-
Články
- Vzdělávání
- Časopisy
Top články
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Volná místa
Doporučené pozice
Reklama- Praxe
The Major Roles of DNA Polymerases Epsilon and Delta at the Eukaryotic Replication Fork Are Evolutionarily Conserved
Coordinated replication of eukaryotic genomes is intrinsically asymmetric, with continuous leading strand synthesis preceding discontinuous lagging strand synthesis. Here we provide two types of evidence indicating that, in fission yeast, these two biosynthetic tasks are performed by two different replicases. First, in Schizosaccharomyces pombe strains encoding a polδ-L591M mutator allele, base substitutions in reporter genes placed in opposite orientations relative to a well-characterized replication origin are strand-specific and distributed in patterns implying that Polδ is primarily involved in lagging strand replication. Second, in strains encoding a polε-M630F allele and lacking the ability to repair rNMPs in DNA due to a defect in RNase H2, rNMPs are selectively observed in nascent leading strand DNA. The latter observation demonstrates that abundant rNMP incorporation during replication can be tolerated and that they are normally removed in an RNase H2-dependent manner. This provides strong physical evidence that Polε is the primary leading strand replicase. Collectively, these data and earlier results in budding yeast indicate that the major roles of Polδ and Polε at the eukaryotic replication fork are evolutionarily conserved.
Published in the journal: . PLoS Genet 7(12): e32767. doi:10.1371/journal.pgen.1002407
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1002407Summary
Coordinated replication of eukaryotic genomes is intrinsically asymmetric, with continuous leading strand synthesis preceding discontinuous lagging strand synthesis. Here we provide two types of evidence indicating that, in fission yeast, these two biosynthetic tasks are performed by two different replicases. First, in Schizosaccharomyces pombe strains encoding a polδ-L591M mutator allele, base substitutions in reporter genes placed in opposite orientations relative to a well-characterized replication origin are strand-specific and distributed in patterns implying that Polδ is primarily involved in lagging strand replication. Second, in strains encoding a polε-M630F allele and lacking the ability to repair rNMPs in DNA due to a defect in RNase H2, rNMPs are selectively observed in nascent leading strand DNA. The latter observation demonstrates that abundant rNMP incorporation during replication can be tolerated and that they are normally removed in an RNase H2-dependent manner. This provides strong physical evidence that Polε is the primary leading strand replicase. Collectively, these data and earlier results in budding yeast indicate that the major roles of Polδ and Polε at the eukaryotic replication fork are evolutionarily conserved.
Introduction
Three DNA polymerases, Polα, Polδ, and Polε, are required for efficient genome replication in eukaryotes [1], [2]. The Polα holoenzyme complex has both primase activity and DNA polymerase activity and is required to initiate each DNA synthesis reaction. The primase subunit first synthesizes a short RNA primer of ∼10 nucleotides and the DNA polymerase subunit then extends this primer using dNTPs for a further 20–30 nucleotides, thus initiating DNA replication. Polδ or Polε then substitutes for Polα and perform the bulk of DNA replication by elongating these primers.
Genomic DNA is replicated faithfully during every cell cycle with an error rate of approximately 1 in 10−10 errors per base pair, ensuring that the genetic blueprint is transmitted largely unaltered through the generations. In eukaryotic cells, DNA replication is initiated bi-directionally from many replication origins. Because of the antiparallel structure of DNA, one strand (leading strand) is replicated continuously in the same direction of the replication fork, while the second strand (lagging strand) is synthesized discontinuously in the opposite direction to that of replication fork progression. The relatively small (200–1000 base) stretches of DNA synthesized during lagging strand replication are known as Okazaki fragments and are rapidly processed and ligated to complete lagging strand replication. The fidelity of replication is ensured by the nucleotide selectivity of replicases to achieve error rates of 10−4–10−5, by exonucleolytic proofreading during replication to increase fidelity about 100-fold, and by post-replication DNA mismatch repair to further increase fidelity and lower the mutation rate to 10−8–10−10 [3].
Polα, Polδ, and Polε all belong to the B family of DNA polymerases. The structure of the active site of B family DNA polymerases is highly conserved throughout evolution. As for most polymerases, the precise geometry of the polymerase active site ensures that mismatches are largely precluded from incorporation [4]. The importance of polymerase active site geometry to replication fidelity is illustrated by the fact that substitutions of conserved active site residues often reduce DNA synthesis fidelity. Relevant to the present study are substitutions in Saccharomyces cerevisiae Polε and Polδ (M644G and L612M, respectively) that increase error rates during DNA synthesis in vitro and also result in elevated spontaneous mutation rates in vivo [5]–[8]. These polymerases have particular value for studies of replication fidelity in vivo because their error rates are preferentially elevated for only one of two possible mismatches that could result in a particular base substitution in a cell. For example Polδ L612M preferentially generates T-dGTP rather than A-dCTP errors, and this preference yields strand specific A–T to G–C mutations during duplex DNA replication in vivo. These biased error rates result in asymmetric mutation profiles in a URA3 reporter gene that is replicated in only one direction due to its close proximity to an active origin. When present in each of the two possible URA3 orientations relative to the origin, the mutational patterns observed in strains harboring the pol2-M644G (polε) and pol3-L612M (polδ) mutator alleles imply that S. cerevisiae Polε and Polδ are the primary leading strand and lagging strand replicase, respectively [9], [10].
The goal of the present study is to identify the major leading and lagging replicases in the fission yeast Schizosaccharomyces pombe. To investigate Polδ, we took advantage of the fact that both S. cerevisiae Polδ L612M [6] and its human equivalent, Polδ L606M [11], [12] have been shown to have biased DNA synthesis fidelity. Here we report that Schizo. pombe Polδ L591M generates asymmetric mutation profiles in vivo that are consistent with Polδ being the primary lagging strand replicase in Schizo. pombe. To investigate Polε, we attempted to generate a Schizo. pombe Polε mutation (polε-M630G) equivalent to that previously studied in S. cerevisiae (encoding Polε M644G). Schizo. pombe strains with this substitution were not viable. We therefore generated a different allele, polε-M630F, because substitution of phenylalanine at the equivalent active site residues in S. cerevisiae Polα [13] and Polζ [14] are viable and have elevated spontaneous mutation rates. We show here that the Schizo. pombe polε-M630F allele is also viable and a spontaneous mutator. Although it did not display a suitable asymmetric mutation profile for strand assignment, we were able to exploit a second infidelity parameter for strand assignment, the propensity to incorporate rNMP into DNA. Previous studies have demonstrated that during DNA synthesis in vitro and in vivo, S. cerevisiae Polε M644G incorporates greater amounts of rNTPs into DNA than does wild type Polε [15], [16]. Here we exploit this same promiscuity with the Schizo. pombe polε-M630F mutant, to provide a physical demonstration that the majority of leading strand synthesis in Schizo. pombe is performed by Polε.
Results
Approach
Which DNA polymerase replicates which strand has only been determined in the budding yeast S. cerevisiae [9], [10]. We thus wished to determine if this division of labour between the main replicative polymerases is conserved in the distantly related eukaryote, the fission yeast Schizo. pombe. Our strategy was to establish the direction of replication for a specific locus, to create mutants in the genes encoding two replicative polymerases, Polδ and Polε, that exhibit specific and characteristic profiles of misincorporation, and to use these to assign each polymerase to one or the other strand (or both) based on the profile of misincorporation at the directionally replicated loci.
The catalytic subunits of Polδ or Polε are encoded by the cdc6 (pol3) and the cdc20 (pol2) genes, respectively. For clarity, here we simply refer to them as polδ and polε. We employed recombination-mediated cassette exchange (RMCE) to create strains that harbor each specific mutant polymerase [17]. Mutant genes introduced into the genome by this method are flanked by lox (P and M3) sequences. Thus, we also created control strains (pol+) that have the gene encoding the wild-type polymerase flanked by the same lox sites.
