-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Nucleosomes Containing Methylated DNA Stabilize DNA Methyltransferases 3A/3B and Ensure Faithful Epigenetic Inheritance
How epigenetic information is propagated during somatic cell divisions is still unclear but is absolutely critical for preserving gene expression patterns and cellular identity. Here we show an unanticipated mechanism for inheritance of DNA methylation patterns where the epigenetic mark not only recruits the catalyzing enzyme but also regulates the protein level, i.e. the enzymatic product (5-methylcytosine) determines the level of the methylase, thus forming a novel homeostatic inheritance system. Nucleosomes containing methylated DNA stabilize de novo DNA methyltransferases, DNMT3A/3B, allowing little free DNMT3A/3B enzymes to exist in the nucleus. Stabilization of DNMT3A/3B on nucleosomes in methylated regions further promotes propagation of DNA methylation. However, reduction of cellular DNA methylation levels creating more potential CpG substrates counter-intuitively results in a dramatic decrease of DNMT3A/3B proteins due to diminished nucleosome binding and subsequent degradation of the unstable free proteins. These data show an unexpected self-regulatory inheritance mechanism that not only ensures somatic propagation of methylated states by DNMT1 and DNMT3A/3B enzymes but also prevents aberrant de novo methylation by causing degradation of free DNMT3A/3B enzymes.
Published in the journal: . PLoS Genet 7(2): e32767. doi:10.1371/journal.pgen.1001286
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1001286Summary
How epigenetic information is propagated during somatic cell divisions is still unclear but is absolutely critical for preserving gene expression patterns and cellular identity. Here we show an unanticipated mechanism for inheritance of DNA methylation patterns where the epigenetic mark not only recruits the catalyzing enzyme but also regulates the protein level, i.e. the enzymatic product (5-methylcytosine) determines the level of the methylase, thus forming a novel homeostatic inheritance system. Nucleosomes containing methylated DNA stabilize de novo DNA methyltransferases, DNMT3A/3B, allowing little free DNMT3A/3B enzymes to exist in the nucleus. Stabilization of DNMT3A/3B on nucleosomes in methylated regions further promotes propagation of DNA methylation. However, reduction of cellular DNA methylation levels creating more potential CpG substrates counter-intuitively results in a dramatic decrease of DNMT3A/3B proteins due to diminished nucleosome binding and subsequent degradation of the unstable free proteins. These data show an unexpected self-regulatory inheritance mechanism that not only ensures somatic propagation of methylated states by DNMT1 and DNMT3A/3B enzymes but also prevents aberrant de novo methylation by causing degradation of free DNMT3A/3B enzymes.
Introduction
DNA methylation is a stable gene silencing mechanism required for key biological processes including embryogenesis, genomic imprinting, X-chromosome inactivation, repression of transposons and maintenance of tissue specific gene expression patterns [1], [2]. Aberrant methylation contributes to tumorigenesis and other diseases [3], [4]. Thus, proper maintenance of DNA methylation patterns is essential for preserving cellular identity and preventing malignant cellular transformation.
In mammals, DNA methylation patterns are generally thought to be established during embryonic development by de novo DNA methyltransferases 3A and 3B [5] and then stably maintained through multiple somatic divisions by the ‘maintenance activity’ of DNMT1 both during and after replication [6]. However, recent studies suggest that DNMT1 alone cannot ensure proper maintenance of methylation patterns [7] and requires co-operative activity of the de novo DNMT3A/3B enzymes [8], [9], [10], which are ubiquitously expressed in somatic cells. A revised model of inheritance was recently proposed assigning DNMT3A/3B to a maintenance role in somatic cells [11]; however, questions still remain regarding the molecular mechanisms guiding the maintenance activity of these de novo enzymes.
In embryonic stem (ES) cells, DNMT3A/3B establish methylation patterns in association with DNMT3L, a regulatory factor which stimulates DNMT3A/3B de novo activity [12] and targets them to nucleosomes containing unmethylated H3K4 residues [13]. Methylated H3K4 containing chromatin regions remain refractory to such DNA methylation [14], [15]. Further, heterochromatin protein 1 (HP1) recruits DNMT3A/3B to H3K9me3 residues, established by histone methyltransferase (HMTase) Suv39h1/2, enabling de novo DNA methylation in pericentric heterochromatin [16]. In euchromatic regions, G9a, another H3K9 HMTase, recruits DNMT3A/3B for de novo methylation of early embryonic gene promoters [17]. UHRF1, which assists DNMT1 in locating to hemimethylated sites [18], also targets DNMT3A/3B for de novo methylation in ES cells [19]. However, DNMT3L is expressed only during gametogenesis and embryonic stages and not in somatic tissues [20], [21]. Further, we and others have recently shown that HP1 and UHRF1 are not required for DNMT3A/3B's association with nucleosomes [22] and G9a does not affect maintenance of DNA methylation in somatic cells [23], [24]. Thus, other mechanisms must exist to ensure proper localization of these enzymes to silent chromatin regions in somatic cells [25], enabling faithful maintenance of methylated states.
We and others have previously shown that the majority of DNMT3A/3B within a somatic cell are strongly anchored to nucleosomes containing methylated DNA with little free DNMT3A/3B proteins existing [22], [26]. Here we show that the presence of such methylated regions is essential for DNMT3A/3B's association with chromatin and quite unexpectedly, also for maintaining the cellular levels of these enzymes. Reduction in DNA methylation levels results in reduced DNMT3A/3B binding to nucleosomes accompanied by selective degradation of the free enzymes by the cellular machinery. Restoration of DNA methylation increases DNMT3A/3B protein levels through their stabilization on nucleosomes. Further, pre-existing methylation stimulates propagation of DNA methylation in vivo by stably anchoring DNMT3A/3B to nucleosomes. DNMT3A/3B work synergistically to propagate methylation patterns with DNMT3B stimulating DNMT3A activity by promoting its association with nucleosomes, similar to DNMT3L. Taken together, these data suggest an inheritance model where DNMT3A/3B remain localized to silent methylated domains by binding to nucleosomes containing methylated DNA, enabling faithful maintenance of methylated states in cooperation with DNMT1; while non-anchored DNMT3A/3B enzymes get selectively degraded preventing spurious de novo methylation.
Results
DNMT3A protein level decreases on depletion of global DNA methylation
In somatic cells, DNMT3A/3B remain bound to nucleosomes containing methylated DNA [22]. To investigate the role of DNA methylation in this binding, we used a series of HCT116 colon cancer cells with homozygous deletions for DNMT1 (DNMT1ΔE2-5; 1KO) [27], [28], DNMT3B (DNMT3B−/−; 3BKO) or both DNMT1 and DNMT3B (DNMT1ΔE2-5/DNMT3B−/−; double knockout, DKO) and consequently different levels of genomic DNA methylation [7]. For the DKO cells, which still contain residual DNMT1 activity [28], we used two clones for our analysis, DKO1 and DKO8, having lost ∼95% and ∼50% DNA methylation respectively [7]. RT-PCR analysis of DNMT3A, DNMT3B and DNMT1 transcript levels in the various HCT116 derivative cell lines showed similar or higher levels of DNMT3A1 transcripts in HCT116 knockout cell lines compared to WT HCT116; reduced levels of DNMT1ΔE2-5 hypomorph transcripts in 1KO and the two DKO clones, with relatively higher expression in the DKO8 clone and no detectable levels of DNMT3B transcripts in 3BKO and both DKO cell lines, consistent with previous data [7], [28] (Figure 1A).
Fig. 1. Transcription-independent decrease in DNMT3A protein level in hypomethylated DKO cells that contain severely impaired DNMT1 activity. 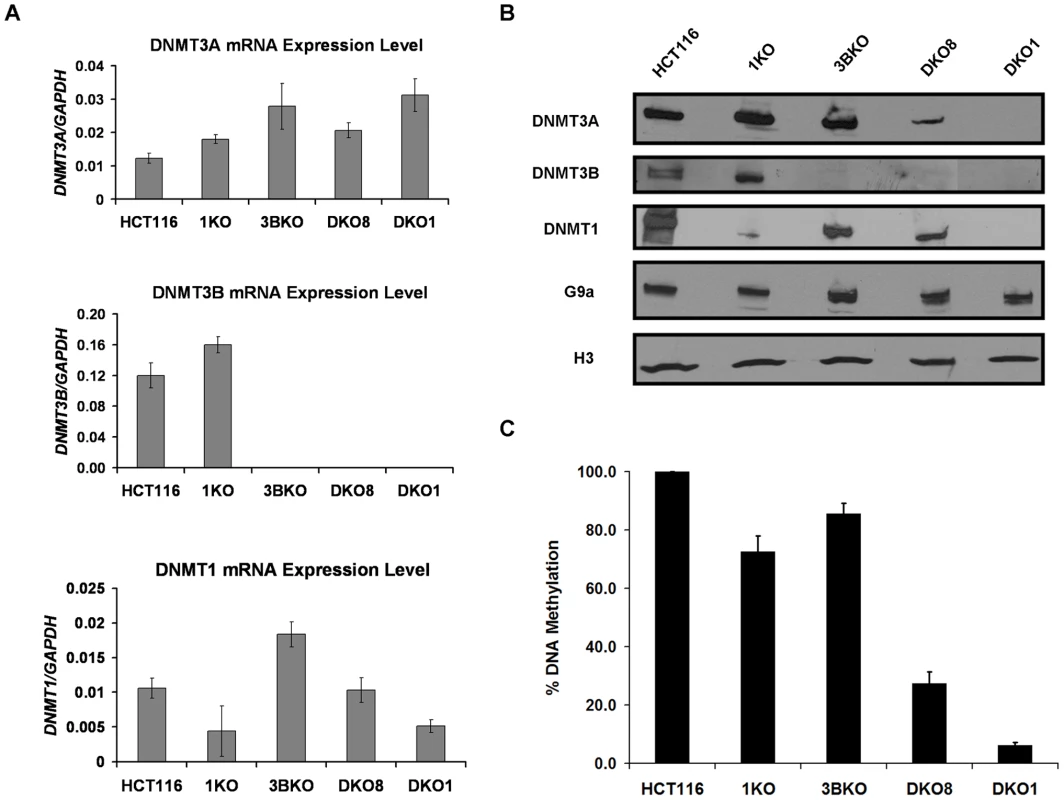
Next we examined DNMT protein levels in these cell lines through immunoblotting of nuclear extracts. Similar to mRNA analysis, DNMT3B and DNMT1 protein levels were severely reduced in the respective knockout cell lines (Figure 1B). Surprisingly, while DNMT3A mRNA levels were higher in both DKO clones, we found dramatically reduced DNMT3A protein in them compared to WT HCT116 cells. Similar reductions in DNMT3A protein levels were observed in whole cell lysates of both DKO cells, suggesting that the reduced nuclear levels are not the result of protein mislocalization (Figure S1). These findings were further confirmed by immunofluorescence analyses of HCT116 and DKO cells which displayed similar reduction in DNMT3A protein levels in DKO cells as observed in western blots of their nuclear extracts. Moreover, the residual DNMT3A protein displayed similar nuclear distribution in DKO cells as in WT HCT116 cells, confirming that its reduced nuclear levels in the DKO cells are not due to protein mislocalization (Figure S2). G9a, another chromatin-modifying protein, did not display such large changes in protein levels in HCT116 knockout cell lines (Figure 1B).
