-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
The Transcriptional Regulator Rok Binds A+T-Rich DNA and Is Involved in Repression of a Mobile Genetic Element in
The rok gene of Bacillus subtilis was identified as a negative regulator of competence development. It also controls expression of several genes not related to competence. We found that Rok binds to extended regions of the B. subtilis genome. These regions are characterized by a high A+T content and are known or believed to have been acquired by horizontal gene transfer. Some of the Rok binding regions are in known mobile genetic elements. A deletion of rok resulted in higher excision of one such element, ICEBs1, a conjugative transposon found integrated in the B. subtilis genome. When expressed in the Gram negative E. coli, Rok also associated with A+T-rich DNA and a conserved C-terminal region of Rok contributed to this association. Together with previous work, our findings indicate that Rok is a nucleoid associated protein that serves to help repress expression of A+T-rich genes, many of which appear to have been acquired by horizontal gene transfer. In these ways, Rok appears to be functionally analogous to H-NS, a nucleoid associated protein found in Gram negative bacteria and Lsr2 of high G+C Mycobacteria.
Published in the journal: . PLoS Genet 6(11): e32767. doi:10.1371/journal.pgen.1001207
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1001207Summary
The rok gene of Bacillus subtilis was identified as a negative regulator of competence development. It also controls expression of several genes not related to competence. We found that Rok binds to extended regions of the B. subtilis genome. These regions are characterized by a high A+T content and are known or believed to have been acquired by horizontal gene transfer. Some of the Rok binding regions are in known mobile genetic elements. A deletion of rok resulted in higher excision of one such element, ICEBs1, a conjugative transposon found integrated in the B. subtilis genome. When expressed in the Gram negative E. coli, Rok also associated with A+T-rich DNA and a conserved C-terminal region of Rok contributed to this association. Together with previous work, our findings indicate that Rok is a nucleoid associated protein that serves to help repress expression of A+T-rich genes, many of which appear to have been acquired by horizontal gene transfer. In these ways, Rok appears to be functionally analogous to H-NS, a nucleoid associated protein found in Gram negative bacteria and Lsr2 of high G+C Mycobacteria.
Introduction
In bacteria, horizontal gene transfer typically occurs through natural transformation, conjugation, and transduction and contributes to rapid evolution and the acquisition of new traits [1], [2]. For example, genes needed for pathogenesis, symbiosis, resistance to antibiotics, and metabolism of various compounds are often acquired through horizontal gene transfer. Even though horizontal gene transfer can confer potential benefits on a recipient cell, there are potentially lethal costs. For example, the acquisition and expression of genes for restriction enzymes (often carried on phage) is potentially lethal. The acquisition of transcriptional regulators could lead to inappropriate rerouting of gene regulatory networks, and the introduction and expression of enzymes could lead to altered metabolic flux, resulting in loss of fitness under some conditions. In addition, the introduction and expression of homologues of essential genes may be detrimental to the cell if the gene products interfere with essential cellular components.
Many organisms have mechanisms to inhibit acquisition and expression of foreign DNA. These mechanisms can help to mitigate some of the potentially deleterious effects of acquisition and expression of foreign genes. For example, the H-NS protein of Gram-negative bacteria is a widely studied nucleoid-associated protein that modulates gene expression and is a negative regulator of some genes and mobile elements acquired by horizontal gene transfer [3]–[5]. The presence of H-NS appears to be confined to proteobacteria. A functional analogue has been recently described from the high-G+C Gram positive actinomycete Mycobacterium tuberculosis [6], [7], but no analogous proteins have been described in low-G+C Gram positive organisms to date.
We found that the transcription factor Rok of the low-G+C (43.5%) Gram positive Bacillus subtilis binds to A+T-rich DNA and helps repress the activity of at least one mobile genetic element. Rok is relatively small (20.7 kD) and is found in several Bacillus species closely related to B. subtilis [8]. Rok was previously identified as a negative regulator of natural genetic competence in B. subtilis [9]. It binds to and represses the promoter of comK [9]. comK encodes a transcriptional activator that is required for expression of the B. subtilis competence machinery needed for uptake of exogenous native and foreign DNA [10], [11]. In addition to repressing transcription of comK, Rok represses transcription of several other genes, including many that encode extracellular functions [8]. Rok binds specifically to the promoters of at least a subset of these genes, although a consensus recognition sequence has not been identified [8].
To better understand the function of Rok in gene regulation and cell physiology, we characterized the association of Rok with chromosomal DNA in vivo. We expected to find Rok binding largely limited to chromosomal regions of genes known to be transcriptionally regulated by Rok [8]. In addition, we found extensive binding of Rok to genomic regions characterized by a high A+T, many of which are believed to have been acquired via horizontal gene transfer. We also found that Rok binds to and represses endogenous excision of the mobile genetic element ICEBs1, an integrative and conjugative element (conjugative transposon) found integrated in the 3′-end of a tRNA gene in B. subtilis [12], [13]. Our results indicate that Rok is a nucleoid-associated protein, is bound to chromosomal regions containing A+T-rich sequences, and inhibits the activity of at least one mobile genetic element. In these ways, Rok helps to repress several genes and elements known or thought to have been acquired by horizontal gene transfer.
Results
Rok is associated with parts of the nucleoid
Rok is a DNA binding protein and appears to have some sequence specificity, although no consensus binding sequence has been identified [8]. We used fluorescence microscopy to visualize the location of Rok in living cells (Figure 1A). We fused rok to yfp, (encoding yellow fluorescent protein, IYFP [14] such that rok-yfp was expressed from its native promoter and was the only source of Rok in the cell. Strains expressing the fusion had normal transformation frequencies, indicating that Rok-YFP was functional (data not shown). As expected for a DNA-binding protein, Rok-YFP (Figure 1A) appeared to be associated with the nucleoid, as visualized with 4′,6-diamidino-2-phenylindole (DAPI) (Figure 1B). However, Rok-YFP was not uniformly associated with the nucleoid (Figure 1).
Fig. 1. Localization of fluorescent Rok and HBsu fusion proteins. 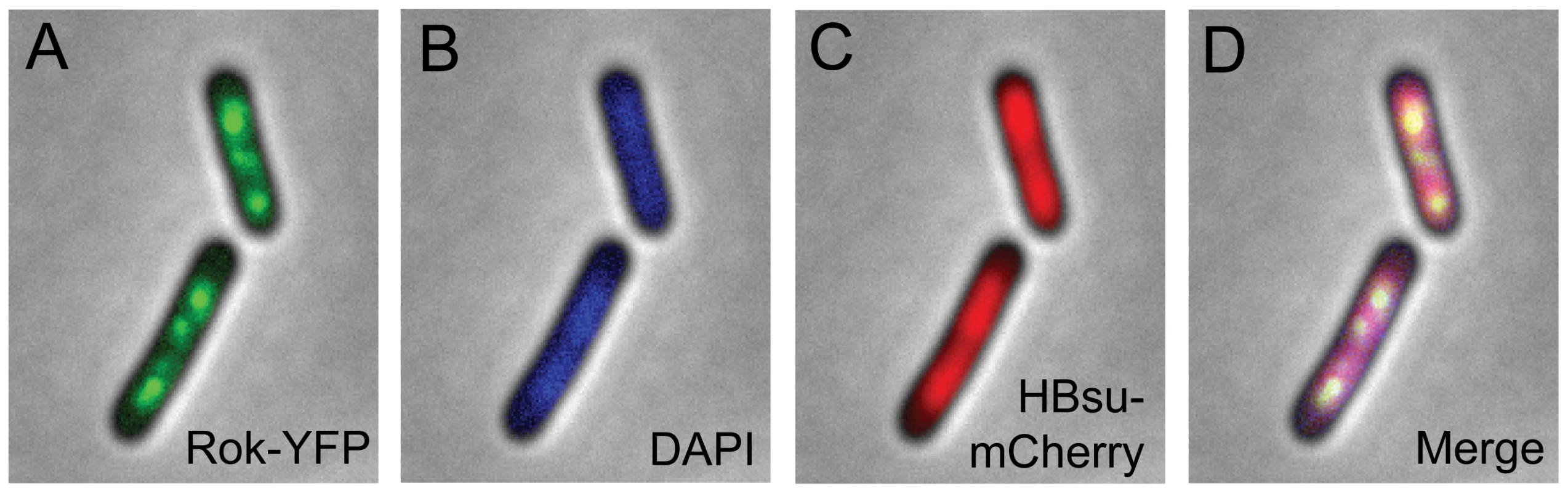
Strain WKS1102 (rok-iyfp; hbs-mcherry) was grown to mid-exponential phase at 32°C in defined minimal medium with glucose. Fluorescence microscopy images are overlaid with the phase contrast image. Images are representative of >95% of the cells. A. Rok-IYFP. B. DAPI. C. HBsu-mCherry. D. Merged image of A–C. In contrast, the nucleoid binding protein HBsu, the B. subtilis homologue of HU from Gram negative organisms [15], appeared to be associated with the nucleoid in a relatively uniform manner (Figure 1C, 1D). We fused HBsu (hbs) to mCherry [16] such that hbs-mcherry was expressed from the endogenous hbs promoter and the fusion was the only source of HBsu in the cell. hbs is essential for cell growth [17], and the hbs-mcherry fusion strain appeared to grow normally, indicating that the fusion was functional. Since HBsu-mCherry appeared to be associated with most or all of the nucleoid, and Rok-YFP was not uniformly associated with the nucleoid (Figure 1), we inferred that the DNA binding preferences of these two proteins are likely to be different. This was further explored in chromatin immunoprecipitation experiments with HBsu and Rok described below.
