-
Články
- Vzdělávání
- Časopisy
Top články
Nové číslo
- Témata
- Kongresy
- Videa
- Podcasty
Nové podcasty
Reklama- Kariéra
Doporučené pozice
Reklama- Praxe
Meta-Analysis of 28,141 Individuals Identifies Common Variants within Five New Loci That Influence Uric Acid Concentrations
Elevated serum uric acid levels cause gout and are a risk factor for cardiovascular disease and diabetes. To investigate the polygenetic basis of serum uric acid levels, we conducted a meta-analysis of genome-wide association scans from 14 studies totalling 28,141 participants of European descent, resulting in identification of 954 SNPs distributed across nine loci that exceeded the threshold of genome-wide significance, five of which are novel. Overall, the common variants associated with serum uric acid levels fall in the following nine regions: SLC2A9 (p = 5.2×10−201), ABCG2 (p = 3.1×10−26), SLC17A1 (p = 3.0×10−14), SLC22A11 (p = 6.7×10−14), SLC22A12 (p = 2.0×10−9), SLC16A9 (p = 1.1×10−8), GCKR (p = 1.4×10−9), LRRC16A (p = 8.5×10−9), and near PDZK1 (p = 2.7×10−9). Identified variants were analyzed for gender differences. We found that the minor allele for rs734553 in SLC2A9 has greater influence in lowering uric acid levels in women and the minor allele of rs2231142 in ABCG2 elevates uric acid levels more strongly in men compared to women. To further characterize the identified variants, we analyzed their association with a panel of metabolites. rs12356193 within SLC16A9 was associated with DL-carnitine (p = 4.0×10−26) and propionyl-L-carnitine (p = 5.0×10−8) concentrations, which in turn were associated with serum UA levels (p = 1.4×10−57 and p = 8.1×10−54, respectively), forming a triangle between SNP, metabolites, and UA levels. Taken together, these associations highlight additional pathways that are important in the regulation of serum uric acid levels and point toward novel potential targets for pharmacological intervention to prevent or treat hyperuricemia. In addition, these findings strongly support the hypothesis that transport proteins are key in regulating serum uric acid levels.
Published in the journal: . PLoS Genet 5(6): e32767. doi:10.1371/journal.pgen.1000504
Category: Research Article
doi: https://doi.org/10.1371/journal.pgen.1000504Summary
Elevated serum uric acid levels cause gout and are a risk factor for cardiovascular disease and diabetes. To investigate the polygenetic basis of serum uric acid levels, we conducted a meta-analysis of genome-wide association scans from 14 studies totalling 28,141 participants of European descent, resulting in identification of 954 SNPs distributed across nine loci that exceeded the threshold of genome-wide significance, five of which are novel. Overall, the common variants associated with serum uric acid levels fall in the following nine regions: SLC2A9 (p = 5.2×10−201), ABCG2 (p = 3.1×10−26), SLC17A1 (p = 3.0×10−14), SLC22A11 (p = 6.7×10−14), SLC22A12 (p = 2.0×10−9), SLC16A9 (p = 1.1×10−8), GCKR (p = 1.4×10−9), LRRC16A (p = 8.5×10−9), and near PDZK1 (p = 2.7×10−9). Identified variants were analyzed for gender differences. We found that the minor allele for rs734553 in SLC2A9 has greater influence in lowering uric acid levels in women and the minor allele of rs2231142 in ABCG2 elevates uric acid levels more strongly in men compared to women. To further characterize the identified variants, we analyzed their association with a panel of metabolites. rs12356193 within SLC16A9 was associated with DL-carnitine (p = 4.0×10−26) and propionyl-L-carnitine (p = 5.0×10−8) concentrations, which in turn were associated with serum UA levels (p = 1.4×10−57 and p = 8.1×10−54, respectively), forming a triangle between SNP, metabolites, and UA levels. Taken together, these associations highlight additional pathways that are important in the regulation of serum uric acid levels and point toward novel potential targets for pharmacological intervention to prevent or treat hyperuricemia. In addition, these findings strongly support the hypothesis that transport proteins are key in regulating serum uric acid levels.
Introduction
Uric acid (UA) is the final catabolic, heterocyclic purine derivative resulting from the oxidation of purines in humans. Due to the loss of hepatic uricase activity during human evolution, UA is excreted as such and is not further metabolized into carbon dioxide and ammonia. A major mechanism underlying hyperuricemia is impaired renal excretion of urate. Most notably, UA is causally involved in the pathogenesis of gouty arthritis that results from deposition of monosodium urate crystals in the joints [1]. Increased UA concentrations have been implicated in cardiovascular disease for more than five decades [2]. In addition, elevated urate is associated with obesity, blood pressure and insulin resistance, and consequently with the metabolic syndrome and type 2 diabetes [2],[3]. However, UA also has a positive role as an antioxidant, and is correlated with longevity in mammals [4]. Thus, human physiology is especially sensitive to the precise range of UA levels.