Assessment of global DNA methylation levels using methylation-sensitive restriction enzymes revealed a direct correlation between the amount of DNMT3A protein and level of methylation retained in the knockout cells, suggesting a possible role of DNA methylation in maintaining cellular DNMT3A levels (Figure 1B, 1C). DKO8 cells, which had retained higher DNA methylation levels, showed higher DNMT3A protein compared to the minimal amount present in the severely hypomethylated DKO1 cells. Since no such decrease in DNMT3A protein was observed in the single DNMT1 and DNMT3B knockout cells (1KO and 3BKO respectively), which retained substantial levels of DNA methylation, maintenance of DNMT3A levels through possible protein-protein interactions with DNMT1 and/or DNMT3B seems unlikely.
Residual DNMT3A protein remains tightly bound to chromatin in the DKO cells
We have previously shown that DNMT3A/3B strongly associate with methylated chromatin regions [22]. To determine whether the residual DNMT3A protein in hypomethylated DKO cells retains similar affinity for chromatin as in WT HCT116 cells, we performed a salt extraction experiment as described previously [22]. Purified nuclei from HCT116 and DKO1 cells were incubated in buffers with increasing concentrations (50 mM to 400 mM) of NaCl. Nuclear pellet and supernatant fractions were independently analyzed through western blot analysis. As expected, similar amounts of core histones remained inside the extracted nuclei under all salt concentrations. In HCT116 cells, the DNMT3A protein level remained almost constant within the nuclei up to 400 mM NaCl indicating a strong binding affinity for chromatin (Figure 2A), whereas other chromatin associated proteins such as EZH2 and G9a showed relatively weaker binding affinities with substantial amounts detected in the supernatant at more than 200 mM NaCl concentrations. Interestingly, the majority of DNMT3A protein present in DKO1 cells, though greatly reduced in comparison to WT HCT116, also remained tightly associated with the chromatin at all salt concentrations (Figure 2A), possibly binding to the few methylated regions remaining in the DKO1 cells. Minimal DNMT3A protein could be detected in the supernatant fractions (50 to 300 mM NaCl) of the DKO1 cells. These data suggest that binding to methylated chromatin regions might be essential for maintaining the stability of DNMT3A protein and that any free protein unable to bind to chromatin in the absence of DNA methylation possibly gets rapidly degraded by the cellular machinery. We did observe some DNMT3A protein dissociating from the chromatin at 400 mM NaCl in DKO1 cells but not in HCT116 cells suggesting a reduction in chromatin binding affinity of DNMT3A in hypomethylated DKO1 cells compared to heavily methylated WT HCT116 cells (Figure 2A). Meanwhile, EZH2 and G9a showed weaker binding to chromatin in DKO1 cells, similar to that observed in WT HCT116. Taken together, these data suggest that binding to methylated chromatin regions may be critical for stabilization of DNMT3A protein.
Fig. 2. DNMT3A chromatin binding affinity and protein stability in WT HCT116 and DKO cells. 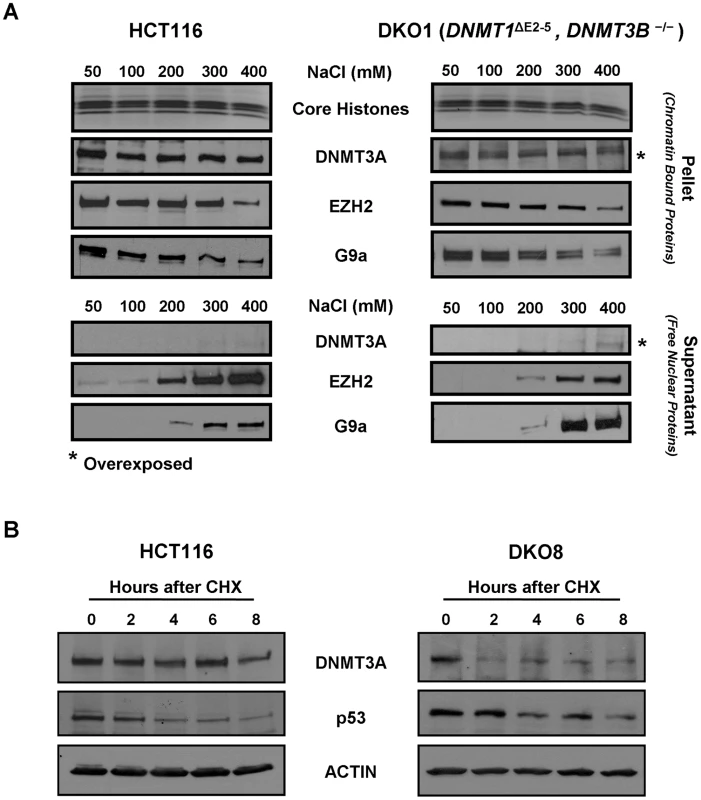
(A) Nuclei purified from WT HCT116 and DKO1 cells were incubated in nondenaturing extraction buffers containing 50 to 400 mM NaCl for 5 min. Equivalent volumes of both supernatant and pellet fractions were subjected to western blot analysis using specific antibodies. Ponceau S staining shows core histones transferred onto the membrane from the SDS/PAGE gel. For detecting low levels of DNMT3A in DKO1 cells, blots for both the supernatant and pellet fractions from DKO1 cells were overexposed for 5 fold more time duration compared to HCT116 cells, as indicated by *. (B) WT HCT116 and DKO8 cells were treated with cycloheximide (CHX) and the levels of DNMT3A protein remaining at different time points after treatment were determined by western blotting of nuclear extracts. p53 and actin were used as positive and loading controls, respectively. Data presented is from a single experiment, representative of two independent biological replicate experiments. Decreased protein stability of DNMT3A in hypomethylated DKO cells
To assess whether the dramatic transcription-independent decrease in steady-state levels of DNMT3A protein observed in hypomethylated DKO cells was due to altered protein stability, we treated WT HCT116 and DKO8 cells with the protein synthesis inhibitor cycloheximide (CHX) [29] and measured the DNMT3A protein remaining at different time points after treatment. DNMT3A was stable in WT HCT116 cells with 93% still remaining after 6 hrs of CHX treatment (Figure 2B). However, in DKO8 cells, DNMT3A was very unstable with its level rapidly decreasing to 49% 2 hrs after treatment. The half-life of DNMT3A protein decreased dramatically from 16 hrs in WT HCT116 to 7 hrs in DKO8 cells (Figure S3). Interestingly, after a rapid initial decrease in DNMT3A protein level in DKO8 cells within the first 2 hrs of CHX treatment, a fraction of DNMT3A protein remained stable thereafter till the 8 hr time point (Figure 2B). This fraction may possibly represent the stable DNMT3A protein bound to the methylated chromatin regions in DKO8 cells, similar to that observed in DKO1 cells (Figure 2A). Taken together, these data indicate that a decrease in DNA methylation results in destabilization of DNMT3A protein, possibly due to reduced chromatin binding in the absence of methylated DNA regions, the main sites of DNMT3A/3B binding [22].