Direction of DNA Replication at the ura4 Locus
The Schizo. pombe ura4+ gene allows for both positive and negative selection. Selecting for loss of ura4 function is achieved by growth on medium containing 5-fluoroorotic acid (5-FOA), which identifies loss-of-function mutants. However, mutations in either the ura4 or the ura5 genes of Schizo. pombe confer 5-FOA resistance, and it has been reported that greater than 50% of spontaneously arising 5-FOA resistant clones harbor mutations in ura5 [18]. In wild type cells, ura4+ is located on chromosome III while ura5+ is located on chromosome II. Therefore, to efficiently identify mutations at a single chromosomal location that confer 5-FOA resistance, we created two artificial loci where ura5+ was placed adjacent to ura4+ on chromosome III. These differ only in the orientation of the ura4+:ura5+ fragment (Figure 1A). We confirmed that this novel ura5+:ura4+ fragment does not function as a replication origin by demonstrating it would not support maintenance of plasmid sequence in cells. We also deleted the genomic ura5+ gene on chromosome II, so that resultant ura4+:ura5+ Δura5 strains have only one copy of the ura4+ and ura5+ genes.
Fig. 1. Polδ L591M shows mutagenic strand bias. 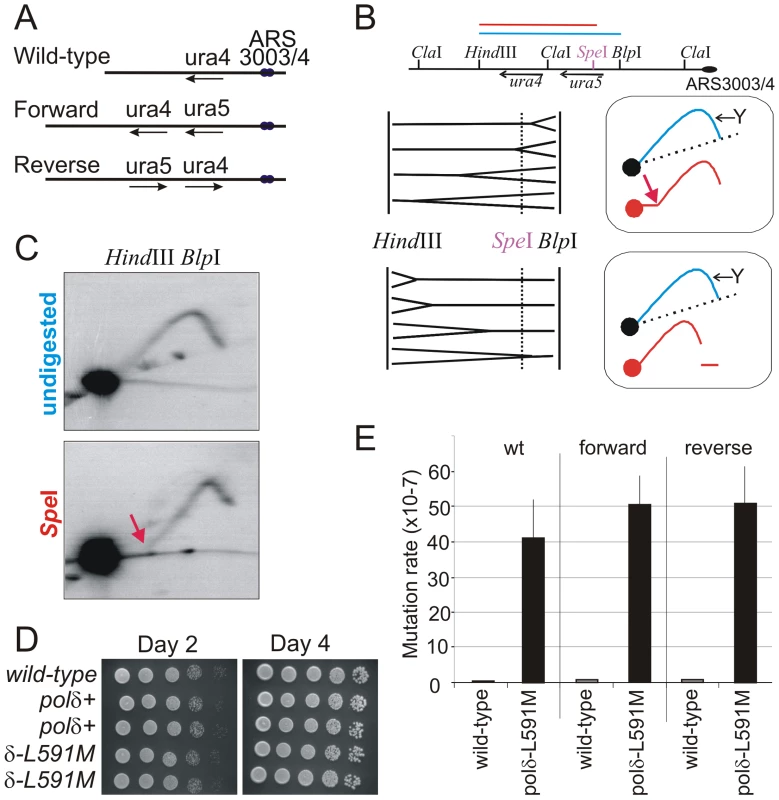
A. Schematic of the wild type ura4+ locus and the two versions of the modified ura4+:ura5+ loci. Forward: the transcribed strand of both ura4 and ura5 corresponds to the lagging strand when replicated from ars3003/3004. Reverse: the orientation of the ura4:ura5 sequences are switched so that the transcribed strand corresponds to the leading strand when replicated from ars3003/3004. Loss of either ura4 or ura5 function results in 5-FOA resistance. B. The direction of replication at the modified ura4+ locus. Top: Schematic of the ura4+:ura5+ locus. Bottom: the principle of directional 2D-gel analysis. Asymmetric digestion of the HindIII-BlpI fragment with SpeI between the running of the first and second dimensions will result in a shift of the Y-arc. The position of the Y arc is indicated by an arrow. The direction of shift is dependent on the direction of replication. C. Comparison of replication intermediates within the HindIII-BlpI region either without (undigested) or after SpeI digestion between dimensions (SpeI). The shifted Y arc following SpeI digestion is indicated with a red arrow, the equivalent of which is also shown in the top frame of panel B. The majority of replication forks run from right to left, corresponding with the efficient initiation from ars3003/ars3004 on the right. D. Cell growth of wild type, polδ+ (wild-type polδ flanked by lox sites) and polδ-L591M. Serial dilutions of cells were spotted on YEA plates, incubated (30°C) for 2 or 4 days and photographed. E. Spontaneous mutation rates in each of the indicated backgrounds for polδ+ and polδ-L591M. Error bars show standard deviation. The ura4+:ura5+ locus is on Chromosome III, near two autonomous replicating sequences; ars3003/3004. Both the ars3003 and ars3004 sequences have been well characterized and are known to be highly efficient at initiating replication [19], [20]. However, more than 50% of Schizo. pombe intergenic regions have the potential to function as origins of replication [21]. Thus, to experimentally determine the direction of DNA replication at the ura4+:ura5+ locus, we employed the method of directional 2-D gel electrophoresis [22]. DNA from an asynchronous population of cells is first digested with HindIII and BlpI and fragments separated in the first dimension without ethidium bromide. The lane is then excised and digested with SpeI, which cleaves within the HindIII-BlpI fragment containing the ura4+ and ura5+ genes. This DNA is then subjected to the second dimension of electrophoresis in the presence of ethidium bromide and DNA in the gel is transferred to a membrane for Southern blot analysis with the ura4-containing HindIII-SpeI fragment. The results revealed the direction of DNA replication, as illustrated in Figure 1B. Most of detectable replication intermediates show the pattern consistent with DNA replication moving from right to left (Figure 1C, see red arrow bottom panel: its equivalent is similarly indicated in the top panel of Figure 1B). Thus, we conclude that a leftward replication fork replicates the ura4+:ura5+ locus in the majority of cells.
Characterization of a polδ-L591M Mutant
We then created the polδ-L591M mutant using RMCE. Schizo. pombe Polδ L591 is equivalent to S. cerevisiae Polδ L612. polδ-L591M cells grow as well as wild type cells (Figure 1D), demonstrating that this mutant of Polδ is proficient for DNA replication in vivo. In wild type and ura4+:ura5+ Δura5 backgrounds, polδ-L591M showed a strong mutator phenotype (Figure 1E). Spontaneous mutation rates are elevated ∼100 fold in polδ-L591M (4–5×10−6/cell division) compared with that in polδ+ (4–7×10−8).
Mutational Bias in polδ L591M Strains
The elevated mutation rates indicate that most of the mutations seen in polδ -L591M cells reflect the error specificity of this mutant polymerase, rather than background mutations. As shown in Table 1, more than half of mutations were point mutations, consistent with elevated base misincorporation observed in vivo for the equivalent S. cerevisiae strain and in vitro for the corresponding mutant version (L612M) of S. cerevisiae Polδ [5], [6], [23] and human (L606M) Polδ [11], [12]. In addition to point mutations, we observed a variety of duplication and deletion mutations. All of these deletions and duplications were observed at repetitive DNA sequences. More than half of the deletions were >100 bp, while the majority of duplications were <100 bp (Table S1). Possible mechanisms by which such mutations may arise are addressed in the Discussion.
