCWuCY 2006 RpoS controls the Vibrio cholerae mucosal escape response. PLoS Pathog 2 doi:10.1371/journal.ppat.0020109 e109
Štítky
Genetika Reprodukční medicína
Článek Uracil-Containing DNA in : Stability, Stage-Specific Accumulation, and Developmental InvolvementČlánek Fuzzy Tandem Repeats Containing p53 Response Elements May Define Species-Specific p53 Target GenesČlánek Preferential Genome Targeting of the CBP Co-Activator by Rel and Smad Proteins in Early EmbryosČlánek Protective Coupling of Mitochondrial Function and Protein Synthesis via the eIF2α Kinase GCN-2Článek Cohesin Proteins Promote Ribosomal RNA Production and Protein Translation in Yeast and Human CellsČlánek TERRA Promotes Telomere Shortening through Exonuclease 1–Mediated Resection of Chromosome EndsČlánek Attenuation of Notch and Hedgehog Signaling Is Required for Fate Specification in the Spinal CordČlánek Genome-Wide Functional Profiling Identifies Genes and Processes Important for Zinc-Limited Growth ofČlánek MicroRNA93 Regulates Proliferation and Differentiation of Normal and Malignant Breast Stem Cells
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2012 Číslo 6- Akutní intermitentní porfyrie
- Transthyretinová amyloidóza z pohledu neurologa a kardiologa aneb jak se vyhnout „misdiagnostice“?
- Mateřský haplotyp KIR ovlivňuje porodnost živých dětí po transferu dvou embryí v rámci fertilizace in vitro u pacientek s opakujícími se samovolnými potraty nebo poruchami implantace
- Psychika ženy, které se nedaří otěhotnět
- Vitamín B6 jako prevence kolorektálního karcinomu
-
Všechny články tohoto čísla
- Rumors of Its Disassembly Have Been Greatly Exaggerated: The Secret Life of the Synaptonemal Complex at the Centromeres
- Mimetic Butterflies Introgress to Impress
- Tipping the Balance in the Powerhouse of the Cell to “Protect” Colorectal Cancer
- Selection-Driven Gene Loss in Bacteria
- Decreased Mitochondrial DNA Mutagenesis in Human Colorectal Cancer
- Parallel Evolution of Auditory Genes for Echolocation in Bats and Toothed Whales
- Diverse CRISPRs Evolving in Human Microbiomes
- The Rad4 ATR-Activation Domain Functions in G1/S Phase in a Chromatin-Dependent Manner
- Stretching the Rules: Monocentric Chromosomes with Multiple Centromere Domains
- Uracil-Containing DNA in : Stability, Stage-Specific Accumulation, and Developmental Involvement
- Fuzzy Tandem Repeats Containing p53 Response Elements May Define Species-Specific p53 Target Genes
- Adaptive Introgression across Species Boundaries in Butterflies
- G Protein Activation without a GEF in the Plant Kingdom
- Synaptonemal Complex Components Persist at Centromeres and Are Required for Homologous Centromere Pairing in Mouse Spermatocytes
- An Engineering Approach to Extending Lifespan in
- Incompatibility and Competitive Exclusion of Genomic Segments between Sibling Species
- Effects of Histone H3 Depletion on Nucleosome Occupancy and Position in
- Patterns of Evolutionary Conservation of Essential Genes Correlate with Their Compensability
- Interplay between Synaptonemal Complex, Homologous Recombination, and Centromeres during Mammalian Meiosis
- Preferential Genome Targeting of the CBP Co-Activator by Rel and Smad Proteins in Early Embryos
- A Mouse Model of Acrodermatitis Enteropathica: Loss of Intestine Zinc Transporter ZIP4 (Slc39a4) Disrupts the Stem Cell Niche and Intestine Integrity
- Protective Coupling of Mitochondrial Function and Protein Synthesis via the eIF2α Kinase GCN-2
- Geographic Differences in Genetic Susceptibility to IgA Nephropathy: GWAS Replication Study and Geospatial Risk Analysis
- Cohesin Proteins Promote Ribosomal RNA Production and Protein Translation in Yeast and Human Cells
- TERRA Promotes Telomere Shortening through Exonuclease 1–Mediated Resection of Chromosome Ends