Restoration of global DNA methylation rescues DNMT3A protein level
To ascertain if depletion of DNA methylation is primarily responsible for the decrease in DNMT3A protein, we sought to restore DNA methylation in the DKO cells. We expressed Myc-tagged DNMT3B1, ΔDNMT3B2 [30] or DNMT3L in DKO1 and DKO8 cells using a lentiviral system and confirmed expression of the relevant proteins by immunoblotting (Figure 3A). We did not use DNMT1 for the restoration of DNA methylation in DKO cells since expression of exogenous DNMT1 in DKO cells has previously been shown to result only in partial increase in DNA methylation [31]. More importantly, DNMT1 expression failed to restore methylation in these cells at the repetitive elements, the key sites of DNMT3A/3B binding [31]. Global DNA methylation levels in infected DKO cells were measured 8 weeks post-infection using methylation-sensitive restriction enzymes. Since DKO cells possess very low levels of a hypomorph of DNMT1 [28], the primary maintenance methyltransferase in the cell, very low levels of DNMT3A protein and no DNMT3B protein, it required a long time (∼8 weeks) to achieve restoration of DNA methylation in these cells. After 8 weeks of infection, we observed increased DNA methylation in both DKO cell lines infected with DNMT constructs compared to empty vector (E/V) controls (Figure 3B). Even though there was equivalent mRNA expression of exogenous DNMT enzymes in the two DKO clones (Figure S4), DKO8 cells, with higher baseline methylation levels, showed a greater increase in methylation compared to hypomethylated DKO1 cells for each individual construct. Moreover, the increase in methylation in the infected DKO cells was preferentially localized to loci having low-levels of pre-existing methylation and minimal de novo methylation of previously unmodified sites could be observed (De Carvalho D. and Sharma S. et. al., unpublished observations), indicating that DKO cells possess similar patterns of chromatin states as present in the parental WT HCT116 cells, including histone modifications (such as H3K4me3 and H2A.Z etc.) which are involved in guiding DNA methylation to specific genomic loci [6]. These results also indicate a stimulatory effect of pre-existing methylation [32] on DNA methylation by DNMTs in vivo, possibly through stabilization of de novo DNMT3A/3B enzymes on methylated nucleosomes as suggested by their higher protein levels in DKO8 cells (Figure 1B, Figure 3A). This process may further be enhanced by the higher levels of DNMT1 hypomorph present in DKO8 cells [33] (Figure 1B). Within each DKO clone, exogenous DNMT3L expressing cells showed the most robust increase in methylation followed by DNMT3B1 and ΔDNMT3B2 expressing cells respectively, re-emphasizing the strong stimulatory effect of DNMT3L on DNMT3A/3B activity observed in ES cells [12]. These methylation data were further confirmed through Illumina Infinium analysis [34] for each infected cell line (data not shown).
Fig. 3. Increase in DNA methylation restores the DNMT3A protein level in DKO cells. 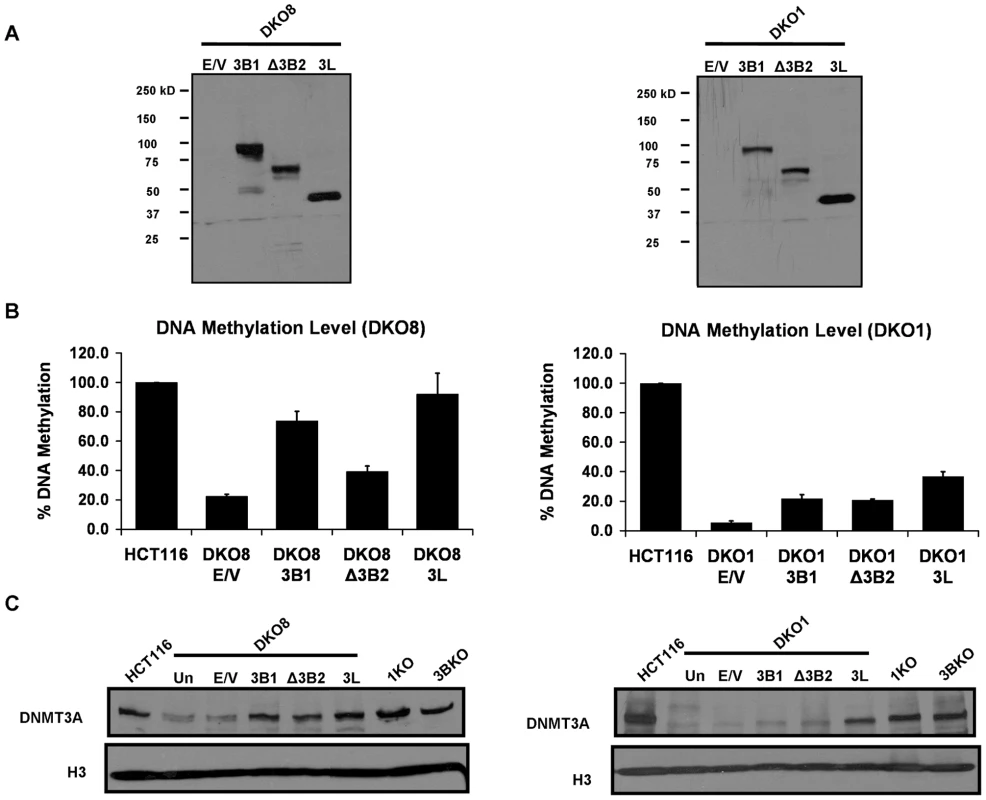
(A) Expression of Myc-tagged DNMT3B1, ΔDNMT3B2 and DNMT3L proteins, infected using a lentiviral system, in DKO cells was confirmed by immunoblotting of nuclear extracts using a Myc antibody. (B) DNA methylation analysis of infected DKO cells using methylation-sensitive restriction enzymes. Genomic DNA was isolated from infected cells eight weeks after infection and methylation level was estimated as described in Figure 1. Data is presented as the percentage of methylation retained compared to WT HCT116 methylation levels. Data represents mean and SEM of three independent replicate experiments. (C) Western blot analysis of nuclear extracts, prepared from infected DKO cells and different HCT116 derivative cell lines, using a DNMT3A antibody. Histone H3 was used as the loading control. Data presented in this figure is representative of two biological replicate experiments. E/V: Empty Vector. Interestingly, immunoblotting of nuclear extracts revealed a substantial transcription-independent increase in DNMT3A protein level in all DNMT infected DKO cell lines (Figure 3C, Figure S5). Moreover, the increase in DNMT3A correlated with the increase in global DNA methylation levels (Figure 3C, 3B). Considering that DNMT3A primarily associates with methylated chromatin regions [22], these data suggest that presence of such methylated regions is required for maintaining its protein level in somatic cells.
DNA methylation–induced DNMT3A increase is mediated by strong anchoring to nucleosomes
To examine whether the increase in DNMT3A protein observed upon restoration of DNA methylation is mediated by binding to nucleosomes, we used sucrose density gradient analysis which allows for the study of in vivo interactions between the chromatin modification enzymes and their actual nucleosomal substrates in the native state [22]. Mononucleosomal digests prepared by extensive micrococcal nuclease (MNase) digestion of nuclei from infected DKO8 cells, expressing either E/V, Myc-tagged DNMT3B1, ΔDNMT3B2 or DNMT3L, were subjected to fractionation on sucrose gradients containing 300 mM NaCl. Western blot analysis showed similar nucleosomal profile in all gradients with mononucleosomes forming a peak at fraction 6 (Figure 4). The DNMT fusion proteins displayed distinct sedimentation profiles indicating different nucleosome binding affinities. DNMT3B1 associated strongly with nucleosomes while the truncated ΔDNMT3B2 variant showed weak association with nucleosomes with a substantial amount of ΔDNMT3B2 sedimenting in nucleosome-free fractions indicating an essential role of the N-terminal region in strong nucleosomal binding, consistent with previous data [22]. However, analysis of various other truncated DNMT3B1 proteins, which contained the N-terminal region but lacked other protein regions (such as the catalytic, PHD and/or PWWP domains), revealed weak nucleosome binding for all truncated proteins (data not shown). These data suggest that DNMT3B requires a full-length protein structure and synergistic activity of its various domains for achieving strong nucleosome binding. DNMT3L showed a bimodal distribution having both nucleosome-free and nucleosome-bound protein (fractions 1–4 and 5–16 respectively). Strikingly, the increased DNMT3A protein in all of the infected cell lines remained strongly associated with nucleosomes similar to that in E/V control, independent of the nucleosome binding affinities of the exogenous proteins, suggesting a nucleosome anchorage dependent stabilization of the protein. DNMT3A formed a peak at fraction 7 in DNMT3B1 and ΔDNMT3B2 expressing cells. In DNMT3L expressing cells, the peak was shifted to fraction 9, indicating the formation of heavier DNMT3A-DNMT3L tetramer encasing the nucleosome [35]. It might also be possible that DNMT3A-DNMT3L tetramer bound nucleosomal regions may be more resistant to MNase digestion and be responsible for this shift. We did not observe any DNMT3A in the nucleosome-free fractions (1–4) co-sedimenting with the unbound pool of ΔDNMT3B2 or DNMT3L fusion proteins (Figure 4), suggesting that the increase in DNMT3A is not due to stabilization through protein-protein interactions with the exogenous proteins but is actually mediated by its binding to nucleosomes upon increase in methylation. Taken together, these data suggest that DNMT3A protein is stabilized by binding to nucleosomes containing its own product (i.e. methylated DNA), which is essential for maintaining its cellular levels.