Rok associates with A+T-rich chromosomal DNA
We determined the genome wide binding profile of Rok (Figure 2A) and HBsu (Figure 2B) in vivo using formaldehyde-mediated crosslinking and immunoprecipitation (ChIP) and hybridization of immunoprecipitated DNA to DNA microarrays (ChIP-chip). We fused a cMyc epitope tag to the 3′ end of rok (rok-myc) and hbs (hbs-myc) such that each fusion is expressed from its native promoter and is the only copy of the gene in the cell. Like Rok-YFP and HBsu-mcherry, both myc-tagged proteins appeared to be functional; rok-myc strains had normal levels of competence and hbs-myc strains were viable (data not shown). We used monoclonal anti-myc antibodies to immunoprecipitate the myc-tagged proteins and detected DNA that was co-precipitated using DNA microarrays containing >99% of the annotated B. subtilis open reading frames as well as a subset of intergenic regions [18].
Fig. 2. Genome-wide binding profile of Rok and Hbs. 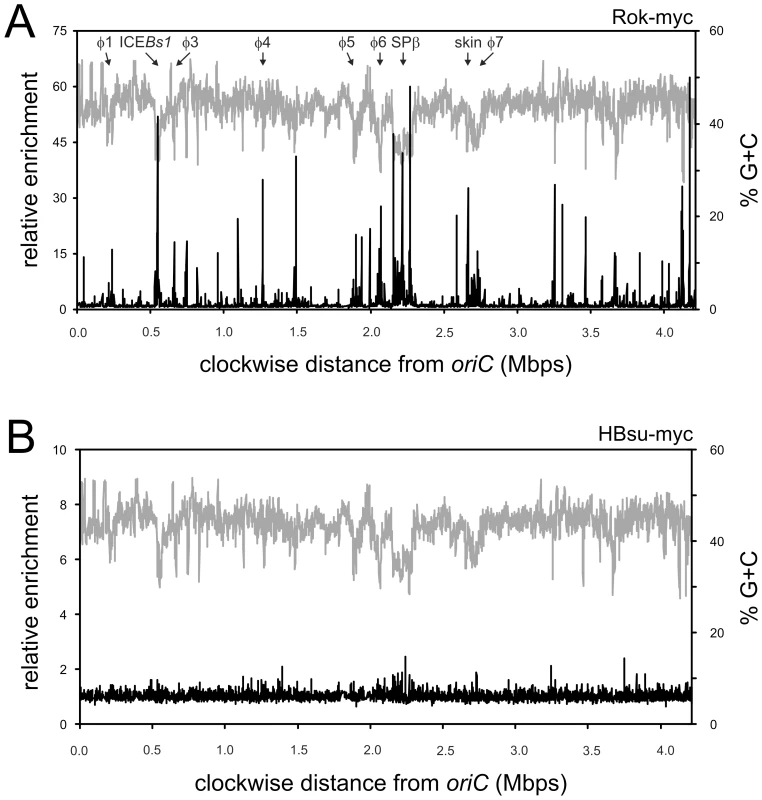
Strains WKS895 (rok-myc) and WKS935 (hbs-myc) were grown to mid-exponential (OD600∼0.6) phase at 32°C in defined minimal medium with glucose and processed as described in Materials and Methods. Relative enrichment of a particular genomic region is shown on the left axis, percent G+C on the right axis, and genome position on the x-axis. Enrichment values are averages from three independent cultures. A. Rok-myc (black curve) in relation to a sliding window analysis of the % G+C of the B. subtilis genome (gray curve). B. ChIP-chip of HBsu-myc (black curve) in relation to a sliding window analysis of the % G+C of the B. subtilis genome (gray curve). We found that Rok was associated with several genomic regions (Figure 2A), many of which correlated with the locations of genetic elements known or thought to have been acquired by horizontal transfer [19], [20]. Since horizontally acquired sequences often have a different G+C content than endogenous genes, we compared the Rok binding profile to the G+C content of the genome (Figure 2A). We calculated the average G+C content in windows of 3,000 bp with a step-size of 1,000 bp (Materials and Methods). The average % G+C across the entire genome is ∼43.5% [19]. Many of the chromosomal regions associated with Rok were strikingly lower in % G+C (higher % A+T) than the rest of the genome (Figure 2A).
The observed binding profile was specific for Rok, and not the myc-epitope tag or nucleoid binding proteins in general as the binding profile of HBsu-myc (Figure 2B) was quite different from that of Rok-myc (Figure 2A). In contrast to Rok, none of the chromosomal regions had >2.5-fold enrichment for HBsu-myc (Figure 2B), consistent with the function of HBsu as a general nucleoid binding protein [21]. Our results indicate that in vivo, HBsu is a relatively non-specific DNA binding protein and Rok binds preferentially to A+T-rich regions of the chromosome.
Analysis of the ChIP data for smaller chromosomal intervals highlights the difference between Rok and HBsu binding (Figure 3). Some of the chromosomal regions have genes characteristic of prophages, or defective prophages and thus appear to be or once have been functional mobile genetic elements likely acquired by horizontal gene transfer. These regions were designated as prophage regions and given a number [19]. Prophage regions 4, 5, and 6 have regions of G+C content significantly less than 40% (using a sliding window of 500 bp and a step size of 100 bp) and these regions are preferentially bound by Rok, but not HBsu (Figure 3A–3C). Similarly, the defective prophage skin [22]) and the prophage regions 1 and 7 also preferentially bind Rok, but not HBsu (Figure 2 and data not shown). Regions of the prophage SPβ [23] were bound by both Rok and HBsu, although binding of Rok appeared to be much greater (Figure 2). The sdp operon (Figure 3D), yybN operon (Figure 3E), and the yefC-yezA region (Figure 3F) are postulated to have been acquired by horizontal gene transfer [20]. Rok binds these regions in vivo, and binding correlates well with the regions of low G+C content (Figure 3D–3F).
Fig. 3. Detailed view of selected chromosomal regions. 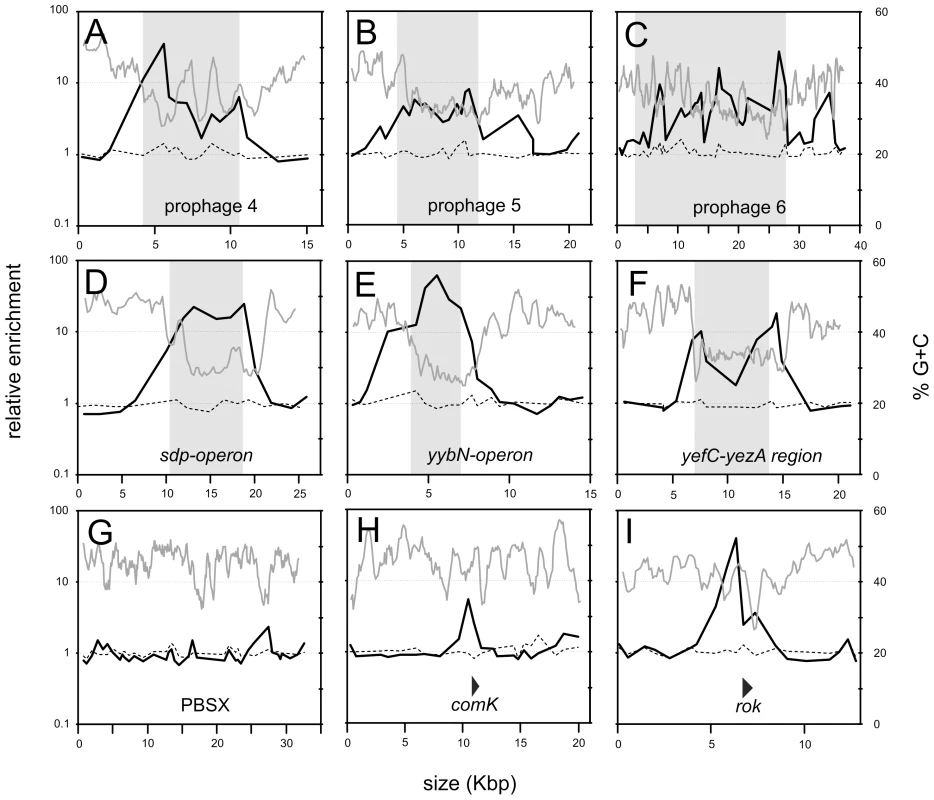
ChIP-chip data from Figure 2 are shown in more detail for selected chromosomal regions. The length of the genomic region is indicated on the x-axis. Enrichment in a Rok-myc immunoprecipitate (black curve) and HBsu-myc immunoprecipitate (dashed line) are given in relation to a sliding window analysis (window of 500 bp and a step size of 100 bp) of the % G+C of the B. subtilis genome (gray curve). A–F. Regions known or proposed to have been acquired by horizontal transfer [20] are indicated with a shaded box. A. Prophage 4 region (yjcJ-manA). B. Prophage 5 region (ynbA-yncB). C. Prophage 6 region (yoaW-yobS). D. sdp region (opuBD-opuCA). E. yybN region (yybP-yybA). F. yefC-yezA region (yeeA-yeeK). G-I. PBSX, a defective prophage, rok, thought to have been acquired by horizontal gene transfer, and comK, required for development of genetic competence. The locations and direction of transcription of comK and rok are indicated by triangles. G. PBSX region (yjqA-ykaA). H. comK region (yhfO-yhjJ). I. rok region (ykuN-yknU). Rok does not bind to all horizontally acquired DNA. The defective prophage PBSX [24] is a mobile genetic element likely acquired by horizontal gene transfer and Rok was not significantly bound to this chromosomal region (Figure 3G). However, the nucleotide composition of the PBSX region is not significantly different from the average of the B. subtilis genome (Figure 3G), consistent with the findings that Rok binds preferentially to regions of relatively low G+C content. Similarly, H-NS in Salmonella species does not significantly associate with mobile genetic elements that have G+C content similar to that of the host [25], [26].