Tab. 1. 5-FOA resistant mutants from polδ-L591M. 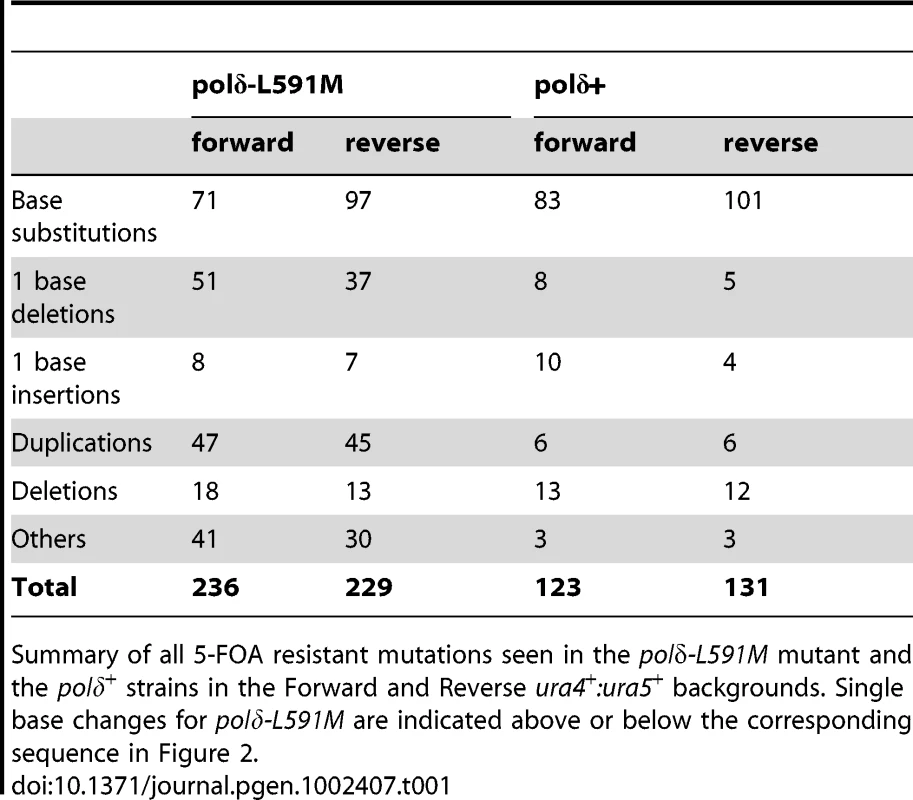
Summary of all 5-FOA resistant mutations seen in the polδ-L591M mutant and the polδ+ strains in the Forward and Reverse ura4+:ura5+ backgrounds. Single base changes for polδ-L591M are indicated above or below the corresponding sequence in Figure 2. Among the point mutations, transition mutations showed significant strand dependence for misincorporation. Figure 2 and Table 2 show the predicted mispairs formed during synthesis of the transcribed strand, which corresponds to lagging or leading strand synthesis in the Forward or Reverse strains, respectively (illustrated in Figure 2A). A:T to G:C mutations can result from either A:dCTP mismatches or T:dGTP mismatches. Depending on which template strand is copied by the mutated polymerase, this will give a bias of mutation resulting in a spectra dependent on the orientation of the DNA sequence (see Figure 2B). We observed that, for A:T to G:C changes, T:dG mispairing is 12.5 fold more frequent than A:dC mispairing in the Forward strain, while A:dC is more frequent in the Reverse strain. Since the misincorporation rate of the corresponding mutant S. cerevisiae and human polymerases are much higher for T:dG than for A:dC [6], [12], the results in Figure 2 imply that Polδ preferentially replicates the lagging strand template. A similar bias was also observed for G:C to A:G mutations. G:dT is ∼3 fold higher than C:dA in the Forward strain while C:dA is ∼3 fold higher in the Reverse strain. Comparing these data with the published in vitro results is also consistent with Polδ being responsible for replicating the lagging strand template. Strand dependence was not observed in polδ+ (Table 3), indicating that the bias seen in polδ -L591M cells reflects base misincorporation by the mutant polymerase rather than sequence context or the transcriptional direction of marker genes. We did not observe strong hotspots for particular mutations, but the total number of occurrences is higher for some mutations, e.g., T to C at ura4 base pair 236 and 76 in the Forward background and C to T at ura4 190 and (−91) in the Reverse background (Figure 3).
Fig. 2. Strand bias for polδ-L591M. 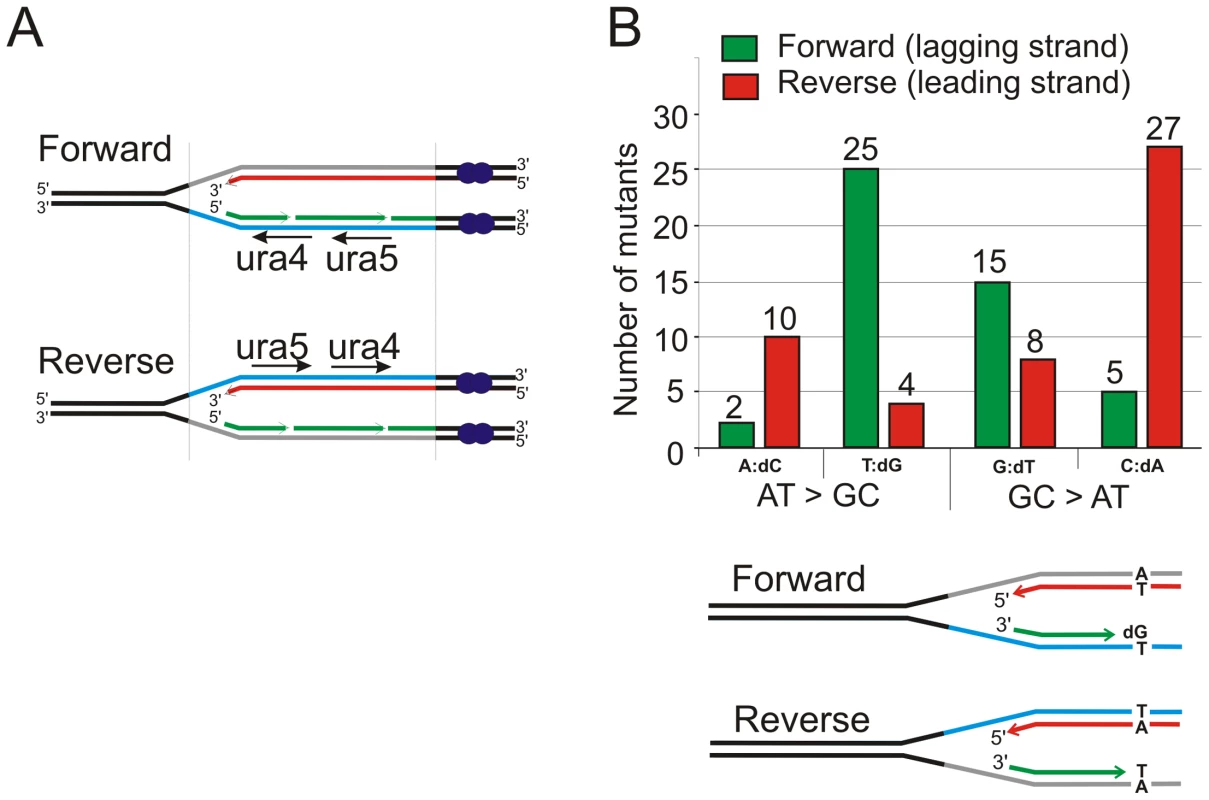
A. Schematic of replication through the Forward and Reverse ura4+:ura5+ loci. Leading strand synthesis shown in red, lagging strand synthesis is shown in green. The transcribed strand is shown in blue for reference. B. Top: the relative numbers of AT>GC and GC>AT mutations identified, classified as resulting from either A:dC or T:dG mispairing (AT>GC) and G:dT or C:dA mispairing (GC>AT). Bottom. Schematic illustration of replication of a specific A:T base pair in both orientations. If the lagging strand polymerase, but not the leading strand polymerase, is prone to mispairing dG opposite T during incorporation, but not dC opposite A, then this A:T base pair will mutate to G:C more frequently in the forward orientation than the reverse orientation. Fig. 3. Mutation spectra. 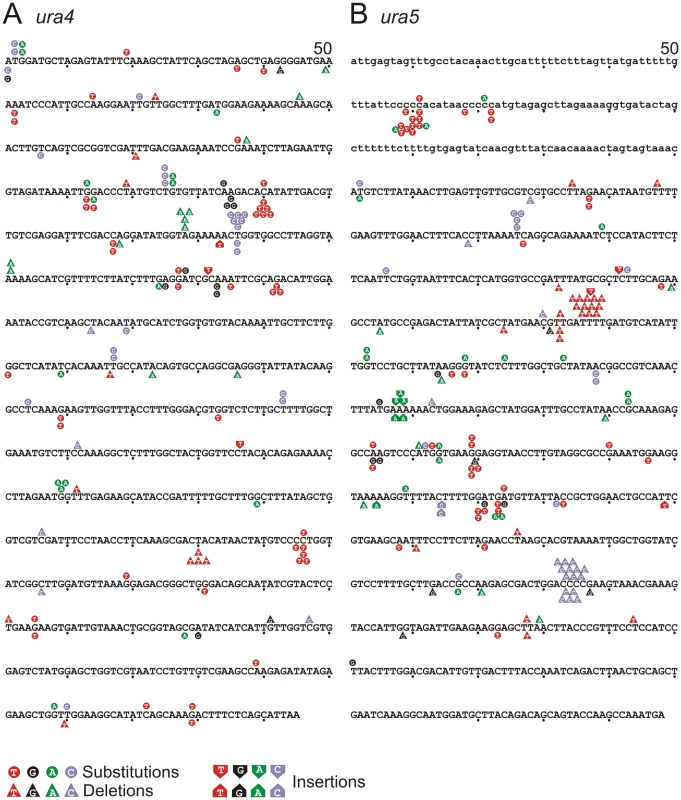