Fig. 4. The increased level of DNMT3A protein in infected DKO cells, which have increased levels of DNA methylation, remains tightly bound to nucleosomes. 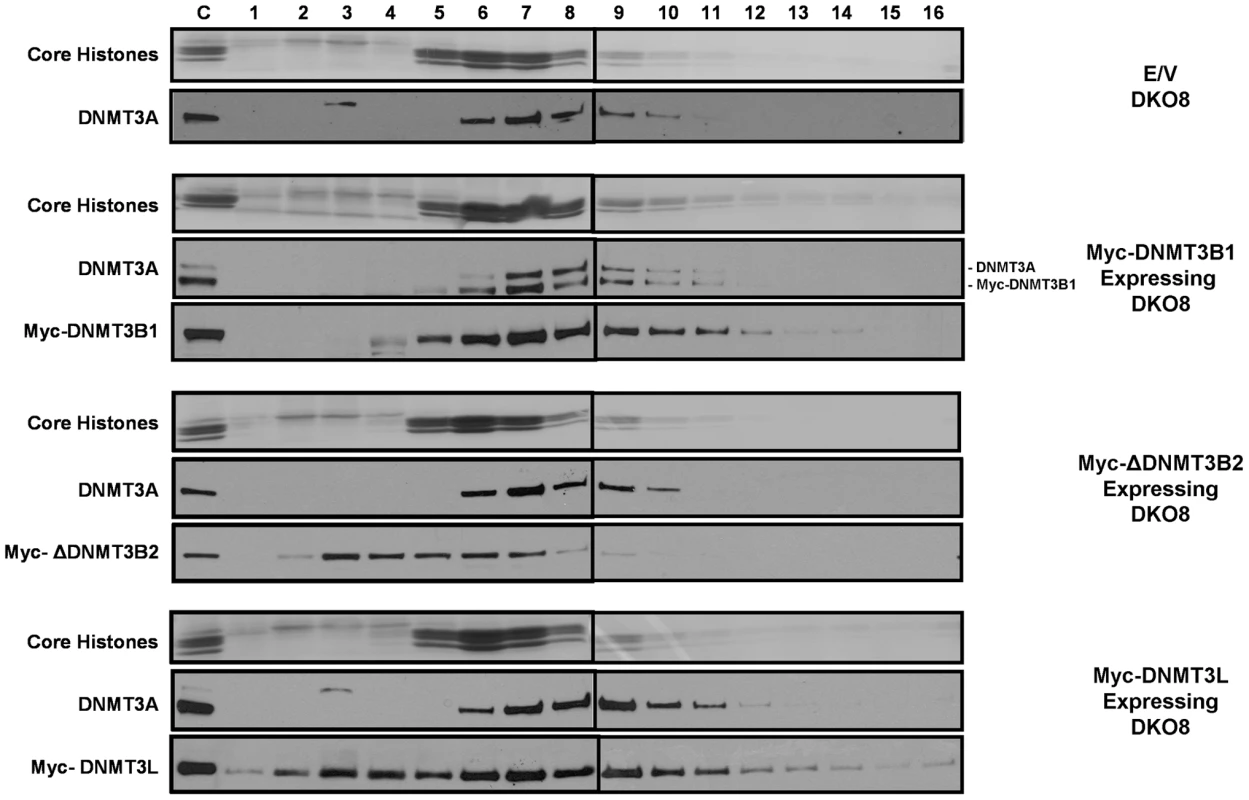
Mononucleosomal digests prepared by extensive MNase digestion of infected DKO8 nuclei, were resolved by ultracentrifugation on a sucrose density gradient (5% to 25%) containing 300 mM NaCl. Gradients were fractioned into 16 aliquots numbered 1–16 starting from the top of the centrifuge tube. To probe the distribution of proteins in each fraction, western blotting was performed with various antibodies after TCA precipitation of proteins from each fraction. Ponceau S staining shows core histones transferred onto the membrane from the SDS/PAGE gel. Mononucleosomes peaked in fraction 6 and the small proportion of higher order oligonucleosomes remaining in the digests sedimented in later fractions. The control lanes on the gels were loaded with unfractionated nuclear extract to monitor the quality of the immunostaining of the membranes. The upper band in the DNMT3A blot for Myc-DNMT3B1 expressing DKO8 cells denotes endogenous DNMT3A. The lower band represents the residual signal of the exogenous Myc-DNMT3B1, which was probed earlier on the same membrane. Reduced nucleosome binding and degradation of unbound DNMT3B upon depletion of DNA methylation
DNMT3B, like DNMT3A, also compartmentalizes to methylated regions in somatic cells via strong anchoring to nucleosomes containing methylated DNA [22], [26]. To examine whether DNMT3B also binds to nucleosomes in a DNA methylation dependent manner, we expressed Myc-tagged DNMT3B1 in three DNMT3B-knockout HCT116 cell lines, 3BKO, DKO8 and DKO1, which possess 86%, 27% and 6% of total genomic DNA methylation respectively (Figure 1C). We first tested mRNA and protein expression of the exogenous DNMT3B1 in these cell lines. Interestingly, while DNMT3B1 mRNA levels were similar in all infected cell lines, we found dramatically reduced DNMT3B1 protein, similar to DNMT3A, in severely hypomethylated DKO1 cells in comparison to 3BKO and DKO8 cells (Figure 5A, 5B).
Fig. 5. Weak nucleosome binding and selective degradation of unbound DNMT3B in the absence of elevated DNA methylation levels. 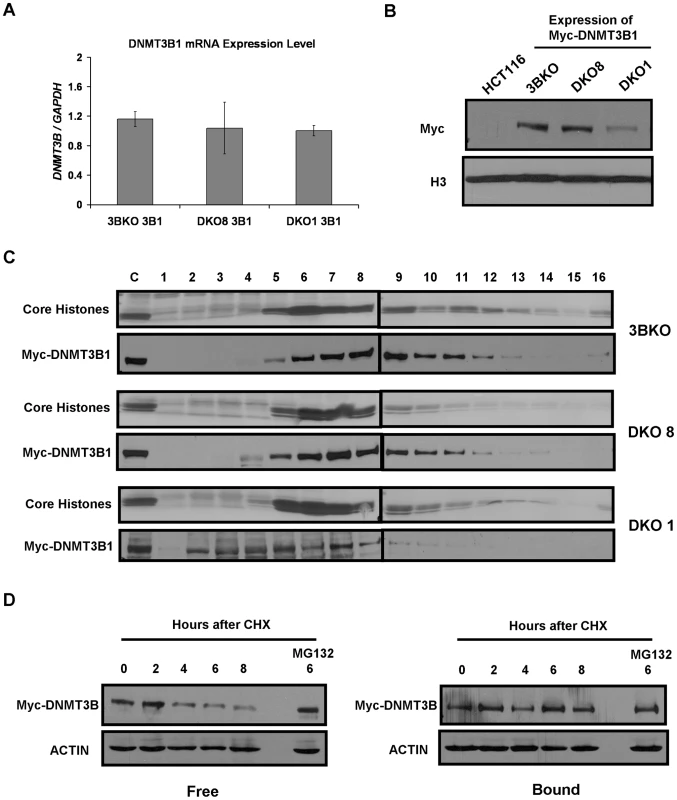
(A) RT-PCR analysis was performed using primers for Myc-DNMT3B1 to assess its mRNA levels in infected 3BKO, DKO8 and DKO1 cells. The results are normalized to GAPDH mRNA levels. Data represents mean and standard deviation of triplicate PCR reactions from a single experiment, representative of two biological replicate experiments. (B) Western blot analysis of nuclear extracts from infected 3BKO, DKO8 and DKO1 cells. Exogenous DNMT3B1 was detected with Myc antibody. (C) Nuclei extracted from infected cells were extensively digested with MNase and mononucleosomes released from them were resolved by ultracentrifugation on a sucrose density gradient (5% to 25%) containing 300 mM NaCl. The gradients were fractionated and analyzed as described previously. (D) DKO1 cells expressing Myc-DNMT3B1 were treated with cycloheximide (CHX) for different time points. The proteosome inhibitor, MG132 was added 2 hr prior to CHX treatment. Nuclei extracted from each sample were then incubated in 500 µl of ice-cold RSB containing 300 mM NaCl, 0.25 M sucrose and protease inhibitors at 4°C for 5 min. Supernatant and nuclear fractions were separated by centrifugation at low speed and equivalent protein amounts from each were subjected to western blot analysis. Data is representative of two biological replicate experiments. To assess whether the decrease in DNMT3B1 resulted from a reduction in binding affinity for nucleosomes in hypomethylated cells, we tested its distribution in mononucleosomal digests fractionated on 300 mM NaCl containing sucrose gradients. In 3BKO and DKO8 cells, the exogeneous DNMT3B1 showed strong association with nucleosomes similar to endogeneous DNMT3A (Figure 5C, Figure 4). However, DNMT3B1 weakly associated with nucleosomes in severely hypomethylated DKO1 cells with the bulk of the overexpressed protein sedimenting in nucleosome-free fractions (2–4), suggesting a dramatic reduction in nucleosome binding affinity upon depletion of DNA methylation. Since the increase in methylation in the infected DKO cells was preferentially localized to the same loci which were originally methylated in the parental HCT116 cells, it suggests that DKO cells possess similar patterns of histone modifications involved in guiding DNA methylation as present in WT HCT116 cells (De Carvalho D. and Sharma S. et. al., unpublished observations). Therefore, taken together, these data strongly suggest that the reduction in nucleosome binding affinity of DNMT3A/3B observed in DKO cells results from depletion of DNA methylation and is not due to clonal variation of global chromatin states.