Rok association with previously defined targets
Previously, Rok was found to control expression of at least 20 different transcription units, either directly or indirectly [8], [9]. At least nine of these appear to be directly regulated by Rok since Rok binds to the promoter regions in vitro [8]. The ChIP-chip data indicate that Rok is also bound to (or near) almost all of these in vivo (Table 1), consistent with a role for Rok in directly repressing their expression.
Tab. 1. Association of Rok in vivo with specific transcription units. 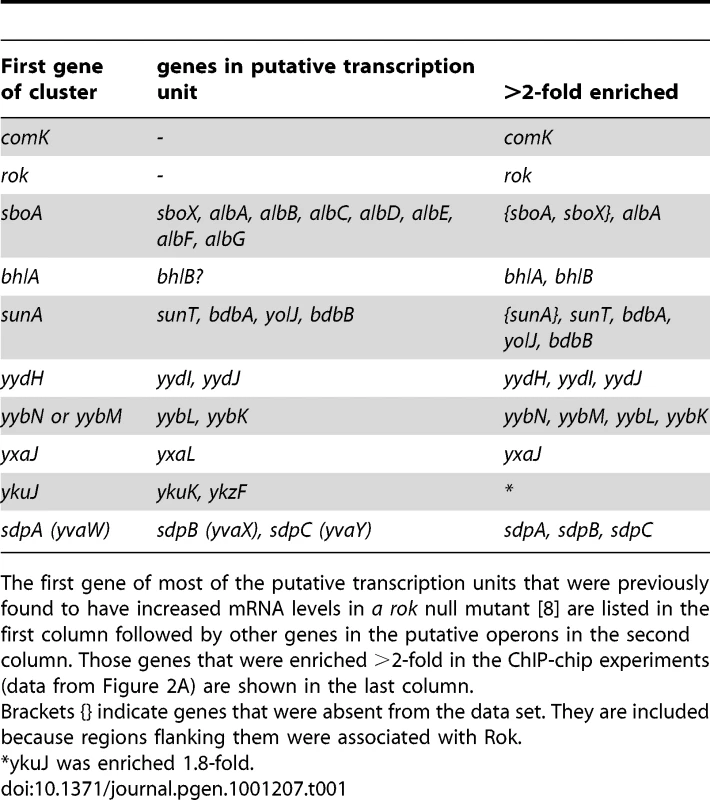
The first gene of most of the putative transcription units that were previously found to have increased mRNA levels in a rok null mutant [8] are listed in the first column followed by other genes in the putative operons in the second column. Those genes that were enriched >2-fold in the ChIP-chip experiments (data from Figure 2A) are shown in the last column. Several of the previously identified Rok targets are located in genomic regions that have extensive binding of Rok in vivo. For instance, prophage 4 contains yjcN, a gene previously identified as a direct target of Rok [8]. Rok binding extends well beyond a simple regulatory region for yjcN (Figure 3A). Similarly, the sdp operon and yybN genomic region were previously identified as direct targets of Rok [8]. Rok binding to these regions in vivo extends greater than 5 kb (Figure 3D and 3E).
Rok also appears to bind to some regions that do not have significantly lower G+C content than the average for the B. subtilis genome. For example, the comK region has normal G+C content and Rok binds to this region in vivo (Figure 3H) and in vitro [9]. This also seemed to be the case for Rok binding to its own promoter region, although there is also some Rok binding in an A+T-rich region of rok (Figure 3I). These results might reflect recruitment of Rok by other DNA binding proteins. Alternatively, they might indicate that Rok has some DNA sequence-specific binding beyond the recognition of low G+C content, and that binding specificity could be obscured by the less specific and widespread association with low G+C content DNA.
Rok associates with A+T-rich DNA in a heterologous host
We found that when expressed in Escherichia coli, Rok also bound preferentially to A+T-rich DNA. We introduced a plasmid expressing full length Rok with a C-terminal myc-tag (Rok-myc) into E. coli and confirmed by Western blotting that the protein accumulated (data not shown). We used ChIP followed by quantitative real time PCR (ChIP-qPCR) to compare the association of Rok-myc with six different chromosomal regions in E. coli, three that are A+T-rich and three that are G+C-rich (Table 2). There was significantly greater association of Rok-myc with the A+T-rich sequences than the G+C-rich sequences (Table 2; Figure 4A). This preferential binding of Rok-myc to A+T-rich sequences in E. coli is consistent with the binding of Rok to AT-rich sequences in B. subtilis as determined by ChIP-chip (Figure 2).