- Stimulation of Host Immune Defenses by a Small Molecule Protects from Bacterial Infection
- A Broad Requirement for TLS Polymerases η and κ, and Interacting Sumoylation and Nuclear Pore Proteins, in Lesion Bypass during Embryogenesis
- Genome-Wide Identification of Ampicillin Resistance Determinants in
- The CCR4-NOT Complex Is Implicated in the Viability of Aneuploid Yeasts
- Gustatory Perception and Fat Body Energy Metabolism Are Jointly Affected by Vitellogenin and Juvenile Hormone in Honey Bees
- Genetic Variants on Chromosome 1q41 Influence Ocular Axial Length and High Myopia
- Is a Key Regulator of Pancreaticobiliary Ductal System Development
- The NSL Complex Regulates Housekeeping Genes in
- Attenuation of Notch and Hedgehog Signaling Is Required for Fate Specification in the Spinal Cord
- Dual-Level Regulation of ACC Synthase Activity by MPK3/MPK6 Cascade and Its Downstream WRKY Transcription Factor during Ethylene Induction in Arabidopsis
- Genome-Wide Functional Profiling Identifies Genes and Processes Important for Zinc-Limited Growth of
- Base-Pair Resolution DNA Methylation Sequencing Reveals Profoundly Divergent Epigenetic Landscapes in Acute Myeloid Leukemia
- MicroRNA93 Regulates Proliferation and Differentiation of Normal and Malignant Breast Stem Cells
- Phylogenomic Analysis Reveals Dynamic Evolutionary History of the Drosophila Heterochromatin Protein 1 (HP1) Gene Family
- Found: The Elusive ANTAR Transcription Antiterminator
- The Mutation in Chickens Constitutes a Structural Rearrangement Causing Both Altered Comb Morphology and Defective Sperm Motility
- Control of CpG and Non-CpG DNA Methylation by DNA Methyltransferases
- Polymorphisms in the Mitochondrial Ribosome Recycling Factor Compromise Cell Respiratory Function and Increase Atorvastatin Toxicity
- Gene Expression Profiles in Parkinson Disease Prefrontal Cortex Implicate and Genes under Its Transcriptional Regulation
- Global Regulatory Functions of the Endoribonuclease III in Gene Expression
- Extensive Evolutionary Changes in Regulatory Element Activity during Human Origins Are Associated with Altered Gene Expression and Positive Selection
- The Regulatory Network of Natural Competence and Transformation of
- Brain Expression Genome-Wide Association Study (eGWAS) Identifies Human Disease-Associated Variants
- Quantifying the Adaptive Potential of an Antibiotic Resistance Enzyme
- Divergence of the Yeast Transcription Factor Affects Sulfite Resistance
- The Histone Demethylase Jhdm1a Regulates Hepatic Gluconeogenesis
- RNA Methylation by the MIS Complex Regulates a Cell Fate Decision in Yeast
- Rare Copy Number Variants Observed in Hereditary Breast Cancer Cases Disrupt Genes in Estrogen Signaling and Tumor Suppression Network
- Genome-Wide Location Analysis Reveals Distinct Transcriptional Circuitry by Paralogous Regulators Foxa1 and Foxa2
- A Mutation Links a Canine Progressive Early-Onset Cerebellar Ataxia to the Endoplasmic Reticulum–Associated Protein Degradation (ERAD) Machinery
- The Mechanism for RNA Recognition by ANTAR Regulators of Gene Expression
- Limits to the Rate of Adaptive Substitution in Sexual Populations
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Rumors of Its Disassembly Have Been Greatly Exaggerated: The Secret Life of the Synaptonemal Complex at the Centromeres
- The NSL Complex Regulates Housekeeping Genes in
- Tipping the Balance in the Powerhouse of the Cell to “Protect” Colorectal Cancer
- Interplay between Synaptonemal Complex, Homologous Recombination, and Centromeres during Mammalian Meiosis
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Revma Focus: Spondyloartritidy
nový kurz
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MUDr. Irena Krčmová, CSc.
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