To further confirm this phenomenon, we subjected Myc-tagged DNMT3B1 expressing DKO1 cells to CHX treatment and analyzed protein stability of the nucleosome-bound and -free fractions of DNMT3B1 protein. Consistent with our previous data on endogenous DNMT3A enzyme, the overexpressed free DNMT3B1 protein underwent rapid degradation compared to the stable nucleosome-bound DNMT3B1 protein, clearly displaying the instability of the unbound protein (Figure 5D). Such degradation was inhibited by treatment with the proteosome inhibitor MG132, indicating the role of proteosomal pathway in this process. However, we could not rescue degradation of DNMT3A protein in DKO cells using MG132 treatment suggesting possible involvement of other mechanisms in its degradation (data not shown). The phenomenon of destabilization and selective degradation of unbound DNMT3A/3B proteins could also be observed in the case of Myc-ΔDNMT3B2 which showed substantially lower protein levels compared to Myc-DNMT3B1 in DKO8 cells even when both genes were expressed at similar mRNA levels (Figure 3A, Figure S4). Since Myc-ΔDNMT3B2 associated weakly with nucleosomes while Myc-DNMT3B1 bound strongly to nucleosomes in DKO8 cells (Figure 4), the reduction in Myc-ΔDNMT3B2 levels possibly results from a decrease in protein stability of the unbound Myc-ΔDNMT3B2 protein, similar to that previously observed for DNMT3A and 3B in DKO1 cells. Taken together, these data show that both DNMT3A/3B require the presence of DNA methylation for tight binding to nucleosomes and subsequent protein stabilization. Such a mechanism would enable faithful inheritance of methylated states through proper compartmentalization of DNMT3A/3B while preventing spurious de novo methylation through selective degradation of the free enzymes.
Synergistic activity of DNMT3A/3B is mediated by their anchoring to nucleosomes
In ES cells, DNMT3A/3B strongly interact and mutually stimulate each other's activity, thus working synergistically to establish genomic DNA methylation patterns during development [36]. To ascertain whether a similar mechanism is involved in propagation of DNA methylation in somatic cells, we expressed a Myc-tagged catalytically-inactive DNMT3B1 mutant, having a cysteine to serine alteration (position 657) which destroys catalytic activity without compromising other functions [37], in DKO8 cells and confirmed its protein expression by immunoblotting (Figure 6A). To determine whether the DNMT3B1 mutant could stimulate DNMT3A activity, we measured the global DNA methylation level in the mutant expressing cells 8 weeks post-infection. We observed a substantial increase in methylation, demonstrating a stimulatory effect of DNMT3B on DNMT3A activity, independent of catalytic activity (Figure 6B). Immunoprecipitation experiments showed that the mutant DNMT3B1 strongly interacted with DNMT3A, similar to WT DNMT3B1, suggesting a DNMT3L-like stimulation mechanism which occurs through physical interaction of the two proteins [12] (Figure S6). Along with an increase in DNA methylation, we observed a substantial increase in endogenous DNMT3A protein levels in mutant DNMT3B1 expressing cells (Figure 6A), similar to WT DNMT3B1 expressing cells, suggesting DNA methylation induced stabilization of DNMT3A protein. Similar results were obtained upon expression of a catalytically-inactive ΔDNMT3B2 mutant in DKO8 cells indicating a stimulatory effect of ΔDNMT3B2 on DNMT3A activity, occurring through its physical interaction with the DNMT3A protein as shown by immunoprecipitation experiments (Figures S7, S6).
Fig. 6. DNMT3B catalytically-inactive mutant stimulates DNA methylation by increasing DNMT3A binding to nucleosomes. 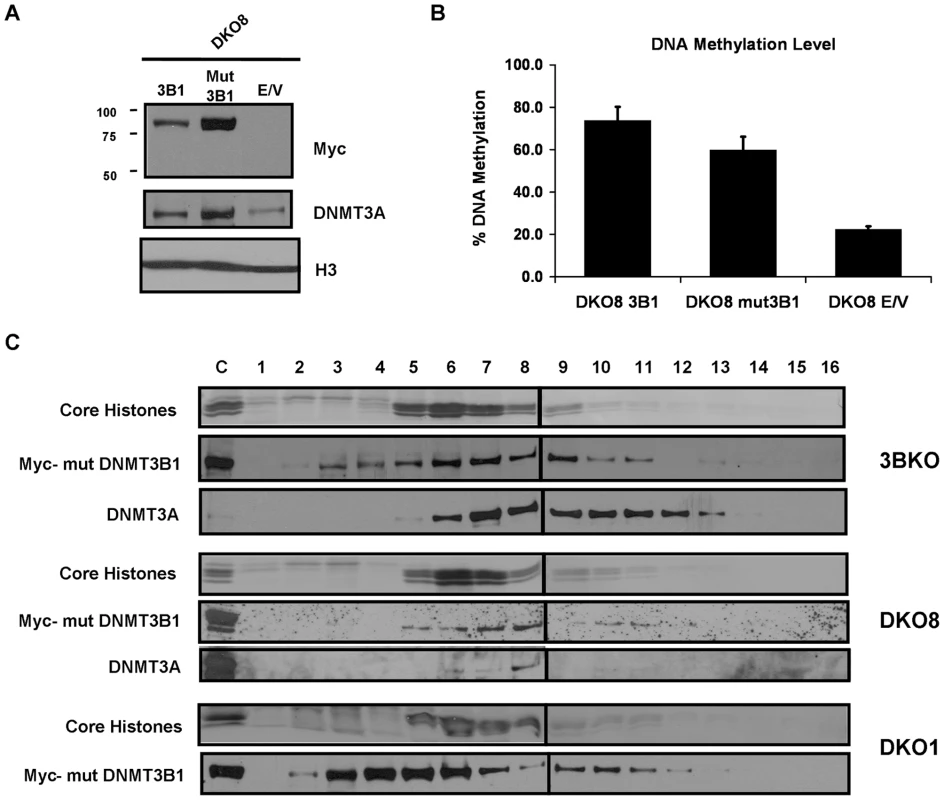
In ES cells, stimulation of DNMT3A/3B activity by DNMT3L partially occurs through increased association of the enzymes with the substrate DNA, allowing these slow acting enzymes to efficiently methylate the substrate [38]. To examine whether stimulation of DNMT3A by DNMT3B in somatic cells occurs through a similar mechanism in a nucleosomal context, we analyzed mononucleosomal digests from DNMT3B1 mutant expressing cells on 300 mM sucrose density gradients. All cellular DNMT3A in infected 3BKO and DKO8 cell lines was found tightly anchored to nucleosomes suggesting that its stimulation by DNMT3B1 occurs through an increased binding to nucleosomes (Figure 6C). We could not detect DNMT3A in DKO1 cells in this assay due to its extremely low levels. In DKO8 cells expressing the mutant ΔDNMT3B2, all cellular DNMT3A protein was found strongly anchored to nucleosomes indicating that the interaction and stimulation of DNMT3A by ΔDNMT3B2 is mediated by their binding to nucleosomes (Figure S7). The DNMT3B1 and ΔDNMT3B2 mutants displayed similar binding affinity for nucleosomes as their WT counterparts suggesting that their catalytic activity has little role in nucleosome binding. Taken together, these data show that in vivo stimulation of DNMT3A by DNMT3B occurs through an increased binding to nucleosomes, similar to that observed with DNMT3L, enabling efficient methylation from these slow acting de novo enzymes and their consequent stabilization through continued association with such methylated regions.
Discussion
Proper maintenance of epigenetic modifications within specific chromatin domains is critical for preserving cellular identity. Recently, a common theme for inheritance of histone marks has emerged where the mark recruits and retains its own modifying enzyme and triggers renewal by stimulating that enzyme through possible allosteric activation mechanisms [39], [40], [41]. Our work suggests involvement of a similar mechanism in maintenance of DNA methylation patterns through DNMT3A/3B in somatic cells.
We and others have previously shown that DNMT3A/3B, but not DNMT1, are strongly anchored to nucleosomes containing methylated DNA in somatic cells [22], [26]. Our current data shows that the presence of DNA methylation is essential for association of DNMT3A/3B with chromatin and also for maintaining the cellular levels of the DNMT3A/3B enzymes, thereby creating a homeostatic inheritance system. Such methylation directed binding stimulates DNA methylation at target loci in vivo ensuring faithful maintenance of methylation patterns, a phenomenon previously observed in inheritance of the polycomb mark [42]. Since DNMT3A/3B are slow acting enzymes compared to DNMT1 [38], stable association with their target methylated regions would be key for their ability to properly maintain methylated states. We further show that DNMT3A/3B work synergistically in this maintenance process and DNMT3B stimulates DNMT3A activity through increased association with nucleosomes, similar to DNMT3L. Thus, promotion of DNA methylation by selective binding of DNMT3A/3B to nucleosomes containing pre-existing methylation may serve as a critical positive feed-back loop mechanism essential for faithful propagation of epigenetic states through somatic cell divisions [25], [43], [44].