Fig. 4. Preferential binding of Rok to A+T-rich DNA in E. coli. 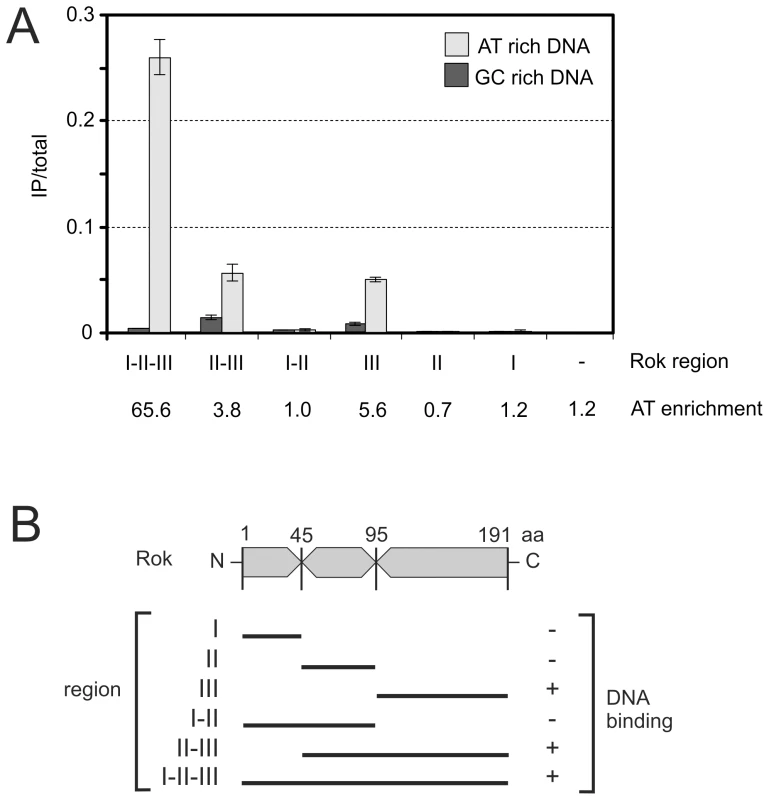
E. coli cells containing the indicated plasmids were grown to mid-exponential phase at 32°C in LB medium and processed for ChIP-PCR as described in Materials and Methods. A. Normalized levels of A+T rich (ybcL; 43.3% G+C; light gray) or G+C rich (phnO; 62.9%G+C; dark gray) DNA in the immunoprecipitates of the myc-tagged region I (pWKS1046), region II (pWKS1048), region III (pWKS1049), region I+II (pWKS1051), region II+III (pWKS1052) and full length Rok – region I+II+III (pWKS1054; same data as in Table 2). The relative enrichment of ybcL (A+T-rich) vs phnO (G+C-rich) is given below the graph. Error bars indicate standard error of the mean (n = 3). B. Schematic representation of the regions of Rok as defined for these experiments. DNA binding is indicated on the right. Tab. 2. Rok preferentially binds A+T-rich DNA in E. coli. 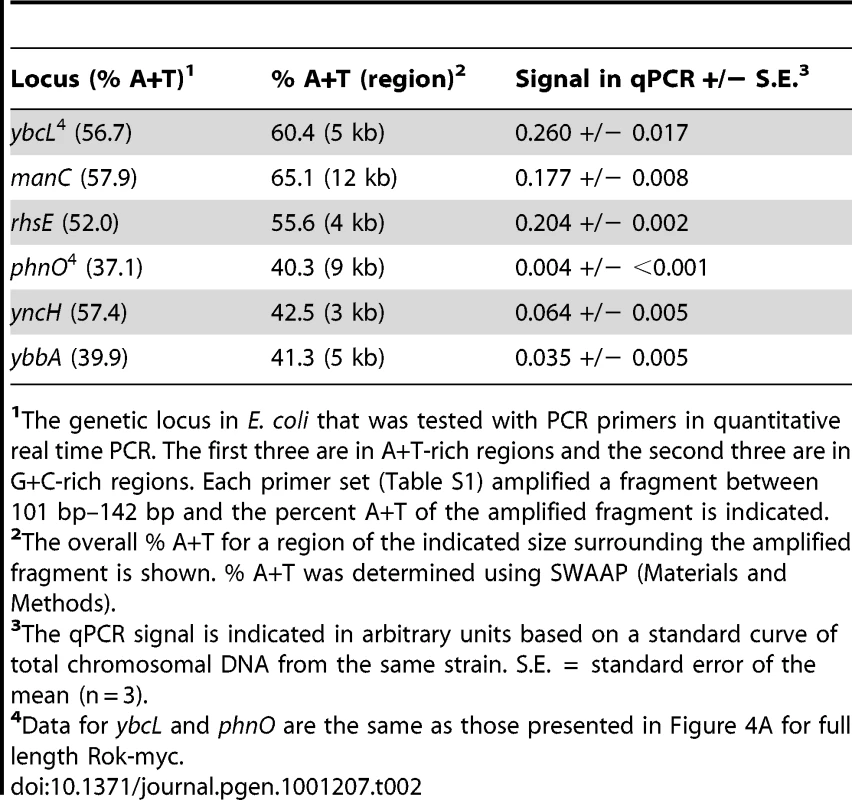
The genetic locus in E. coli that was tested with PCR primers in quantitative real time PCR. The first three are in A+T-rich regions and the second three are in G+C-rich regions. Each primer set (Table S1) amplified a fragment between 101 bp–142 bp and the percent A+T of the amplified fragment is indicated. The C-terminal domain of Rok is sufficient for DNA binding
It is not known which region of Rok is needed for DNA binding [8], [9], [27]. Using various tools for sequence analyses combined with rok deletions and ChIP-qPCR, we found that the DNA binding activity of Rok is contained in the C-terminal region of the protein. We used a ClustalW2 alignment {http://www.ebi.ac.uk/clustalw/} of primary amino acid sequences of Rok homologs from B. subtilis, B. amyloliquefaciens, B. pumilus, B. licheniformis, B. coagulans and B. pseudomycoides to define conserved regions of the protein (Figure S1). The analysis revealed three distinct regions: (I) a highly conserved N-terminal region; (II) a moderately conserved central region; and (III) a highly conserved C-terminal region (Figure 4B and Figure S1). On the basis of the presence of many positively charged residues, we used the third putative domain to query the I-TASSER server for protein structure prediction [28]. The in silico analysis predicted that the C-terminal region might have some structural relatedness (Tm-score 0.6145) to winged helix DNA-binding domains of proteins such as FurB from Mycobacterium tuberculosis (PDB 2o03_A) {Lucarelli 2007, FurB}. The C-terminal region of Rok was also classified as containing a possible DNA/RNA-binding 3-helical bundle from the Winged Helix Superfamily, with an estimated precision of 20% by the PHYRE Protein Fold Recognition server [29].
Based on the in silico sequence analysis, we tested for the ability of the three regions of Rok to bind DNA. We fused each of these regions to a myc-epitope tag (Figure 4B) and used ChIP-qPCR to measure the ability of each fusion to associate with a region of E. coli DNA with high or low A+T content (as described above). Each of the fusion proteins accumulated to similar levels in E. coli and was detectable with anti-myc antibodies (data not shown). We found that any variant that contained the C-terminal region (region III) of Rok was able to bind DNA, and there was a preference for DNA with high A+T content (Figure 4A). In contrast, variants that contained region I, region II, or both, did not appear to have significant DNA binding activity (Figure 4A, 4B). However, association of full length Rok with A+T-rich DNA was greater than that of region III alone (Figure 4A). Based on these results, we conclude that region III of Rok is the DNA binding region and that this region alone has some preference for A+T-rich DNA. We suspect that region I, either alone or in combination with region II, likely contributes to DNA binding and the specificity for A+T-rich DNA, perhaps by affecting dimerization and/or multimerization of Rok and potentially contributing to cooperativity. This would be similar to H-NS and H-NS-like proteins where the N-terminal region affects multimerization [30].
Rok contributes to stability of ICEBs1
Rok binds to regions of the mobile element ICEBs1, and the regions with the most binding, at the left and right ends of ICEBs1, have the lowest G+C content (Figure 5A). The genes encoding the ICEBs1 site-specific recombinase (int) and the excisionase (xis) that allow excision from the genome are in the left end, and the recombination reaction occurs between 17 bp sequences found at the ends of the integrated element (attL and attR) [12]. During normal exponential growth, ICEBs1 gene expression is repressed by the element's repressor ImmR and there is very little excision of ICEBs1 [31]. Upon production of active RapI, a cell sensory protein, or during the RecA-dependent SOS response, ImmR is inactivated and ICEBs1 gene expression is derepressed. This leads to rapid production of excisionase and efficient excision of ICEBs1 from the chromosome [31], [32]. Because Rok was associated with genes and sequences at the ends of ICEBs1, we determined the effects of a rok null mutation on the stability of ICEBs1.
Fig. 5. Association of Rok with ICEBs1. 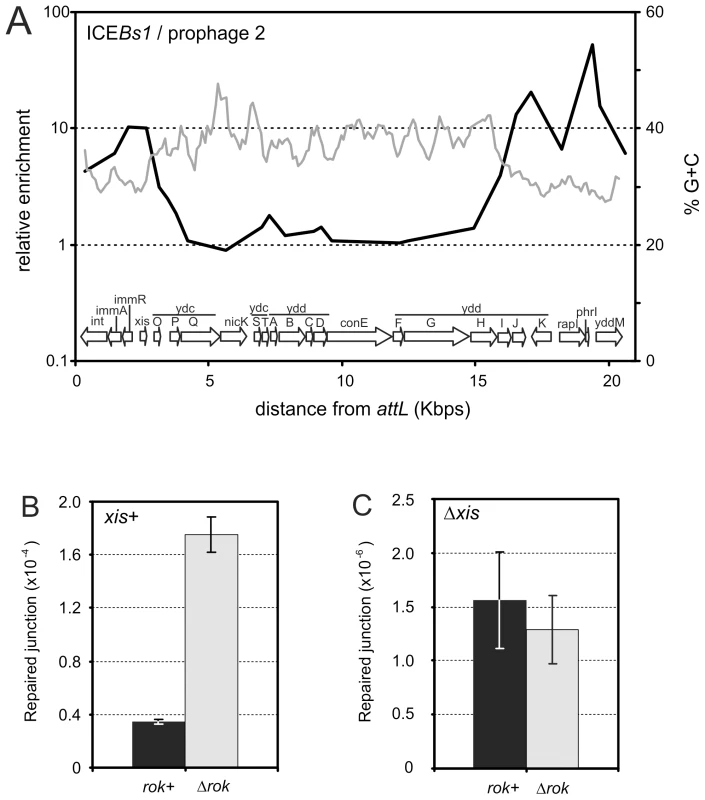
A. Association of Rok-myc with ICEBs1 in relation to a sliding window analysis of its %G+C. The genomic organization of ICEBs1 is given in the graph. Data are from Figure 2. B. Excision of ICEBs1 in rok+ (MMB869) and rok− (WKS1112) strains. C. Excision of an ICEBs1 xis null mutant in rok+ (CAL873) and rok− (WKS1093) strains. Strains used in panels B and C contained an (uninduced) amyE::Pxyl-rapI construct. Induction of this construct eliminated the effect of a rok mutation (data not shown). For panels B and C, error bars indicate the standard error of the mean (n = 3). We found that endogenous excision of ICEBs1 increased approximately 4-fold in a rok null mutant (Figure 5B). Excision of ICEBs1 by site-specific recombination between the left and right attachment sites (attL and attR) leaves behind an empty bacterial attachment site, attB, that is readily detected by PCR using appropriate primers [12]. The occurrence of attB in a population, compared to a nearby chromosomal locus ydbT, is a measure of the frequency of excision of ICEBs1 [33], [34]. During exponential growth, the excision frequency of ICEBs1 in rok+ cells was ∼4×10−5 (Figure 4B), consistent with previous measurements [33], [34]. In a rok null mutant, this frequency increased to ∼1.6×10−4 (Figure 5B).