A. ura4 and B. ura5. The promoter region of ura5 (lower case) and the ORF's of both rad4 and ura5 (upper case) are shown. The position of each mapped mutation is indicated. Mutations arising in the “forward” background (transcribed strand replicated by lagging strand synthesis) are shown above the sequence, whereas the mutations arising in the “reverse” background (transcribed strand replicated by leading strand synthesis) are shown below the sequence. Tab. 2. Strand bias of mutants from polδ-L591M. 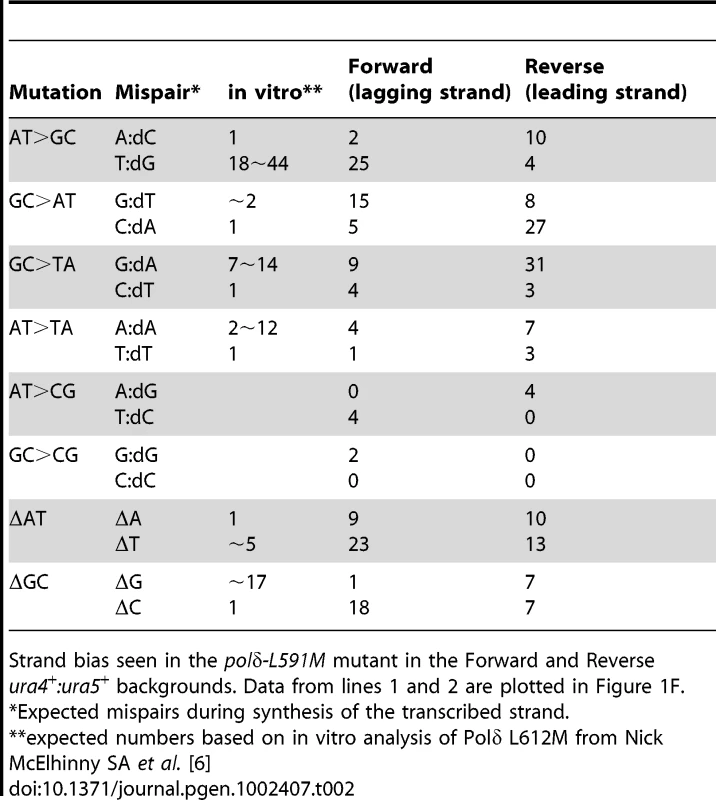
Strand bias seen in the polδ-L591M mutant in the Forward and Reverse ura4+:ura5+ backgrounds. Data from lines 1 and 2 are plotted in Figure 1F. Tab. 3. Lack of strand bias of mutants from polδ+. 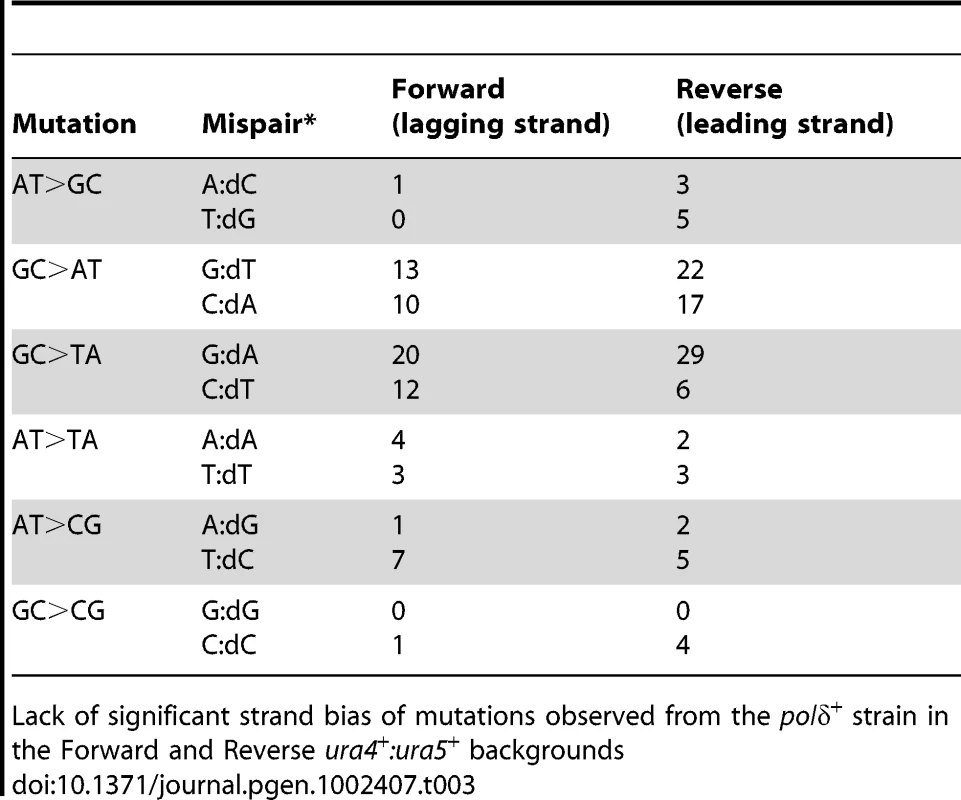
Lack of significant strand bias of mutations observed from the polδ+ strain in the Forward and Reverse ura4+:ura5+ backgrounds Characterization of the polε-M630F Mutant
S. cerevisiae Polε M644G shows strong bias between A:dA and T:dT mispairs in vitro and the spontaneous mutation rates of the corresponding mutant cells are significantly higher than that of wild type and exhibit strand bias [8]. However, we found that the equivalent Schizo. pombe polε-M630G mutation is lethal, as was a polε-M630K mutation. Analysis of strains expressing Polε M630G or Polε-M630K from an ectopic integrated copy in a polε+ background (Figure S1) suggest this is largely due to catalytic inactivity, as mutation frequencies were not dramatically increased. Thus, we created strains harboring polε-M630F as an alternative. The decision to substitute to phenylalanine was based on earlier studies showing that Polα L868F is error prone in vitro and mutagenic in vivo [13], Polε M644F is error-prone in vitro with a weak bias in error rates [7], and Polζ L979F is error prone in vitro [24] and mutagenic in vivo [14]. The Schizo. pombe polε-M630F that we created using the RMCE methodology grows slightly more slowly than polε+, although the size of mutant colonies becomes comparable to that of wild type after prolonged incubation (Figure 4A). Strains harboring polε-M630F did not exhibit a substantial increase in spontaneous mutation rate in a mismatch repair proficient background. In strains wherein mismatch repair is inactivated by deleting the msh2 gene, polε-M630F increased the mutation rate by 4–5 fold (Figure 4C). However, upon sequencing ura5 and ura4 from 5-FOA resistant clones, a strand bias sufficient to infer which strand is copied by the mutant Polε was not observed (Table S2).