Another key finding of our work is the selective degradation of free DNMT3A/3B proteins which could not bind to chromatin in the absence of pre-existing DNA methylation in somatic cells. In ES cells and PGCs (primordial germ cells), DNMT3A/3B are required for establishment of global DNA methylation patterns. Therefore, in these cells, DNMT3A/3B are highly expressed at the transcriptional level and their methylation activity is strongly stimulated by DNMT3L [12], [45]. However, in somatic cells, the main role of the de novo DNMT3A/3B enzymes is to assist DNMT1 in proper maintenance of pre-established DNA methylation patterns and prevention of de novo methylation of previously unmethylated regions is required [11], [45]. Therefore, DNMT3A/3B mRNA expression is substantially downregulated and DNMT3L is not expressed in differentiated somatic tissues in order to prevent any aberrant de novo methylation [20], [45]. Our data suggest that to further regulate this maintenance process, DNMT3A/3B protein levels are post-translationally regulated by the levels of pre-existing DNA methylation in somatic cells. Selective degradation of free DNMT3A/3B enzymes may help explain how somatic cells, which still express low levels of de novo DNMT3A/3B enzymes, prevent aberrant de novo methylation of CpG islands. Our data suggests that once DNMT3A/3B are recruited to methylated chromatin domains, pre-existing methylation stabilizes their binding to such regions and enables faithful propagation of methylated states. However, in absence of DNA methylation, as would be the case with unmethylated CpG islands, these slow acting enzymes are unable to stably bind to the chromatin. The resulting free de novo enzymes, which could potentially cause spurious methylation, are then selectively degraded by the cellular machinery possibly through recognition of an altered conformation in the unbound state (Figure 7). As shown in the model, in hypomethylated DKO (DNMT1ΔE2-5/DNMT3B−/−) cells, DNMT3A loses its ability to bind to nucleosomes resulting in destabilization and selective degradation of free DNMT3A protein while the residual DNMT3A remains bound to the remaining few methylated chromatin regions (Figure 7). Since exogenous Myc-DNMT3B1 also displayed a similar DNA methylation-dependent stabilization upon nucleosomes, it suggests that a similar model might apply to the regulation of DNMT3B enzyme in somatic cells. Our data indicates that unbound DNMT3B1 is degraded through the proteosomal pathway but how DNMT3A is selectively targeted for degradation in somatic cells is still unclear. Future studies are required to further understand the exact mechanisms involved in the selective degradation of unbound DNMT3A/3B enzymes. Histone methyltransferases, however, are not regulated in such a manner and have been found to exist in both free and chromatin-bound forms within nuclei. This difference can be partially explained by the fact that histone marks are far more dynamic in nature, actively regulated by the combined action of histone methyltransferases and demethylases [46], compared to DNA methylation which is still believed to be a relatively stable mark in differentiated tissues [6].
Fig. 7. Model for selective stabilization of DNMT3A/3B through anchoring to nucleosomes containing methylated DNA. 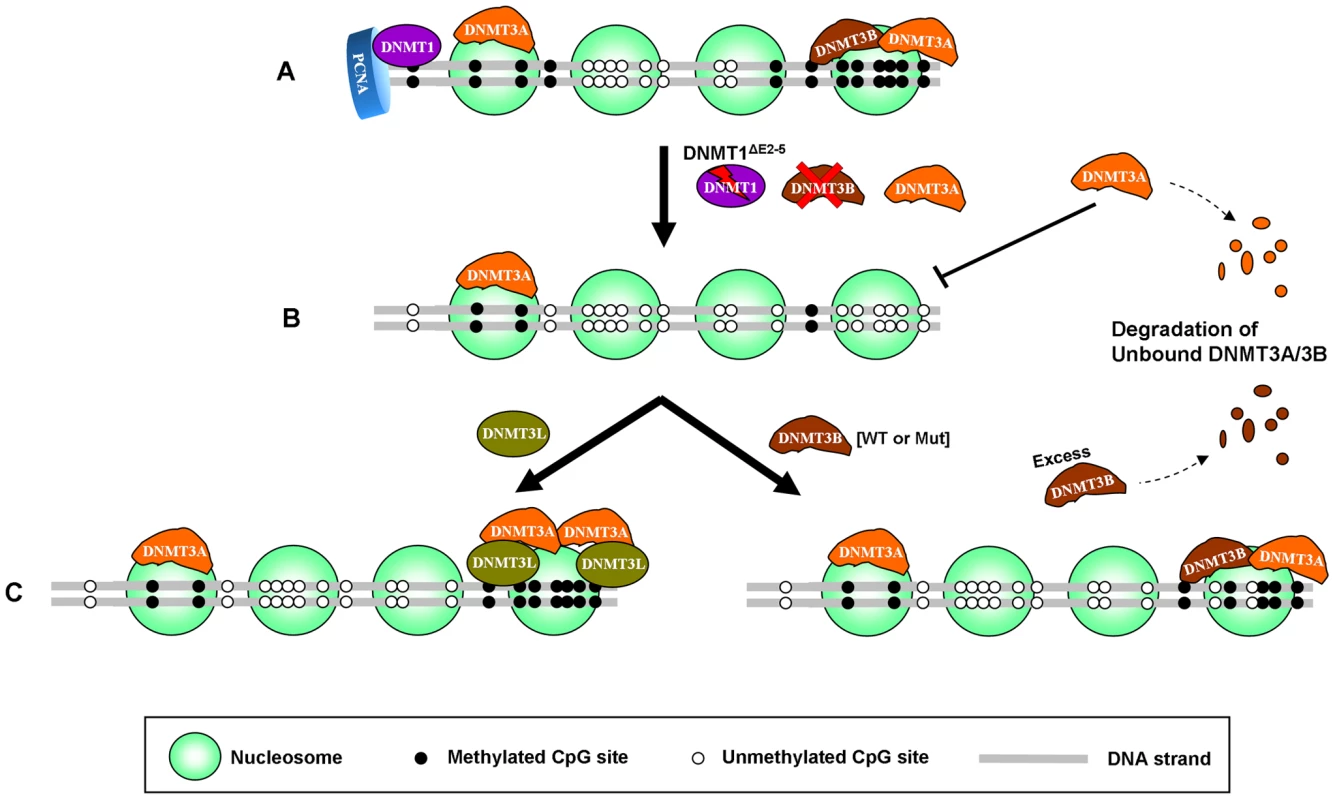
(A) In somatic cells, DNMT3A/3B remain bound to nucleosomes containing methylated DNA, enabling proper maintenance of methylated states in co-operation with DNMT1, the maintenance enzyme, which copies the methylation pattern during replication by associating with the proliferating cell nuclear antigen (PCNA). (B) When DNA methylation is lowered by genetic disruption of DNMT1 and DNMT3B in DKO cells, DNMT3A loses its ability to bind to nucleosomes which results in destabilization and subsequent degradation of the protein. (C) Restoration of DNA methylation in such hypomethylated cells, through expression of exogenous DNMT3B (WT or mut) or DNMT3L, increases DNMT3A protein levels by enabling it to bind to nucleosomes again which results in stabilization of DNMT3A protein. Exogenous DNMT3B (WT or mut) also binds strongly to nucleosomes in the presence of DNA methylation and synergistically increases methylation along with DNMT3A while the excess free DNMT3B protein, which could not anchor to the nucleosomes, gets degraded by the proteosomal machinery. mut: mutant. While initial recruitment of DNMT3A/3B to methylated regions may involve other proteins, our data strongly suggests that their anchoring to chromatin primarily depends upon pre-existing DNA methylation. However, in addition to DNA methylation, certain histone modifications and accessory proteins may also help in selective compartmentalization of these enzymes. For instance, unmethylated H3K4, recently shown to bind DNMT3A [47], may assist in stable binding to silent domains. Recruitment of DNMT3A/3B to such domains may involve UHRF1 [19]. On the other hand, proteins like H2A.Z, CTCF and H3K4me3 etc. which are antagonistic to DNA methylation [14], [48], [49], may occlude binding of DNMT3A/3B to active/poised regions, thus constraining their activities to silent methylated domains only. Recently, Witcher and Emerson [50] have shown that loss of such boundary elements indeed results in aberrant spreading of DNA methylation beyond methylated domains. Our data suggests that these aberrations may involve DNMT3A/3B enzymes which remain bound to methylated regions [22]. During tumorigenesis, these de novo enzymes may progressively override the chromatin boundaries, gradually spreading methylation beyond their specific domains to the entire region [51] resulting in aberrant methylation of genes in clusters – a common feature of cancer-specific hypermethylation [52], [53]. Such a mechanism may also help explain why CpG island loci having pre-existing methylation in a normal tissue are more susceptible to undergo de novo methylation in cancer [54]. Moreover, ectopic de novo methylation, correlated with overexpression of DNMT3A/3B in several types of cancer [3], may also be maintained and propagated through continued association of DNMT3A/3B with such regions. DNA methylation inhibitors like 5-aza-CdR, widely used to inhibit aberrant methylation in cancer, target DNMTs by trapping them on DNA [55]. Since these hypomethylating drugs trap DNMTs onto the DNA, it is not feasible to use them for studying the dissociation of DNMT3A/3B from nucleosomes and destabilization upon loss in DNA methylation observed in our experiments. Nevertheless, our data suggests that destabilization of DNMT3A/3B upon removal of DNA methylation may provide another mechanism for depletion of these enzymes upon treatment with such hypomethylating drugs. However, future studies are required to further understand these mechanisms, focusing on factors determining proper compartmentalization of DNMT3A/3B to methylated regions and mechanisms responsible for selective degradation of the unbound protein.
In conclusion, our data suggests a model for epigenetic inheritance of DNA methylation in somatic tissues where pre-existing methylation triggers its renewal by recruiting and stabilizing DNMT3A/3B on methylated chromatin domains, which then work synergistically to propagate DNA methylation in co-operation with DNMT1. Such a mechanism not only ensures faithful maintenance of methylated states but also guards against aberrant methylation from the de novo DNMT3A/3B enzymes.
Materials and Methods
Cell culture and drug treatment
HCT116 derivative cell lines were maintained in McCoy's 5A medium containing 10% inactivated fetal bovine serum, 100 units/ml penicillin and 100 µg/ml streptomycin. Puromycin was included in the culture medium at 3 µg/ml to maintain infected HCT116 derivative cell lines. When indicated, cycloheximide (Sigma) was added to a final concentration of 50 µg/ml. The proteosome inhibitor MG132 (Calbiochem) was used at 10 µM for 2 h prior to CHX treatment.