The increased excision frequency of ICEBs1 in the rok null mutant could be due to an increase in integrase - and excisionase-mediated site-specific recombination, or possibly homologous recombination between the 60 bp direct repeats that mark the ends of ICEBs1. We found that the increased excision in the rok null mutant was dependent on the ICEBs1 excisionase. An ICEBs1 xis (excisionase) null mutation reduced the excision frequency to ∼1-2×10−6 (Figure 5C), consistent with previous findings [33]. This frequency did not significantly change in a rok null mutant (Figure 5C), indicating that the increased spontaneous excision observed in the rok mutant requires excisionase and is not due to increased homologous recombination between the 60 bp direct repeats at the ends of ICEBs1. Based on these results, we conclude that Rok contributes to keeping ICEBs1 quiescent in the genome.
Discussion
Results presented here show that B. subtilis Rok binds A+T-rich DNA, that the DNA binding activity resides in its C-terminal region, and that Rok helps inhibit the mobile genetic element ICEBs1. Together with previous work on the effects of rok on gene expression [8], these properties indicate that Rok is analogous to the nucleoid-associated protein H-NS from E. coli and other Gram negative organisms [3]–[5], and the recently described Lsr2 of Mycobacterium and related high-G+C actinomycetes [6].
Targets of Rok and horizontal gene transfer
Rok is a negative regulator of genes involved in horizontal gene transfer in at least two ways: 1) as a repressor of competence development, and 2) as an inhibitor of expression of genes in A+T-rich chromosomal regions. rok was initially identified as a repressor of of comK [9]. The comK gene product is a transcriptional activator and the master regulator of competence development and the K-state in B. subtilis [10], [11], [35]. By virtue of repressing comK, Rok also indirectly represses expression of the genes activated by ComK and is a therefore a strong negative regulator of competence development, thus inhibiting the ability of cells to acquire foreign DNA.
Rok also binds to chromosomal regions that are known or thought to have been acquired by horizontal gene transfer and are characterized by a high A+T content. One of these regions contains ICEBs1, an integrative and conjugative element integrated in the B. subtlis chromosome [12]. Rok binds to ICEBs1 and helps prevent spontaneous excision. Many other horizontally acquired genes are bound by Rok in vivo and expression of some of these were previously found to be de-repressed in a rok null mutant [8]. Interestingly, rok itself appears to have been recently acquired in the B. subtilis - B. amyloliquefaciens - B. licheniformis clade as it is inserted in and interrupts an otherwise conserved genomic arrangement [8]. rok is auto-regulated, repressing its own expression. That is, transcription of rok increases in the absence of functional Rok protein. Thus, when the concentration of Rok decreases at its own promoter, its expression will increase, perhaps allowing cells to adjust the levels of Rok in response to acquisition of new DNA that has a high A+T content.
Effects of Rok on gene expression
A null mutation in rok causes increased expression of at least 20 transcription units, either directly or indirectly [8], separate from genes that are activated by ComK. In vitro, Rok binds to sequences upstream of at least a subset of these genes [8]. We found that in vivo Rok is associated with most of these genes, consistent with the notion that Rok directly represses their transcription.
Our results indicate that the number of chromosomal genes bound by Rok is s significantly greater than the number of genes whose expression is detectably altered in a rok null mutant [8]. There are two ways to explain this difference. First, many genes bound by Rok are not expected to be expressed under the conditions used for analysis of mRNA levels [8]. This is particularly true for genes that might be controlled by other regulators, and for regions containing the mobile genetic elements that are strongly repressed during normal growth. Second, Rok may be bound to regions but not properly positioned to have an effect on transcription.
Multiple ways for Rok to bind DNA?
There is a general correlation between chromosomal regions with low G+C content and Rok binding. However, it is notable that at least a few of the chromosomal regions bound by Rok appear to have a G+C content much closer to or greater than the norm, including the comK and rok regulatory regions. Rok might be recruited to these regions by other DNA binding proteins. Alternatively, Rok might be capable of some sequence-specific binding somewhat different from binding to A+T-rich DNA, and these possibilities are not mutually exclusive. A discriminative MEME motif search [36] using the few regions of Rok binding that are close to the average G+C content identified a motif that appears to be overrepresented (Figure S2). This motif could be a binding site for another protein that possibly interacts with Rok, or could indicate a specific binding sequence for Rok. Further genetic and functional dissection of Rok and this motif should help determine how Rok is associated with this DNA. Binding of H-NS to DNA also appears to be complex. H-NS can interact with or bind to regions bound by other DNA binding proteins, might have some site-specific binding, and can switch from stimulating DNA bridging to causing DNA stiffening {e.g., [5], [37]–[40]}.
Rok and the nucleoid-associated proteins H-NS and Lsr2
All bacteria appear to have nucleoid-associated proteins (NAPs) [41] that are abundant, bind relatively non-specifically to extended regions of the chromosome, and often cause changes in DNA topology. The most highly conserved nucleoid-associated protein is the “heat-unstable” protein HU (and the related integration host factor IHF) found in both Gram negative and Gram positive organisms. In contrast, the heat-stable nucleoid restructuring protein H-NS is found in several Gram negative bacteria, but there are no obvious homologues in Gram positive bacteria (reviewed in [42]).
Despite the lack of sequence similarity, the recently characterized Lsr2 protein from Mycobacerium and related high G+C Gram positive bacteria is a functional analogue of H-NS [6]. Like H-NS, Lsr2 binds A+T-rich DNA, including regions acquired by horizontal gene transfer [7], [43], can repress transcription [43], is capable of bridging DNA [6], [44], and can partly substitute for H-NS in E. coli [6]. Neither H-NS nor Lsr2 homologues have been found in low G+C content Gram positive species.
There are several functional similarities between Rok of B. subtilis and H-NS (and Lsr2). Both Rok and H-NS act as negative regulators of transcription and bind extended chromosomal regions with high A+T content. H-NS causes significant changes in DNA topology, and this has also been postulated for Rok [27]. Most notably, both Rok and H-NS help silence foreign DNA with a high A+T content. Transcriptional repression exerted by H-NS can be reversed by certain anti-silencing mechanisms {reviewed in [45]}. Likewise, auto-activation of comK transcription is accomplished by preventing Rok-mediated repression [27]. Rok and H-NS are both relatively small (20.7 kDa and 15.4 kDa, respectively), although H-NS appears to be more abundant than Rok. There are approximately 20,000 molecules per cell of H-NS in growing cells [4]. In contrast, we estimate that there are approximately 1,000–3,000 molecules of Rok per genome of exponentially growing B. subtilis cells (see Materials and Methods). Based on the several similarities in function, we propose that Rok of B. subtilis and its close relatives is functionally analogous to H-NS of Gram negative bacteria and Lsr2 of Mycobacterium and related high G+C actinomycetes. We suspect that other organisms have H-NS analogues that are not readily recognized by sequence similarities.
Materials and Methods
Media and growth conditions
For routine growth and manipulations, E. coli and B. subtilis cells were grown in LB medium. For most experiments, B. subtilis cells were grown in the MOPS buffered S750 defined minimal medium [46] with 0.1% glutamate, supplemented with required amino acids (typically tryptophan and phenylalanine), 1% glucose or arabinose as a carbon source, and 1 mM IPTG or 1% xylose as inducer as necessary. Strains with plasmids integrated into the chromosome by single crossover were grown with appropriate antibiotic to maintain selection for the integrated plasmid.
Strains and alleles
B. subtilis strains used are listed in Table 3. PCR Primers used in strain constructions are listed in Table S1. B. subtilis strains were constructed by transformation using standard procedures [35], [47] Previously described alleles affecting ICEBs1 include Δxis190, a deletion of the excisionase gene of ICEBs1 [33], and Pxyl-rapI [48], used to induce efficient ICEBs1 gene expression.