Fig. 4. Mutation rate and dNMP incorporation by polε-M630F. 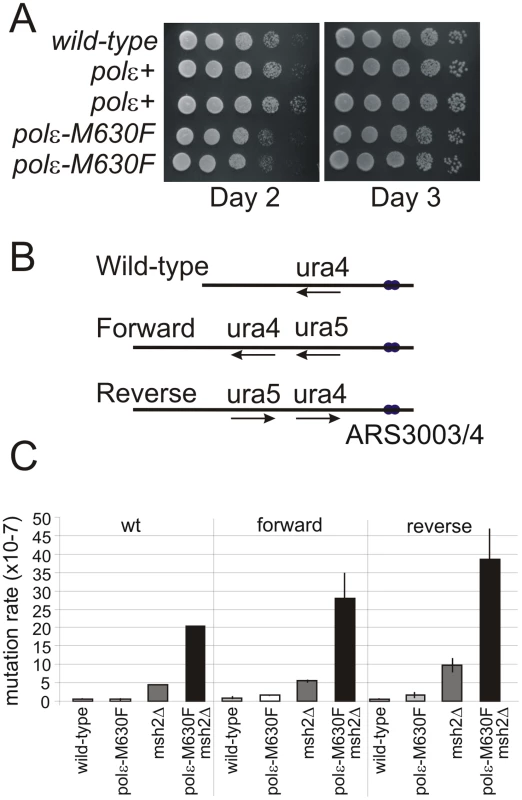
A. Cell growth of wild type cells and polε+ (wild-type polε flanked by lox sites) and polε-M630F strains. Serial dilutions of cells were spotted on YEA plates, incubated (30°C) for 2 or 3 days and photographed. B. Schematic of the wild type ura4+ locus and the two versions of the modified ura4+:ura5+ loci. C. Spontaneous mutation rates for polε+, msh2Δ, polε-M630F and polε-M630F msh2Δ double mutant cells in the ura4+ (wild type), Forward and Reverse backgrounds. Error bars are standard deviations. rNMP Incorporation into DNA by Polε M630F
Mutations at Polε M644 in S. cerevisiae affect the rate of rNMP incorporation into DNA [15]. We thus tested this possibility in Schizo. pombe. rNMPs incorporated into DNA are rapidly excised by the activity of RNase H2, whose catalytic subunit is encoded by the rnh201 gene of Schizo. pombe. Since increased rNMP incorporation increases alkali-dependent DNA fragmentation, we assayed for increased gel mobility of DNA from the endogenous ura4+ locus using Southern blot analysis. As anticipated, genomic DNA prepared from polε-M630F was not particularly sensitive to alkali treatment when compared to genomic DNA from the polε+ strain (Figure 5A, lanes 1 and 2). However, it becomes significantly sensitive compared to polε+ when rnh201 is deleted (lanes 3 and 4). This indicates that Polε M630F incorporates rNMP into DNA at higher rate than wild type Polε and that these are largely removed by RNase H2 activity.
Fig. 5. Strand bias for rNMP incorporation. 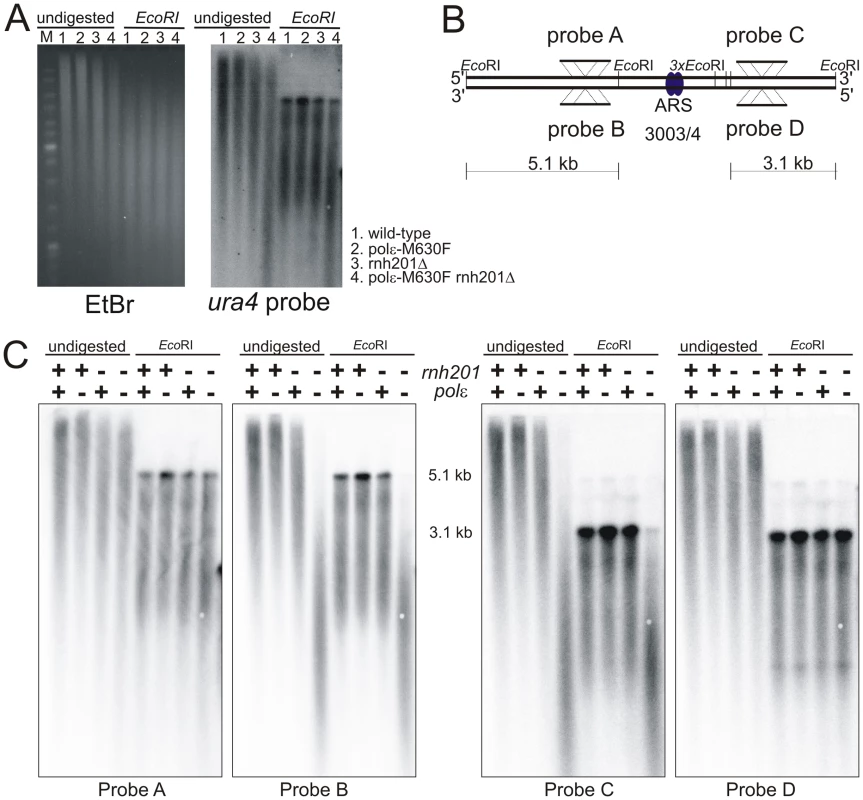
A. Alkali sensitivity of genomic DNA. Genomic DNA from the indicated strains was either digested with EcoRI or left undigested, then treated with alkali and separated by alkaline agarose gel electrophoresis. DNA was revealed with ethidium bromide following neutralization and then processed for Southern analysis and probed with a ura4 probe that reveals both DNA strands. B. Schematic of the loci either side of ars3003/3004 indicating the positions of the EcoRI sites, plus the location and strand specificity of the probes used. C. Alkali sensitivity of each strand, either on the left of ars3003/3004 (probes A and B) or on the right (probes C and D). Strains were either polε+ (+) or polε-M630F (−) with or without concomitant deletion of rnh201, as indicated. The membrane from Figure 3D was stripped and hybridized with the indicated single stranded probes. Probe A and C hybridize with the top strand, probes B and D hybridize with the bottom strand. Based on this observation, we chose to test the strand specificity of rNMP incorporation using alkali treatment and subsequent probing for either the leading or lagging strand using the appropriate single-stranded probes. We prepared two pairs of probe across ars3003/3004 (Figure 5B). The top strand is detected by probe A and C, while the bottom strand is detected by probe B and D. As shown in Figure 5C, only one of each of the two strands from rnh201Δ polε-M630F was sensitive to alkali at each probed site. The alkali sensitive strand was the bottom strand on the left side of the origin, while the top was sensitive on the right side (Figure 5B and 5C). Since those probed sites are inferred to be copied by replication forks emerged at ars3003/3004, the alkali-sensitive strands correspond to the nascent leading strand products of replication. Similar results were obtained at another origin (Figure S2). These results strongly suggest that Polε replicates the leading strand template.
Discussion
An understanding of the fundamental mechanism of DNA replication is an important aspect of appreciating how replication and the errors made during replication influence evolution and human disease. While we have a breadth of knowledge about the proteins involved in eukaryotic DNA replication, including those that move with the active replisome, we do not have an unambiguous view of the architecture of the replication machine itself. Indeed, only recently has genetic data from the budding yeast S. cerevisiae linked the key replicative polymerases Polε and Polδ to leading and lagging strand synthesis, respectively. While these assignments are consistent with a number of additional observations, such as the role of Polδ in the maturation of lagging strands [25]–[27], it is important to provide additional evidence to reinforce these assignments, as well as to establish if they are evolutionarily conserved.