RNA isolation and RT-PCR
Detailed methods are described in Text S1.
Expression vector construction
Human 3B1, ΔDNMT3B2 and DNMT3L cDNA sequences having the Myc tag DNA sequence ligated to their 5′ ends were amplified from the pIRESpuro/Myc constructs [22] (a modified version of the pIRESpuro3 vector, Clontech), a generous gift from Allen Yang (USC), using polymerase chain reaction (PCR). Myc-tagged catalytically-inactive mutants of DNMT3B1 and ΔDNMT3B2, having a cysteine to serine alteration in the catalytic domain corresponding to position 657 of DNMT3B1 protein, were prepared using a site-directed mutagenesis kit (Stratagene). The mutation was confirmed by sequencing both strands of the constructs. For preparation of the constructs, the lentivirus vector pLJM1 was linearized using AgeI and EcoRI restriction enzymes and the Myc tagged DNMT cDNAs were cloned in it using In-fusion advantage PCR cloning kit (Clontech) following manufacturer's protocol. For lentivirus production, the vesicular stomatitis virus envelope protein G expression construct pMD.G1, the packaging vector pCMV ΔR8.91 and the transfer vector pLJM1 were used as described previously [56]. Infected HCT116 derivative cells, stably expressing various DNMTs, were selected in the presence of 3 µg/ml puromycin for three weeks.
Nuclear and whole-cell lysates preparation
Detailed methods are described in Text S1.
Salt extraction of nuclei
Nuclei from 5×106 cells were incubated in 500 µl of ice-cold RSB containing 0.25 M sucrose, protease inhibitors and various concentrations of NaCl for 5 min at 4°C. Nuclei were then harvested by microcentrifugation, separating the supernatant and the pellet fractions. Nuclear pellets were resuspended in RIPA buffer and subjected to sonication. Proteins in the supernatant were concentrated using TCA precipitation and later resuspended in RIPA buffer. Equivalent volumes of supernatant and pellet fractions were added to SDS loading buffer and subjected to Western blotting.
MNase digestion and sucrose density gradient centrifugation
MNase digestion and sucrose gradients experiments were performed as described previously [22]. For details, see Text S1.
DNA methylation analysis
Genomic DNA (10 µg) isolated from various HCT116 derivative cell lines was digested with methylation-sensitive restriction enzymes, HpaII or MspI (New England Biolabs), at 37°C over night. The digested DNA was run on an agarose gel at low voltage for 8 hrs in order to achieve good separation. The undigested DNA band in each lane was then quantified using the ImageQuant software. Percentage of genomic methylation present was calculated using the formula: where H = undigested with HpaII; M = undigested with MspI and G = genomic DNA.
Supporting Information
Zdroje
1. BirdA
2002 DNA methylation patterns and epigenetic memory. Genes Dev 16 6 21
2. SuzukiMM
BirdA
2008 DNA methylation landscapes: provocative insights from epigenomics. Nat Rev Genet 9 465 476
3. JonesPA
BaylinSB
2007 The epigenomics of cancer. Cell 128 683 692
4. SharmaS
KellyTK
JonesPA
2010 Epigenetics in cancer. Carcinogenesis 31 27 36
5. OkanoM
BellDW
HaberDA
LiE
1999 DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99 247 257
6. LawJA
JacobsenSE
Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet 11 204 220
7. RheeI
BachmanKE
ParkBH
JairKW
YenRW
2002 DNMT1 and DNMT3b cooperate to silence genes in human cancer cells. Nature 416 552 556
8. LiangG
ChanMF
TomigaharaY
TsaiYC
GonzalesFA
2002 Cooperativity between DNA methyltransferases in the maintenance methylation of repetitive elements. Mol Cell Biol 22 480 491
9. ChenT
UedaY
DodgeJE
WangZ
LiE
2003 Establishment and maintenance of genomic methylation patterns in mouse embryonic stem cells by Dnmt3a and Dnmt3b. Mol Cell Biol 23 5594 5605
10. RiggsAD
XiongZ
2004 Methylation and epigenetic fidelity. Proc Natl Acad Sci U S A 101 4 5
11. JonesPA
LiangG
2009 Rethinking how DNA methylation patterns are maintained. Nat Rev Genet 10 805 811
12. GowherH
LiebertK
HermannA
XuG
JeltschA
2005 Mechanism of stimulation of catalytic activity of Dnmt3A and Dnmt3B DNA-(cytosine-C5)-methyltransferases by Dnmt3L. J Biol Chem 280 13341 13348
13. OoiSK
QiuC
BernsteinE
LiK
JiaD
2007 DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature 448 714 717
14. OkitsuCY
HsiehCL
2007 DNA methylation dictates histone H3K4 methylation. Mol Cell Biol 27 2746 2757
15. WeberM
HellmannI
StadlerMB
RamosL
PaaboS
2007 Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nat Genet 39 457 466
16. LehnertzB
UedaY
DerijckAA
BraunschweigU
Perez-BurgosL
2003 Suv39h-mediated histone H3 lysine 9 methylation directs DNA methylation to major satellite repeats at pericentric heterochromatin. Curr Biol 13 1192 1200
17. Epsztejn-LitmanS
FeldmanN
Abu-RemailehM
ShufaroY
GersonA
2008 De novo DNA methylation promoted by G9a prevents reprogramming of embryonically silenced genes. Nat Struct Mol Biol 15 1176 1183
18. SharifJ
MutoM
TakebayashiS
SuetakeI
IwamatsuA
2007 The SRA protein Np95 mediates epigenetic inheritance by recruiting Dnmt1 to methylated DNA. Nature 450 908 912
19. MeilingerD
FellingerK
BultmannS
RothbauerU
BonapaceIM
2009 Np95 interacts with de novo DNA methyltransferases, Dnmt3a and Dnmt3b, and mediates epigenetic silencing of the viral CMV promoter in embryonic stem cells. EMBO Rep 10 1259 1264
20. AapolaU
KawasakiK
ScottHS
OllilaJ
VihinenM
2000 Isolation and initial characterization of a novel zinc finger gene, DNMT3L, on 21q22.3, related to the cytosine-5-methyltransferase 3 gene family. Genomics 65 293 298
21. Bourc'hisD
XuGL
LinCS
BollmanB
BestorTH
2001 Dnmt3L and the establishment of maternal genomic imprints. Science 294 2536 2539
22. JeongS
LiangG
SharmaS
LinJC
ChoiSH
2009 Selective anchoring of DNA methyltransferases 3A and 3B to nucleosomes containing methylated DNA. Mol Cell Biol 29 5366 5376
23. LinkPA
GangisettyO
JamesSR
Woloszynska-ReadA
TachibanaM
2009 Distinct roles for histone methyltransferases G9a and GLP in cancer germ-line antigen gene regulation in human cancer cells and murine embryonic stem cells. Mol Cancer Res 7 851 862
24. KondoY
ShenL
AhmedS
BoumberY
SekidoY
2008 Downregulation of histone H3 lysine 9 methyltransferase G9a induces centrosome disruption and chromosome instability in cancer cells. PLoS ONE 3 e2037 doi:10.1371/journal.pone.0002037
25. BachmanKE
RountreeMR
BaylinSB
2001 Dnmt3a and Dnmt3b are transcriptional repressors that exhibit unique localization properties to heterochromatin. J Biol Chem 276 32282 32287
26. SchlesingerY
StraussmanR
KeshetI
FarkashS
HechtM
2007 Polycomb-mediated methylation on Lys27 of histone H3 pre-marks genes for de novo methylation in cancer. Nat Genet 39 232 236
27. RheeI
JairKW
YenRW
LengauerC
HermanJG
2000 CpG methylation is maintained in human cancer cells lacking DNMT1. Nature 404 1003 1007
28. EggerG
JeongS
EscobarSG
CortezCC
LiTW
2006 Identification of DNMT1 (DNA methyltransferase 1) hypomorphs in somatic knockouts suggests an essential role for DNMT1 in cell survival. Proc Natl Acad Sci U S A 103 14080 14085
29. VegaFM
SevillaA
LazoPA
2004 p53 Stabilization and accumulation induced by human vaccinia-related kinase 1. Mol Cell Biol 24 10366 10380
30. WangL
WangJ
SunS
RodriguezM
YueP
2006 A novel DNMT3B subfamily, DeltaDNMT3B, is the predominant form of DNMT3B in non-small cell lung cancer. Int J Oncol 29 201 207
31. JairKW
BachmanKE
SuzukiH
TingAH
RheeI
2006 De novo CpG island methylation in human cancer cells. Cancer Res 66 682 692
32. KimGD
NiJ
KelesogluN
RobertsRJ
PradhanS
2002 Co-operation and communication between the human maintenance and de novo DNA (cytosine-5) methyltransferases. EMBO J 21 4183 4195
33. VilkaitisG
SuetakeI
KlimasauskasS
TajimaS
2005 Processive methylation of hemimethylated CpG sites by mouse Dnmt1 DNA methyltransferase. J Biol Chem 280 64 72
34. Laird PW Principles and challenges of genome-wide DNA methylation analysis. Nat Rev Genet 11 191 203
35. JiaD
JurkowskaRZ
ZhangX
JeltschA
ChengX
2007 Structure of Dnmt3a bound to Dnmt3L suggests a model for de novo DNA methylation. Nature 449 248 251
36. LiJY
PuMT
HirasawaR
LiBZ
HuangYN
2007 Synergistic function of DNA methyltransferases Dnmt3a and Dnmt3b in the methylation of Oct4 and Nanog. Mol Cell Biol 27 8748 8759
37. HsiehCL
1999 In vivo activity of murine de novo methyltransferases, Dnmt3a and Dnmt3b. Mol Cell Biol 19 8211 8218
38. YokochiT