Tab. 3. <i>B. subtilis</i> strains. 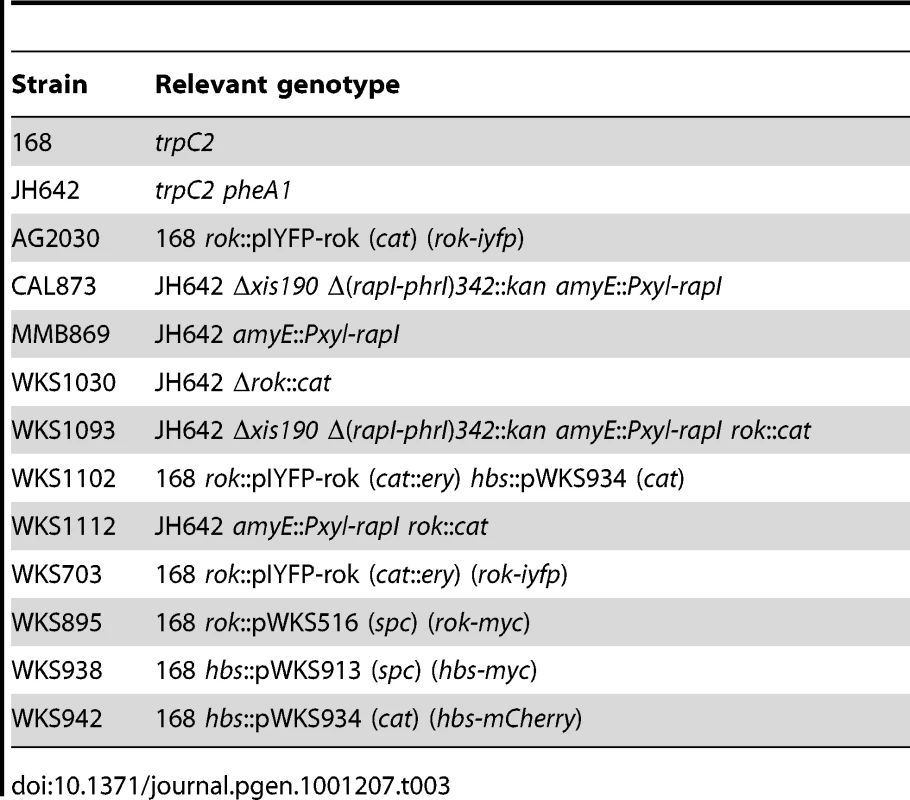
rok-iyfp
The 3′ end of the rok open reading frame was amplified using primers rok-gfp-KpnI and rok-gfp -EcoRI. After digestion with KpnI and EcoRI, this fragment was ligated into similarly digested pIYFP [14], yielding pIYFP-Rok. To allow for dual labeling, the chloramphenicol resistance gene of AG2030 (rok-yfp) was replaced with a erythromycin resistance gene using plasmid pCm::Em [49].
hbs-mCherry
The 3′ end of the hbs open reading frame was amplified using primers oWKS-213 and oWKS-215. After digestion with EcoRI and XhoI, the fragments were ligated into similarly digested pWKS553, yielding pWKS934. pWKS553 is derived from pGEMcat [50] and contains mCherry inserted between the KpnI and SphI sites. mCherry was amplified with primers oWKS-114 and oWKS-115 from pMMB1010 (from M. Berkman, Suffolk University, Boston).
rok-myc and hbs-myc
The 3′ end of rok and hbs were amplified using primers oWKS-121 or oWKS-213 and oWKS-215, respectively. After digestion with EcoRI and XhoI, the fragments were ligated into similarly digested pCAL812 (a vector for making C-terminal fusions to 3x-myc and encoding spectinomycin-resistance for selection of single cross-over integrants in B. subtilis), yielding pWKS516 and pWKS913, respectively. These plasmids were introduced by single crossover into the B. subtilis chromosome by natural transformation selecting for spectinomycin-resistance.
Δrok::cat
We used long flanking homology PCR [51] to construct a deletion-insertion that replaces the rok open reading frame with a chloramphenicol resistance cassette. Fragments ∼1 kb upstream and downstream of rok were amplified using primers oWKS-245, oWKS-246, and oWKS-247 and oWKS-248, respectively, resulting in two products with regions of complementarity to a chloramphenicol resistance cassette (cat). These products were used as megaprimers in an expand Long Template PCR reaction (Roche) with plasmid pGEMcat (Youngman, 1989) as template, followed by a second Expand Long Template PCR using primers oWKS-245 and oWKS-248. Purified PCR product of the expected size was transformed into a wild type strain, and double crossover replacement of the rok gene with the cat cassette was verified by PCR.
rok-myc alleles in E. coli
E. coli strains expressing C-terminally myc-tagged variants of Rok were constructed as follows. Region I (N-terminal) was amplified using primers oWKS-235 and oWKS-239, region II (central) using primers oWKS-236 and oWKS-240, region III (C-terminal) using primers oWKS-237 and 238, region I-II using primers oWKS-235 and oWKS-240, region II-III using primers oWKS-236 and oWKS-238, and full length rok (region I-II-III) was amplified using primers oWKS-235 and oWKS-238. The resulting PCR products (that contain an artificial ribosome binding site and a sequence encoding a C-terminal cMyc epitope tag) were digested with SalI and SphI and cloned into similarly digested pDR110, yielding pWKS1046 (region I), pWKS1048 (region II), pWKS1049 (region III), pWKS1051 (region I+II), pWKS1052 (region II+III) and pWKS1054 (full length). After initial transformation into an intermediary E. coli host, the plasmids were recovered and introduced into E. coli BL21(DE3) [52] for the ChIP experiments.
Fluorescence microscopy
Cells were grown in defined minimal medium, placed on agarose (1.5%) pads containing Spizizen minimal salts [47]. DAPI was added to a final concentration of ∼80 ng/ml five minutes prior to visualization. Images were acquired using Nikon Ti-E inverted microscope under a 100× phase oil objective. Fluorescence images were acquired using Nikon Intensilight mercury illuminator and appropriate sets of excitation and emission filters (49008 for mCherry, 49003 for YFP and 49000 for DAPI, Chroma). Images were recorded using a CoolSNAP HQ camera (Photometrics) and processed using NIS-Elements Advanced Research 3.10 Software. TIFF images were processed in Adobe Photoshop CS3 and Figure 1 was prepared in Adobe Illustrator CS3.
Chromatin immunoprecipitation (ChIP) experiments
Chromatin immunoprecipitation of DNA bound to the various proteins in B. subtilis was done essentially as described [53], except that DNA was precipitated in the presence of glycogen (20 µg) as a carrier. For ChIP experiments in E. coli, crosslinking was done at room temperature for 20 minutes. Myc-tagged proteins were immunoprecipitated using monoclonal anti-cMyc antibodies (Zymed). Both Rok-myc and HBsu-myc were functional in B. subtilis (see results) and both were detected in Western blots (data not shown). Preliminary comparisons between Rok-myc and HBsu-myc in Western blots with anti-myc monoclonal antibodies indicated that there is about 20-50-fold more HBsu in the cell than Rok. Since the cellular concentration of HBsu is about 50,000 molecules per genome [54], we estimate that there are approximately 1,000–3,000 molecules of Rok per genome. For Rok-myc and HBsu-myc, we verified that the anti-myc antibodies actually immunoprecipitated the protein by depleting it from an extract (data not shown). Even though there was little or no significant enrichment of specific chromosomal regions in the HBsu-myc ChIP experiments, the protein appeared to crosslink to DNA as the signals on the microarrays were significantly above background.
ChIP-chip analyses were performed as described previously using printed DNA microarrays with PCR products corresponding to most open reading frames and many intergenic regions [55]. The microarray data discussed in this publication have been deposited in NCBI's Gene Expression Omnibus [56] and are accessible through GEO Series accession number GSE23199 (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE23199).
qPCR was performed on a Roche LightCycler 480 II. Samples (2 µl) of immunoprecipitated DNA were analyzed in duplicate in a 20 µl reaction volume that contained Sybr green. Signals were analyzed using the LightCycler 480 SW 1.5 software (Roche), according to the manufacturer (Absolute quantification; 2nd derivative of Max). Signals were normalized against a 12-point standard curve obtained from a dilution series of total chromosomal DNA of B. subtilis JMA222 [34] or BL21(DE3) [52]. Primers used for real time PCRs are listed in Table S1.
For all conditions at least three independent biological replicates were analyzed and data shown represent the average of these replicates.
ICEBs1 excision assay
Excision of ICEBs1 was monitored by quantitative real time PCR, as described previously [34] using primers CLO261, CLO262, CLO283, and CLO284 (Table S1).