Lagging Strand Synthesis by Polδ
To investigate the role of Schizo. pombe Polδ during DNA replication, we created strains that replicate using a Polδ L591M mutant protein. We showed that Polδ L591M is highly mutagenic and induced various types of mutations in Schizo. pombe. Strand dependence in transition mutations allowed us to conclude that the main role of Polδ is during lagging strand synthesis (Figure 2 and Table 2). However, the mutational bias seen in this mutant is weaker than would be predicted from the in vivo and in vitro studies of the equivalent S. cerevisiae mutant. Because we used mismatch repair proficient cells for this study (the double mutant was lethal), the mutation spectra we observed here reflect mispairs that have escaped mismatch detection and repair. This may influence our interpretations. For example, bacterial MutS protein has variable affinity for different mismatches, with G:T being one of the best substrates [28], [29]. Thus, the specificity of mismatch repair might have partially masked the bias of misincorporation induced by Polδ L591M. It is also possible that the mutation spectra were affected by spontaneous base damage that results in mismatches that escape mismatch repair. These caveats mean that, while our results are consistent with a function of Polδ as a lagging strand polymerase, we cannot exclude the possibility that Polδ partly participates in leading strand synthesis or that Polε (or indeed other polymerases) may partially replicate the lagging strand [10], [30].
In addition to point mutations that were expected from in vitro studies of S. cerevisiae and human polymerases, we also observed significantly enhanced formation of deletions and duplications in polδ -L591M cells (Table 1). All deletions and duplications occurred at repetitive DNA sequences. The majority of duplications involved <100 bases (Table S1), reminiscent of the mutation spectra for S. cerevisiae rad27 mutants. Jin et al. showed that duplication rates were enhanced by mutations in the Polδ exonuclease domain [25] and S. cerevisiae polδ -L612M cells require functional Rad27 for viability [31]. These studies are consistent with our observations and add support to the premise that Polδ is involved directly in lagging strand synthesis in Schizo. pombe.
The size of deletions we observed was relatively larger than that of duplications. More than half of the deletions were loss of >100 bp of sequence. Cai et al. have observed that exonuclease deficient E. coli DNA polymerase II generates similar deletions flanked by direct repeat sequences [32]. They proposed a model in which a mismatch made by a mutator polymerase during replication of the first direct repeat promotes primer relocation to the second direct repeat. Furthermore, we observed a low frequency of inversions flanked by inverted repeat sequences and most of these inversions were associated with deletion, duplication, and/or gene conversion. These events can be explained by template switching. Taken together, these observations suggest that a mismatch formed during DNA replication can cause various kinds of genome rearrangements. Interestingly, chromosome abnormalities such as chromatid breaks are substantially elevated in Pold1+/L604G and Pold1+/L604K mouse cells [33].
Leading Strand Synthesis by Polε
To investigate a role of Polε during normal DNA replication, we utilized the observation that S. cerevisiae Polε M644G increased rNMP incorporation [15], [34]. We first demonstrated that Schizo. pombe polε-M630F cells incorporate rNMP into DNA at higher frequency than polε+ cells (Figure 5A). This property of the mutant polymerase made it possible to determine the strand that is copied by the mutant Polε. Incorporation of rNMP in the leading strand was strikingly higher in polε-M630F mutant cells compared to polε+ cells (Figure 5B and Figure S1). This result strongly suggests that Polε synthesizes the leading strand. On the other hand, we failed to observe a significant difference in rNMP incorporation in the lagging strand. This suggests that Polε has, at most, a limited role in lagging strand synthesis.
Schizo. pombe cells that harbor polε-M630G were not viable, while the corresponding mutation does not cause lethality in S. cerevisiae. Interestingly, the N-terminal catalytic domain of Polε can be entirely deleted in both yeasts [35], [36], while a catalytically dead Polε, that retains the full-length protein, is inviable. Our mutation frequency analysis of cells expressing Polε-M630G in a polε+ background (Figure S1) suggest the inviability of polε-M630G is because the corresponding protein is catalytically dead, rather than because it increases the mutation burden beyond that which is sustainable.
Incorporation and Repair of rNMPs in DNA
In addition to supporting a role for Polε in leading strand replication, the results in Figure 5 extend to Schizo. pombe two important conclusions derived from earlier studies in S. cerevisiae, namely that large numbers of rNTPs can be incorporated into the nascent leading strand during replication without strongly affecting growth (Figure 4A) and the rNMPs that are stably incorporated into the Schizo. pombe genome by a eukaryotic replicase are efficiently repaired in a RNase H2-dependent manner. In S. cerevisiae, unrepaired rNMPs in DNA promote formation of short deletions between short, tandemly repeated DNA sequences, by a mechanism that is unaffected by mismatch repair status [34] and is initiated by topoisomerase 1-dependent cleavage of rNMPs [37]. Many of the deletions occur in a manner that depends on the orientation of the reporter gene in relation to the closest origin of replication [15], indicating that they result from rNMPs incorporated into the nascent leading strand by Polε. The characteristics of the Schizo. pombe polε-M630F strains described here offer the opportunity to determine if these consequences are conserved in fission yeast, and to also test whether mating type switching, which depends on rNMPs in DNA [38], is affected by increased rNMP incorporation by replicases and/or by RNase H2 or topoisomerase status.
DNA Replication in Schizo. pombe
In this study, we examined roles of Polδ and Polε during normal DNA replication in Schizo. pombe using two different methods. The first method was a genetic analysis of mutation spectra asymmetry in polδ mutant cells. The second was a physical rNMP incorporation assay using polε mutant cells. The combination of these analyses indicates that genomic DNA is replicated in Schizo. pombe in similar manner as has been suggested for S. cerevisiae. Because Schizo. pombe and S. cerevisiae are highly diverged in evolutionary terms [39], [40] our results strengthen the interpretation that replication in all eukaryotes follows similar rules. We also add a physical assay to the previous genetic data, increasing the likelihood that the interpretation of the genetics is indeed correct. We mainly examined DNA replication at the genomic ura4 locus, because replication initiation at this locus is known to be highly efficient (Figure 1B). However, a similar result was obtained for a second independent locus using the physical method for assigning Polε activity (Figure S1). Thus, it is reasonable to suggest that DNA replication occurs in similar manner throughout the genome. However, it remains possible that cells utilize these two polymerases in a different manner in some specific situations or at some specific loci.
Materials and Methods
Schizo. pombe Strains, Media, and Methods
Schizo. pombe cells were grown in yeast extract (YE) medium. Standard genetic and molecular procedures were employed as described previously [41]. To examine cell growth on plates, serial dilutions of cells were spotted on YEA (YE agar) plates, and incubated at 30°C.
Generating DNA Polymerase Mutant Strains
The cdc6+ and cdc20+ genes were amplified by PCR and cloned into pUC19. cdc6-L591F and cdc20-M630F mutant genes were constructed by PCR-meditated site-directed mutagenesis and sequenced to ensure that only the desired mutation was introduced. Both wild-type and mutant genes were introduced into Schizo. pombe at their native loci by recombination-mediated cassette exchange (RMCE) [17].
Determining Spontaneous Mutation Rates
Spontaneous mutation rates were determined by fluctuation assay as described previously [42]. Briefly, 11 independent single colonies were suspended in 5 ml YEP (YE+polypeptone) medium and grown to saturation at 30°C. Cells were diluted appropriately and plated on YEA or YEA containing 0.1% 5-fluoroorotic acid (5-FOA). Colonies were counted after 4 days incubation at 30°C. Mutation rates were calculated by the method of median [43]. Genomic DNA from a single 5-FOA resistant colony was isolated and the ura5-ura4 construct was amplified by PCR to be sequenced.
2-D Gel Analysis
Directional 2-D gel analysis was performed as described previously [22] with modifications. Genomic DNA was extracted and digested with HindIII and BlpI as described in [44]. After the first dimension electrophoresis, DNA was digested with SpeI in a gel slice and subjected to the second dimension electrophoresis. Replication intermediates were detected by Southern blot.
Detecting Alkali-Sensitive Sites in Genomic DNA
Genomic DNA was extracted from exponentially growing cells and purified by Qiagen genomic-tip 100/G. 5 µg of undigested or EcoRI digested DNA was incubated in 0.3 M NaOH at 55°C for 2 hours and subjected to 1% alkaline agarose gel electrophoresis [15]. Gels were neutralized and stained with ethidium bromide, followed by Southern blot.