RobertsonKD
2002 Preferential methylation of unmethylated DNA by Mammalian de novo DNA methyltransferase Dnmt3a. J Biol Chem 277 11735 11745
39. HansenKH
BrackenAP
PasiniD
DietrichN
GehaniSS
2008 A model for transmission of the H3K27me3 epigenetic mark. Nat Cell Biol 10 1291 1300
40. FelsenfeldG
GroudineM
2003 Controlling the double helix. Nature 421 448 453
41. CollinsRE
NorthropJP
HortonJR
LeeDY
ZhangX
2008 The ankyrin repeats of G9a and GLP histone methyltransferases are mono - and dimethyllysine binding modules. Nat Struct Mol Biol 15 245 250
42. MargueronR
JustinN
OhnoK
SharpeML
SonJ
2009 Role of the polycomb protein EED in the propagation of repressive histone marks. Nature 461 762 767
43. ChodavarapuRK
FengS
BernatavichuteYV
ChenPY
StroudH
Relationship between nucleosome positioning and DNA methylation. Nature
44. MargueronR
ReinbergD
Chromatin structure and the inheritance of epigenetic information. Nat Rev Genet 11 285 296
45. OkanoM
XieS
LiE
1998 Cloning and characterization of a family of novel mammalian DNA (cytosine-5) methyltransferases. Nat Genet 19 219 220
46. ShiY
WhetstineJR
2007 Dynamic regulation of histone lysine methylation by demethylases. Mol Cell 25 1 14
47. OtaniJ
NankumoT
AritaK
InamotoS
AriyoshiM
2009 Structural basis for recognition of H3K4 methylation status by the DNA methyltransferase 3A ATRX-DNMT3-DNMT3L domain. EMBO Rep 10 1235 1241
48. ZilbermanD
Coleman-DerrD
BallingerT
HenikoffS
2008 Histone H2A.Z and DNA methylation are mutually antagonistic chromatin marks. Nature 456 125 129
49. ZlatanovaJ
CaiafaP
2009 CTCF and its protein partners: divide and rule? J Cell Sci 122 1275 1284
50. WitcherM
EmersonBM
2009 Epigenetic silencing of the p16(INK4a) tumor suppressor is associated with loss of CTCF binding and a chromatin boundary. Mol Cell 34 271 284
51. GraffJR
HermanJG
MyohanenS
BaylinSB
VertinoPM
1997 Mapping patterns of CpG island methylation in normal and neoplastic cells implicates both upstream and downstream regions in de novo methylation. J Biol Chem 272 22322 22329
52. KeshetI
SchlesingerY
FarkashS
RandE
HechtM
2006 Evidence for an instructive mechanism of de novo methylation in cancer cells. Nat Genet 38 149 153
53. CoolenMW
StirzakerC
SongJZ
StathamAL
KassirZ
Consolidation of the cancer genome into domains of repressive chromatin by long-range epigenetic silencing (LRES) reduces transcriptional plasticity. Nat Cell Biol 12 235 246
54. HuangTH
PerryMR
LauxDE
1999 Methylation profiling of CpG islands in human breast cancer cells. Hum Mol Genet 8 459 470
55. EggerG
LiangG
AparicioA
JonesPA
2004 Epigenetics in human disease and prospects for epigenetic therapy. Nature 429 457 463
56. OuCY
KimJH
YangCK
StallcupMR
2009 Requirement of cell cycle and apoptosis regulator 1 for target gene activation by Wnt and beta-catenin and for anchorage-independent growth of human colon carcinoma cells. J Biol Chem 284 20629 20637
Štítky
Genetika Reprodukční medicína
Článek Break to Make a ConnectionČlánek A New Testing Strategy to Identify Rare Variants with Either Risk or Protective Effect on DiseaseČlánek The Architecture of Gene Regulatory Variation across Multiple Human Tissues: The MuTHER Study
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2011 Číslo 2- Akutní intermitentní porfyrie
- Farmakogenetické testování pomáhá předcházet nežádoucím efektům léčiv
- Růst a vývoj dětí narozených pomocí IVF
- Pilotní studie: stres a úzkost v průběhu IVF cyklu
- Intrauterinní inseminace a její úspěšnost
-
Všechny články tohoto čísla
- Break to Make a Connection
- A New Testing Strategy to Identify Rare Variants with Either Risk or Protective Effect on Disease
- Single-Tissue and Cross-Tissue Heritability of Gene Expression Via Identity-by-Descent in Related or Unrelated Individuals
- Pervasive Adaptive Protein Evolution Apparent in Diversity Patterns around Amino Acid Substitutions in
- The Architecture of Gene Regulatory Variation across Multiple Human Tissues: The MuTHER Study
- MiRNA Control of Vegetative Phase Change in Trees
- New Functions of Ctf18-RFC in Preserving Genome Stability outside Its Role in Sister Chromatid Cohesion
- Genome-Wide Association Studies of the PR Interval in African Americans
- Mapping of the Disease Locus and Identification of As a Candidate Gene in a Canine Model of Primary Open Angle Glaucoma
- Mapping a New Spontaneous Preterm Birth Susceptibility Gene, , Using Linkage, Haplotype Sharing, and Association Analysis
- A Population Genetic Approach to Mapping Neurological Disorder Genes Using Deep Resequencing
- and Genes Modulate the Switch between Attraction and Repulsion during Behavioral Phase Change in the Migratory Locust
- Targeted Sister Chromatid Cohesion by Sir2
- Correlated Evolution of Nearby Residues in Drosophilid Proteins
- Parallel Evolution of a Type IV Secretion System in Radiating Lineages of the Host-Restricted Bacterial Pathogen
- Lipophorin Receptors Mediate the Uptake of Neutral Lipids in Oocytes and Imaginal Disc Cells by an Endocytosis-Independent Mechanism
- Genome-Wide Association Study of Coronary Heart Disease and Its Risk Factors in 8,090 African Americans: The NHLBI CARe Project
- The Evolution of Host Specialization in the Vertebrate Gut Symbiont
- Genome-Wide Association of Familial Late-Onset Alzheimer's Disease Replicates and and Nominates in Interaction with
- Risk Alleles for Systemic Lupus Erythematosus in a Large Case-Control Collection and Associations with Clinical Subphenotypes
- Association between Common Variation at the Locus and Changes in Body Mass Index from Infancy to Late Childhood: The Complex Nature of Genetic Association through Growth and Development
- AID Induces Double-Strand Breaks at Immunoglobulin Switch Regions and Causing Chromosomal Translocations in Yeast THO Mutants
- A Study of CNVs As Trait-Associated Polymorphisms and As Expression Quantitative Trait Loci
- Whole-Genome Comparison Reveals Novel Genetic Elements That Characterize the Genome of Industrial Strains of
- Prevalence of Epistasis in the Evolution of Influenza A Surface Proteins
- Srf1 Is a Novel Regulator of Phospholipase D Activity and Is Essential to Buffer the Toxic Effects of C16:0 Platelet Activating Factor
- Two Frizzled Planar Cell Polarity Signals in the Wing Are Differentially Organized by the Fat/Dachsous Pathway
- Phosphoinositide Regulation of Integrin Trafficking Required for Muscle Attachment and Maintenance
- Pathogenic VCP/TER94 Alleles Are Dominant Actives and Contribute to Neurodegeneration by Altering Cellular ATP Level in a IBMPFD Model
- Meta-Analysis of Genome-Wide Association Studies in Celiac Disease and Rheumatoid Arthritis Identifies Fourteen Non-HLA Shared Loci
- A Genome-Wide Study of DNA Methylation Patterns and Gene Expression Levels in Multiple Human and Chimpanzee Tissues
- Nucleosomes Containing Methylated DNA Stabilize DNA Methyltransferases 3A/3B and Ensure Faithful Epigenetic Inheritance
- Mutations in Zebrafish Result in Adult-Onset Ocular Pathogenesis That Models Myopia and Other Risk Factors for Glaucoma
- [], the Prion Formed by the Chromatin Remodeling Factor Swi1, Is Highly Sensitive to Alterations in Hsp70 Chaperone System Activity
- Characterization of Transcriptome Remodeling during Cambium Formation Identifies and As Opposing Regulators of Secondary Growth
- The Cardiac Transcription Network Modulated by Gata4, Mef2a, Nkx2.5, Srf, Histone Modifications, and MicroRNAs
- Epistatic Interaction Maps Relative to Multiple Metabolic Phenotypes
- Quantitative Models of the Mechanisms That Control Genome-Wide Patterns of Transcription Factor Binding during Early Development
- Genome-Wide Transcript Profiling of Endosperm without Paternal Contribution Identifies Parent-of-Origin–Dependent Regulation of
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Meta-Analysis of Genome-Wide Association Studies in Celiac Disease and Rheumatoid Arthritis Identifies Fourteen Non-HLA Shared Loci
- MiRNA Control of Vegetative Phase Change in Trees
- Risk Alleles for Systemic Lupus Erythematosus in a Large Case-Control Collection and Associations with Clinical Subphenotypes
- The Cardiac Transcription Network Modulated by Gata4, Mef2a, Nkx2.5, Srf, Histone Modifications, and MicroRNAs
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MUDr. Irena Krčmová, CSc.
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