In silico analyses
Conserved regions of Rok from B. subtilis 168 (NP_389307.1), B. amyloliquefaciens FZB42 (YP_001420994.1), B. pumilus ATCC7061 (ZP_03052836.1) and SAFR-032 (YP_001486564.1), B. licheniformis ATCC14580 (YP_078814.1 and YP_079175.1), B. psuedomycoides DSM12442 (ZP_04153718.1) and B. coagulans 36D1 (ZP_04433348.1) were identified using ClustalW2 (http://www.ebi.ac.uk/Tools/clustalw2/index.html). Amino acids 96–191 of B. subtilis Rok were used to query the I-Tasser server (http://zhanglab.ccmb.med.umich.edu/I-TASSER) [57] and the PHYRE Protein Fold Recognition Server (http://www.sbg.bio.ic.ac.uk/~phyre/) [29] for secondary structure predictions. Sliding window analyses were performed using SWAAP 1.0.3 (http://asiago.stanford.edu/SWAAP/SwaapPage.htm) [58] on genome sequences of Bacillus subtilis (accession number AL009126) and E. coli BL21(DE3) (accession number CP001665) retrieved from GenBank (ftp://ftp.ncbi.nih.gov/genbank/genomes/Bacteria/). A discriminative MEME motif search [36] was used to detect overrepresented motifs in selected regions that show Rok binding in vivo.
Supporting Information
Zdroje
1. FrostLS
LeplaeR
SummersAO
ToussaintA
2005 Mobile genetic elements: the agents of open source evolution. Nat Rev Microbiol 3 722 732
2. WozniakRA
WaldorMK
2010 Integrative and conjugative elements: mosaic mobile genetic elements enabling dynamic lateral gene flow. Nat Rev Microbiol 8 552 563
3. NavarreWW
McClellandM
LibbySJ
FangFC
2007 Silencing of xenogeneic DNA by H-NS-facilitation of lateral gene transfer in bacteria by a defense system that recognizes foreign DNA. Genes Dev 21 1456 1471
4. DormanCJ
2007 H-NS, the genome sentinel. Nat Rev Microbiol 5 157 161
5. FangFC
RimskyS
2008 New insights into transcriptional regulation by H-NS. Curr Opin Microbiol 11 113 120
6. GordonBR
ImperialR
WangL
NavarreWW
LiuJ
2008 Lsr2 of Mycobacterium represents a novel class of H-NS-like proteins. J Bacteriol 190 7052 7059
7. GordonBR
LiY
WangL
SintsovaA
van BakelH
2010 Lsr2 is a nucleoid-associated protein that targets AT-rich sequences and virulence genes in Mycobacterium tuberculosis. Proc Natl Acad Sci U S A 107 5154 5159
8. AlbanoM
SmitsWK
HoLT
KraigherB
Mandic-MulecI
2005 The Rok protein of Bacillus subtilis represses genes for cell surface and extracellular functions. J Bacteriol 187 2010 2019
9. HoaTT
TortosaP
AlbanoM
DubnauD
2002 Rok (YkuW) regulates genetic competence in Bacillus subtilis by directly repressing comK. Mol Microbiol 43 15 26
10. BerkaRM
HahnJ
AlbanoM
DraskovicI
PersuhM
2002 Microarray analysis of the Bacillus subtilis K-state: genome-wide expression changes dependent on ComK. Mol Microbiol 43 1331 1345
11. OguraM
YamaguchiH
KobayashiK
OgasawaraN
FujitaY
2002 Whole-genome analysis of genes regulated by the Bacillus subtilis competence transcription factor ComK. J Bacteriol 184 2344 2351
12. AuchtungJM
LeeCA
MonsonRE
LehmanAP
GrossmanAD
2005 Regulation of a Bacillus subtilis mobile genetic element by intercellular signaling and the global DNA damage response. Proc Natl Acad Sci U S A 102 12554 12559
13. BurrusV
PavlovicG
DecarisB
GuedonG
2002 The ICESt1 element of Streptococcus thermophilus belongs to a large family of integrative and conjugative elements that exchange modules and change their specificity of integration. Plasmid 48 77 97
14. VeeningJW
SmitsWK
HamoenLW
JongbloedJD
KuipersOP
2004 Visualization of differential gene expression by improved cyan fluorescent protein and yellow fluorescent protein production in Bacillus subtilis. Appl Environ Microbiol 70 6809 6815
15. MickaB
GrochN
HeinemannU
MarahielMA
1991 Molecular cloning, nucleotide sequence, and characterization of the Bacillus subtilis gene encoding the DNA-binding protein HBsu. J Bacteriol 173 3191 3198
16. ShanerNC
CampbellRE
SteinbachPA
GiepmansBN
PalmerAE
2004 Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat Biotechnol 22 1567 1572
17. MickaB
MarahielMA
1992 The DNA-binding protein HBsu is essential for normal growth and development in Bacillus subtilis. Biochimie 74 641 650
18. BreierAM
GrossmanAD
2007 Whole-genome analysis of the chromosome partitioning and sporulation protein Spo0J (ParB) reveals spreading and origin-distal sites on the Bacillus subtilis chromosome. Mol Microbiol 64 703 718
19. KunstF
OgasawaraN
MoszerI
AlbertiniAM
AlloniG
1997 The complete genome sequence of the gram-positive bacterium Bacillus subtilis. Nature 390 249 256
20. NicolasP
BizeL
MuriF
HoebekeM
RodolpheF
2002 Mining Bacillus subtilis chromosome heterogeneities using hidden Markov models. Nucleic Acids Res 30 1418 1426
21. KohlerP
MarahielMA
1997 Association of the histone-like protein HBsu with the nucleoid of Bacillus subtilis. J Bacteriol 179 2060 2064
22. TakemaruK
MizunoM
SatoT
TakeuchiM
KobayashiY
1995 Complete nucleotide sequence of a skin element excised by DNA rearrangement during sporulation in Bacillus subtilis. Microbiology 141 323 327
23. LazarevicV
DusterhoftA
SoldoB
HilbertH
MauelC
1999 Nucleotide sequence of the Bacillus subtilis temperate bacteriophage SPbetac2. Microbiology 145 Pt 5 1055 1067
24. KroghS
O'ReillyM
NolanN
DevineKM
1996 The phage-like element PBSX and part of the skin element, which are resident at different locations on the Bacillus subtilis chromosome, are highly homologous. Microbiology 142 Pt 8 2031 2040
25. LucchiniS
RowleyG
GoldbergMD
HurdD
HarrisonM
2006 H-NS mediates the silencing of laterally acquired genes in bacteria. PLoS Pathog 2 e81 doi:10.1371/journal.ppat.0020081
26. NavarreWW
PorwollikS
WangY
McClellandM
RosenH
2006 Selective silencing of foreign DNA with low GC content by the H-NS protein in Salmonella. Science 313 236 238
27. SmitsWK
HoaTT
HamoenLW
KuipersOP
DubnauD
2007 Antirepression as a second mechanism of transcriptional activation by a minor groove binding protein. Mol Microbiol 64 368 381
28. ZhangY
2008 I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9 40
29. KelleyLA
SternbergMJ
2009 Protein structure prediction on the Web: a case study using the Phyre server. Nat Protoc 4 363 371
30. DormanCJ
HintonJC
FreeA
1999 Domain organization and oligomerization among H-NS-like nucleoid-associated proteins in bacteria. Trends Microbiol 7 124 128
31. AuchtungJM
LeeCA
GarrisonKL
GrossmanAD
2007 Identification and characterization of the immunity repressor (ImmR) that controls the mobile genetic element ICEBs1 of Bacillus subtilis. Mol Microbiol 64 1515 1528
32. BoseB
AuchtungJM
LeeCA
GrossmanAD
2008 A conserved anti-repressor controls horizontal gene transfer by proteolysis. Mol Microbiol 70 570 582
33. LeeCA
AuchtungJM
MonsonRE
GrossmanAD
2007 Identification and characterization of int (integrase), xis (excisionase), and chromosomal attachment sites of the integrative and conjugative element ICEBs1 of Bacillus subtilis. Mol Microbiol 66 1356 1369
34. LeeCA
BabicA
GrossmanAD
2010 Autonomous plasmid-like replication of a conjugative transposon. Mol Microbiol 75 268 279
35. HamoenLW
SmitsWK
de JongA
HolsappelS
KuipersOP
2002 Improving the predictive value of the competence transcription factor (ComK) binding site in Bacillus subtilis using a genomic approach. Nucleic Acids Res 30 5517 5528
36. BaileyTL
BodenM
BuskeFA
FrithM
GrantCE
2009 MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37 W202 208
37. TippnerD
WagnerR
1995 Fluorescence analysis of the Escherichia coli transcription regulator H-NS reveals two distinguishable complexes dependent on binding to specific or nonspecific DNA sites. J Biol Chem 270 22243 22247
38. ShindoH
OhnukiA
GinbaH
KatohE
UeguchiC
1999 Identification of the DNA binding surface of H-NS protein from Escherichia coli by heteronuclear NMR spectroscopy. FEBS Lett 455 63 69
39. LangB
BlotN
BouffartiguesE