Southern Blotting
Southern blotting was performed according to [45]. DNA fragments of interest were amplified by PCR from Schizo. pombe genomic DNA and used as templates to obtain labeled probes. Radioactive nucleotides were incorporated into DNA using Ready-To-Go DNA Labeling beads (GE Healthcare) or strand specific primers and TaKaRa Ex Taq (TAKARA BIO).
Supporting Information
Zdroje
1. GargPBurgersPM 2005 DNA polymerases that propagate the eukaryotic DNA replication fork. Crit Rev Biochem Mol Biol 40 115 128
2. JohnsonAO'DonnellM 2005 Cellular DNA replicases: components and dynamics at the replication fork. Annu Rev Biochem 74 283 315
3. KunkelTA 2009 Evolving views of DNA replication (in)fidelity. Cold Spring Harb Symp Quant Biol 74 91 101
4. KunkelTABebenekK 2000 DNA replication fidelity. Annu Rev Biochem 69 497 529
5. Nick McElhinnySAGordeninDAStithCMBurgersPMKunkelTA 2008 Division of labor at the eukaryotic replication fork. Mol Cell 30 137 144
6. Nick McElhinnySAStithCMBurgersPMKunkelTA 2007 Inefficient proofreading and biased error rates during inaccurate DNA synthesis by a mutant derivative of Saccharomyces cerevisiae DNA polymerase delta. J Biol Chem 282 2324 2332
7. PursellZFIsozILundstromEBJohanssonEKunkelTA 2007 Regulation of B family DNA polymerase fidelity by a conserved active site residue: characterization of M644W, M644L and M644F mutants of yeast DNA polymerase epsilon. Nucleic Acids Res 35 3076 3086
8. PursellZFIsozILundstromEBJohanssonEKunkelTA 2007 Yeast DNA polymerase epsilon participates in leading-strand DNA replication. Science 317 127 130
9. BurgersPM 2009 Polymerase dynamics at the eukaryotic DNA replication fork. J Biol Chem 284 4041 4045
10. KunkelTABurgersPM 2008 Dividing the workload at a eukaryotic replication fork. Trends Cell Biol 18 521 527
11. SchmittMWVenkatesanRNPillaireMJHoffmannJSSidorovaJM 2010 Active site mutations in mammalian DNA polymerase delta alter accuracy and replication fork progression. J Biol Chem 285 32264 32272
12. SchmittMWMatsumotoYLoebLA 2009 High fidelity and lesion bypass capability of human DNA polymerase delta. Biochimie 91 1163 1172
13. NiimiALimsirichaikulSYoshidaSIwaiSMasutaniC 2004 Palm mutants in DNA polymerases alpha and eta alter DNA replication fidelity and translesion activity. Mol Cell Biol 24 2734 2746
14. SakamotoANStoneJEKisslingGEMcCullochSDPavlovYI 2007 Mutator alleles of yeast DNA polymerase zeta. DNA Repair (Amst) 6 1829 1838
15. Nick McElhinnySAKumarDClarkABWattDLWattsBE 2010 Genome instability due to ribonucleotide incorporation into DNA. Nat Chem Biol 6 774 781
16. Nick McElhinnySAWattsBEKumarDWattDLLundstromEB 2010 Abundant ribonucleotide incorporation into DNA by yeast replicative polymerases. Proc Natl Acad Sci U S A 107 4949 4954
17. WatsonATGarciaVBoneNCarrAMArmstrongJ 2008 Gene tagging and gene replacement using recombinase-mediated cassette exchange in Schizosaccharomyces pombe. Gene 407 63 74
18. FraserJLNeillEDaveyS 2003 Fission yeast Uve1 and Apn2 function in distinct oxidative damage repair pathways in vivo. DNA Repair (Amst) 2 1253 1267
19. DubeyDDZhuJCarlsonDLSharmaKHubermanJA 1994 Three ARS elements contribute to the ura4 replication origin region in the fission yeast, Schizosaccharomyces pombe. EMBO J 13 3638 3647
20. PatelPKArcangioliBBakerSPBensimonARhindN 2006 DNA replication origins fire stochastically in fission yeast. Mol Biol Cell 17 308 316
21. DaiJChuangRYKellyTJ 2005 DNA replication origins in the Schizosaccharomyces pombe genome. Proc Natl Acad Sci U S A 102 337 342
22. FriedmanKLBrewerBJ 1995 Analysis of replication intermediates by two-dimensional agarose gel electrophoresis. Methods Enzymol 262 613 627
23. VenkatesanRNHsuJJLawrenceNAPrestonBDLoebLA 2006 Mutator phenotypes caused by substitution at a conserved motif A residue in eukaryotic DNA polymerase delta. J Biol Chem 281 4486 4494
24. StoneJEKisslingGELujanSARogozinIBStithCM 2009 Low-fidelity DNA synthesis by the L979F mutator derivative of Saccharomyces cerevisiae DNA polymerase zeta. Nucleic Acids Res 37 3774 3787
25. JinYHObertRBurgersPMKunkelTAResnickMA 2001 The 3′→5′ exonuclease of DNA polymerase delta can substitute for the 5′ flap endonuclease Rad27/Fen1 in processing Okazaki fragments and preventing genome instability. Proc Natl Acad Sci U S A 98 5122 5127
26. GargPStithCMSabouriNJohanssonEBurgersPM 2004 Idling by DNA polymerase delta maintains a ligatable nick during lagging-strand DNA replication. Genes Dev 18 2764 2773
27. PavlovYIFrahmCNick McElhinnySANiimiASuzukiM 2006 Evidence that errors made by DNA polymerase alpha are corrected by DNA polymerase delta. Curr Biol 16 202 207
28. BrownJBrownTFoxKR 2001 Affinity of mismatch-binding protein MutS for heteroduplexes containing different mismatches. Biochem J 354 627 633
29. SuSSLahueRSAuKGModrichP 1988 Mispair specificity of methyl-directed DNA mismatch correction in vitro. J Biol Chem 263 6829 6835
30. PavlovYIShcherbakovaPV 2010 DNA polymerases at the eukaryotic fork-20 years later. Mutat Res 685 45 53
31. LiLMurphyKMKanevetsUReha-KrantzLJ 2005 Sensitivity to phosphonoacetic acid: a new phenotype to probe DNA polymerase delta in Saccharomyces cerevisiae. Genetics 170 569 580
32. CaiHYuHMcEnteeKKunkelTAGoodmanMF 1995 Purification and properties of wild-type and exonuclease-deficient DNA polymerase II from Escherichia coli. J Biol Chem 270 15327 15335
33. VenkatesanRNTreutingPMFullerEDGoldsbyRENorwoodTH 2007 Mutation at the polymerase active site of mouse DNA polymerase delta increases genomic instability and accelerates tumorigenesis. Mol Cell Biol 27 7669 7682
34. ClarkABLujanSAKisslingGEKunkelTA 2011 Mismatch repair-independent tandem repeat sequence instability resulting from ribonucleotide incorporation by DNA polymerase varepsilon. DNA Repair (Amst) 10 476 482
35. KestiTFlickKKeranenSSyvaojaJEWittenbergC 1999 DNA polymerase epsilon catalytic domains are dispensable for DNA replication, DNA repair, and cell viability. Mol Cell 3 679 685
36. FengWD'UrsoG 2001 Schizosaccharomyces pombe cells lacking the amino-terminal catalytic domains of DNA polymerase epsilon are viable but require the DNA damage checkpoint control. Mol Cell Biol 21 4495 4504
37. KimNHuangSNWilliamsJSLiYCClarkAB 2011 Mutagenic processing of ribonucleotides in DNA by yeast topoisomerase I. Science 332 1561 1564
38. VengrovaSDalgaardJZ 2004 RNase-sensitive DNA modification(s) initiates Schizo. pombe mating-type switching. Genes Dev 18 794 804
39. KuramaeEERobertVSnelBBoekhoutT 2006 Conflicting phylogenetic position of Schizosaccharomyces pombe. Genomics 88 387 393
40. SipiczkiM 2000 Where does fission yeast sit on the tree of life? Genome Biol 1 REVIEWS 1011 1011.1014
41. MorenoSKlarANurseP 1991 Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol 194 795 823
42. FosterPL 2006 Methods for determining spontaneous mutation rates. Methods Enzymol 409 195 213
43. Lea DaCCA 1949 The distribution of the numbers of mutants in bacterial populations. J Genetics 49 264 285
44. ArcangioliB 1998 A site - and strand-specific DNA break confers asymmetric switching potential in fission yeast. EMBO J 17 4503 4510
45. LambertSWatsonASheedyDMMartinBCarrAM 2005 Gross chromosomal rearrangements and elevated recombination at an inducible site-specific replication fork barrier. Cell 121 689 702
Štítky
Genetika Reprodukční medicína
Článek A Complex Genomic Rearrangement Involving the Locus Causes Dermal Hyperpigmentation in the ChickenČlánek Genome Instability and Transcription Elongation Impairment in Human Cells Depleted of THO/TREXČlánek A Population Genetics-Phylogenetics Approach to Inferring Natural Selection in Coding SequencesČlánek Interspecific Sex in Grass Smuts and the Genetic Diversity of Their Pheromone-Receptor SystemČlánek Genomic Distribution and Inter-Sample Variation of Non-CpG Methylation across Human Cell Types
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2011 Číslo 12- Akutní intermitentní porfyrie
- IVF a rakovina prsu – zvyšují hormony riziko vzniku rakoviny?