BuckleM
GeertzM
2007 High-affinity DNA binding sites for H-NS provide a molecular basis for selective silencing within proteobacterial genomes. Nucleic Acids Res 35 6330 6337
40. LiuY
ChenH
KenneyLJ
YanJ
2010 A divalent switch drives H-NS/DNA-binding conformations between stiffening and bridging modes. Genes Dev 24 339 344
41. DillonSC
DormanCJ
2010 Bacterial nucleoid-associated proteins, nucleoid structure and gene expression. Nat Rev Microbiol 8 185 195
42. DormanCJ
2010 Horizontally acquired homologues of the nucleoid-associated protein H-NS: implications for gene regulation. Mol Microbiol 75 264 267
43. ColangeliR
HelbD
VilchezeC
HazbonMH
LeeCG
2007 Transcriptional regulation of multi-drug tolerance and antibiotic-induced responses by the histone-like protein Lsr2 in M. tuberculosis. PLoS Pathog 3 e87 doi:10.1371/journal.ppat.0030087
44. ChenJM
RenH
ShawJE
WangYJ
LiM
2008 Lsr2 of Mycobacterium tuberculosis is a DNA-bridging protein. Nucleic Acids Res 36 2123 2135
45. StoebelDM
FreeA
DormanCJ
2008 Anti-silencing: overcoming H-NS-mediated repression of transcription in Gram-negative enteric bacteria. Microbiology 154 2533 2545
46. JaacksKJ
HealyJ
LosickR
GrossmanAD
1989 Identification and characterization of genes controlled by the sporulation-regulatory gene spo0H in Bacillus subtilis. J Bacteriol 171 4121 4129
47. HarwoodCR
CuttingSM
1990 Molecular biological methods for Bacillus. Chichester, England John Wiley & Sons
48. BerkmenMB
LeeCA
LovedayEK
GrossmanAD
2010 Polar Positioning of a Conjugation Protein from the Integrative and Conjugative Element ICEBs1 of Bacillus subtilis. J Bacteriol 192 38 45
49. SteinmetzM
RichterR
1994 Plasmids designed to alter the antibiotic resistance expressed by insertion mutations in Bacillus subtilis, through in vivo recombination. Gene 142 79 83
50. YoungmanP
PothH
GreenB
YorkK
OlmedoG
1989 Methods for Genetic Manipulation, Cloning, and Functional Analysis of Sporulation Genes in Bacillus subtilis.
SmithI
SlepeckyRA
SetlowP
Regulation of Procaryotic Development Washington, D.C. ASM Press 65 87
51. WachA
1996 PCR-synthesis of marker cassettes with long flanking homology regions for gene disruptions in S. cerevisiae. Yeast 12 259 265
52. StudierFW
MoffattBA
1986 Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J Mol Biol 189 113 130
53. SmitsWK
GoranovAI
GrossmanAD
2010 Ordered association of helicase loader proteins with the Bacillus subtilis origin of replication in vivo. Mol Microbiol 75 452 461
54. RagkousiK
CowanAE
RossMA
SetlowP
2000 Analysis of nucleoid morphology during germination and outgrowth of spores of Bacillus species. J Bacteriol 182 5556 5562
55. BreierAM
GrossmanAD
2009 Dynamic association of the replication initiator and transcription factor DnaA with the Bacillus subtilis chromosome during replication stress. J Bacteriol 191 486 493
56. EdgarR
DomrachevM
LashAE
2002 Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30 207 210
57. RoyA
KucukuralA
ZhangY
2010 I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5 725 738
58. PrideDT
WassenaarTM
GhoseC
BlaserMJ
2006 Evidence of host-virus co-evolution in tetranucleotide usage patterns of bacteriophages and eukaryotic viruses. BMC Genomics 7 8
Štítky
Genetika Reprodukční medicína
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2010 Číslo 11- Akutní intermitentní porfyrie
- Farmakogenetické testování pomáhá předcházet nežádoucím efektům léčiv
- Růst a vývoj dětí narozených pomocí IVF
- Pilotní studie: stres a úzkost v průběhu IVF cyklu
- Vliv melatoninu a cirkadiálního rytmu na ženskou reprodukci
-
Všechny články tohoto čísla
- Genome-Wide Association Meta-Analysis of Cortical Bone Mineral Density Unravels Allelic Heterogeneity at the Locus and Potential Pleiotropic Effects on Bone
- Beyond QTL Cloning
- An Evolutionary Framework for Association Testing in Resequencing Studies
- Genome-Wide Association Study Identifies Two Novel Regions at 11p15.5-p13 and 1p31 with Major Impact on Acute-Phase Serum Amyloid A
- The Functional Interplay between Protein Kinase CK2 and CCA1 Transcriptional Activity Is Essential for Clock Temperature Compensation in Arabidopsis
- Endogenous Viral Elements in Animal Genomes
- Analysis of the 10q11 Cancer Risk Locus Implicates and in Human Prostate Tumorigenesis
- DNA Methylation and Normal Chromosome Behavior in Neurospora Depend on Five Components of a Histone Methyltransferase Complex, DCDC
- Sarcomere Formation Occurs by the Assembly of Multiple Latent Protein Complexes
- Genetic Basis of Growth Adaptation of after Deletion of , a Major Metabolic Gene
- Nomadic Enhancers: Tissue-Specific -Regulatory Elements of Have Divergent Genomic Positions among Species
- The Parental Non-Equivalence of Imprinting Control Regions during Mammalian Development and Evolution
- CTCF-Dependent Chromatin Bias Constitutes Transient Epigenetic Memory of the Mother at the Imprinting Control Region in Prospermatogonia
- Systematic Dissection and Trajectory-Scanning Mutagenesis of the Molecular Interface That Ensures Specificity of Two-Component Signaling Pathways
- Nucleolin Is Required for DNA Methylation State and the Expression of rRNA Gene Variants in
- The Complex Genetic Architecture of the Metabolome
- ATM Limits Incorrect End Utilization during Non-Homologous End Joining of Multiple Chromosome Breaks
- Mutation Disrupts Synaptonemal Complex Formation, Recombination, and Chromosome Segregation in Mammalian Meiosis
- Mismatch Repair–Independent Increase in Spontaneous Mutagenesis in Yeast Lacking Non-Essential Subunits of DNA Polymerase ε
- The Kinesin-3 Motor UNC-104/KIF1A Is Degraded upon Loss of Specific Binding to Cargo
- Epigenetic Silencing of Spermatocyte-Specific and Neuronal Genes by SUMO Modification of the Transcription Factor Sp3
- A Coastal Cline in Sodium Accumulation in Is Driven by Natural Variation of the Sodium Transporter AtHKT1;1
- Cyclin B3 Is Required for Multiple Mitotic Processes Including Alleviation of a Spindle Checkpoint–Dependent Block in Anaphase Chromosome Segregation
- Altered DNA Methylation in Leukocytes with Trisomy 21
- Human-Specific Evolution and Adaptation Led to Major Qualitative Differences in the Variable Receptors of Human and Chimpanzee Natural Killer Cells
- Leptotene/Zygotene Chromosome Movement Via the SUN/KASH Protein Bridge in
- RACK-1 Acts with Rac GTPase Signaling and UNC-115/abLIM in Axon Pathfinding and Cell Migration
- Genome-Wide Effects of Long-Term Divergent Selection
- Endless Forms Most Viral
- Conflict between Noise and Plasticity in Yeast
- Essential Functions of the Histone Demethylase Lid
- The Transcriptional Regulator Rok Binds A+T-Rich DNA and Is Involved in Repression of a Mobile Genetic Element in
- The Cellular Robustness by Genetic Redundancy in Budding Yeast
- Localization of a Guanylyl Cyclase to Chemosensory Cilia Requires the Novel Ciliary MYND Domain Protein DAF-25
- A Buoyancy-Based Screen of Drosophila Larvae for Fat-Storage Mutants Reveals a Role for in Coupling Fat Storage to Nutrient Availability
- A Functional Genomics Approach Identifies Candidate Effectors from the Aphid Species (Green Peach Aphid)
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Genome-Wide Association Study Identifies Two Novel Regions at 11p15.5-p13 and 1p31 with Major Impact on Acute-Phase Serum Amyloid A
- Analysis of the 10q11 Cancer Risk Locus Implicates and in Human Prostate Tumorigenesis
- The Parental Non-Equivalence of Imprinting Control Regions during Mammalian Development and Evolution
- Genome-Wide Effects of Long-Term Divergent Selection
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MUDr. Irena Krčmová, CSc.
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