- Souvislost haplotypu M2 genu pro annexin A5 s opakovanými reprodukčními ztrátami
- Vliv dlouhodobě užívané hormonální antikoncepce na schopnost otěhotnět
- Farmakogenetické testování pomáhá předcházet nežádoucím efektům léčiv
-
Všechny články tohoto čísla
- The Connection between Space and Thinking: An Interview with Rafael Viñoly
- An Assessment of the Individual and Collective Effects of Variants on Height Using Twins and a Developmentally Informative Study Design
- Widespread Cotranslational Formation of Protein Complexes
- Genomes Reveal Transition of Bacteria from Aquatic to Terrestrial Environments
- A Complex Genomic Rearrangement Involving the Locus Causes Dermal Hyperpigmentation in the Chicken
- Plasticity of BRCA2 Function in Homologous Recombination: Genetic Interactions of the PALB2 and DNA Binding Domains
- Transcription Is Required to Establish Maternal Imprinting at the Prader-Willi Syndrome and Angelman Syndrome Locus
- Substitutions in the Amino-Terminal Tail of Neurospora Histone H3 Have Varied Effects on DNA Methylation
- MAPK/ERK Signaling Regulates Insulin Sensitivity to Control Glucose Metabolism in
- A Comprehensive Analysis of Shared Loci between Systemic Lupus Erythematosus (SLE) and Sixteen Autoimmune Diseases Reveals Limited Genetic Overlap
- Genome Instability and Transcription Elongation Impairment in Human Cells Depleted of THO/TREX
- Genome-Wide Meta-Analysis of Five Asian Cohorts Identifies as a Susceptibility Locus for Corneal Astigmatism
- A Population Genetics-Phylogenetics Approach to Inferring Natural Selection in Coding Sequences
- HIF-1 Regulates Iron Homeostasis in by Activation and Inhibition of Genes Involved in Iron Uptake and Storage
- Ror2 Enhances Polarity and Directional Migration of Primordial Germ Cells
- DNA Methylation of the Gonadal Aromatase () Promoter Is Involved in Temperature-Dependent Sex Ratio Shifts in the European Sea Bass
- A Genetic Screening Strategy Identifies Novel Regulators of the Proteostasis Network
- Interspecific Sex in Grass Smuts and the Genetic Diversity of Their Pheromone-Receptor System
- The Synthetic Multivulva Genes Prevent Ras Pathway Activation by Tightly Repressing Global Ectopic Expression of EGF
- Mining the Allelic Spectrum Reveals the Contribution of Rare and Common Regulatory Variants to HDL Cholesterol
- Identification of a Genomic Reservoir for New Genes in Primate Genomes
- Genomic Distribution and Inter-Sample Variation of Non-CpG Methylation across Human Cell Types
- Identification of Evolutionarily Conserved Exons as Regulated Targets for the Splicing Activator Tra2β in Development
- Acute Multiple Organ Failure in Adult Mice Deleted for the Developmental Regulator Wt1
- Age-Related Neuronal Degeneration: Complementary Roles of Nucleotide Excision Repair and Transcription-Coupled Repair in Preventing Neuropathology
- Target Site Recognition by a Diversity-Generating Retroelement
- Ancestral Components of Admixed Genomes in a Mexican Cohort
- Targeted Proteolysis of Plectin Isoform 1a Accounts for Hemidesmosome Dysfunction in Mice Mimicking the Dominant Skin Blistering Disease EBS-Ogna
- Autosomal Recessive Dilated Cardiomyopathy due to Mutations Results from Abnormal Dystroglycan O-Mannosylation
- SREBP Coordinates Iron and Ergosterol Homeostasis to Mediate Triazole Drug and Hypoxia Responses in the Human Fungal Pathogen
- The RNA Silencing Enzyme RNA Polymerase V Is Required for Plant Immunity
- An Anti-Checkpoint Activity for Rif1
- The FGFR4-G388R Polymorphism Promotes Mitochondrial STAT3 Serine Phosphorylation to Facilitate Pituitary Growth Hormone Cell Tumorigenesis
- Common Variants Show Predicted Polygenic Effects on Height in the Tails of the Distribution, Except in Extremely Short Individuals
- The Fission Yeast Stress-Responsive MAPK Pathway Promotes Meiosis via the Phosphorylation of Pol II CTD in Response to Environmental and Feedback Cues
- Integrating Genome-Wide Genetic Variations and Monocyte Expression Data Reveals -Regulated Gene Modules in Humans
- Repetitive Elements May Comprise Over Two-Thirds of the Human Genome
- A Novel Checkpoint and RPA Inhibitory Pathway Regulated by Rif1
- Hierarchical Generalized Linear Models for Multiple Groups of Rare and Common Variants: Jointly Estimating Group and Individual-Variant Effects
- The Major Roles of DNA Polymerases Epsilon and Delta at the Eukaryotic Replication Fork Are Evolutionarily Conserved
- A High-Resolution Whole-Genome Map of Key Chromatin Modifications in the Adult
- A Densely Interconnected Genome-Wide Network of MicroRNAs and Oncogenic Pathways Revealed Using Gene Expression Signatures
- A Functional Phylogenomic View of the Seed Plants
- Histone H3K9 Trimethylase Eggless Controls Germline Stem Cell Maintenance and Differentiation
- Ribosomal Protein Mutants Control Tissue Growth Non-Autonomously via Effects on the Prothoracic Gland and Ecdysone
- , , and Are Required to Activate or Delimit the Spread of the Transcriptional Response to Epidermal Wounds in
- Mechanisms Establishing TLR4-Responsive Activation States of Inflammatory Response Genes
- Candidate Gene Screen in the Red Flour Beetle Reveals as Ancient Regulator of Anterior Median Head and Central Complex Development
- Charcot-Marie-Tooth–Linked Mutant GARS Is Toxic to Peripheral Neurons Independent of Wild-Type GARS Levels
- The RNA–Methyltransferase Misu (NSun2) Poises Epidermal Stem Cells to Differentiate
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Targeted Proteolysis of Plectin Isoform 1a Accounts for Hemidesmosome Dysfunction in Mice Mimicking the Dominant Skin Blistering Disease EBS-Ogna
- The RNA Silencing Enzyme RNA Polymerase V Is Required for Plant Immunity
- The FGFR4-G388R Polymorphism Promotes Mitochondrial STAT3 Serine Phosphorylation to Facilitate Pituitary Growth Hormone Cell Tumorigenesis
- Target Site Recognition by a Diversity-Generating Retroelement
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Revma Focus: Spondyloartritidy
nový kurz
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MUDr. Irena Krčmová, CSc.
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