Besides environmental factors, there is evidence for a strong genetic influence upon serum UA concentrations, with heritability estimates of up to 73% [5]. Recently, genome-wide association (GWA) studies have identified single nucleotide polymorphisms (SNPs) in the SLC2A9 gene (solute carrier family 2, member 9 gene), a putative glucose transporter, which are strongly associated with serum UA concentrations and gout [6]–[9]. This novel gene locus functions as a high-capacity urate transporter in humans [8],[10]. This emphasises the power of GWA studies in expanding our understanding at the molecular level of disease mechanisms and in pointing to innovative therapeutic strategies.
The power of GWA studies to detect common variants with modest effects directly depends on the size of the study group. Therefore, the present study sought to detect novel genetic variants related to serum UA levels by conducting a meta-analysis of GWA findings from 14 studies (BRIGHT, CoLaus, CROATIA, Health 2000, KORA F3, KORA S4, ORCADES, PROCARDIS, NSPHS, SardiNIA, SHIP, SSAGA, MICROS, and TwinsUK) totalling 28,141 participants. In addition, the meta-analysis was performed independently on sex specific GWA results to address the pronounced gender differences in the regulation of UA concentrations that have previously been reported [1],[6]. Identified variants were further analyzed for association with metabolite profiles.
Results
The sample size and participant characteristics for each participating study are shown in Table S1. Meta-analysis of GWA data of 28,141 individuals of European ancestry yielded 954 SNPs (full list is provided in Table S4) that exceeded the genome-wide significance threshold of 5×10−8 (Figure 1A).
Fig. 1. Genome-wide association results. 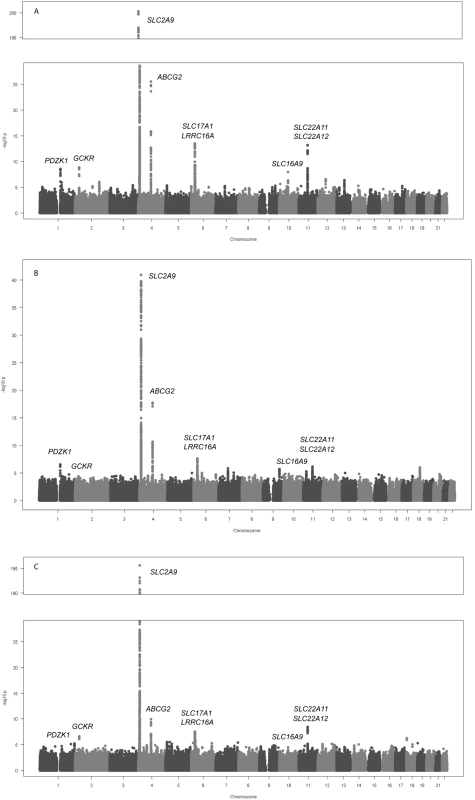
Manhattan plots showing significance of association of all SNPs in the meta-analysis for (A) men and women combined, (B) men only, and (C) women only. SNPs are plotted on the x-axis according to their position on each chromosome against association with uric acid concentrations on the y-axis (shown as −log10 p-value). Those SNPs cluster around nine loci (Table 1), four of which are well known regulators of serum UA levels: SLC2A9 (p = 5.2×10−201), ABCG2 (p = 3.1×10−26), SLC17A1 (p = 3.0×10−14), and SLC22A12 (p = 2.0×10−9). The first, SLC2A9, was identified in previous GWA scans (Figure 2C) [6]–[9]. A total of 788 SNPs reached the genome-wide significance threshold at the SLC2A9 locus. The strongest associated marker was the intronic SNP rs734553 (p = 5.2×10−201, Table 1), which is in high linkage disequilibrium (r2 = 0.88) with the missense mutation rs16890979 previously described [11]. The second locus was on chromosome 4q22, harbouring the ABCG2 gene (Figure 2D). In accordance with previous results, the strongest observed association was at rs2231142 (p = 3.1×10−26, Table 1), a coding SNP leading to a glutamine-to-lysine amino acid change at position 141 [11]. The third previously implicated locus influencing UA levels was on chromosome 6p23-p21.3, which contains the SLC17A3 gene (Figure 2F) [11]. The top associated marker was SNP rs1183201 (p = 3.0×10−14, Table 1), intronic of SLC17A1, but the association signal encompassed a larger region including the SLC17A1, SLC17A3, SLC17A4 genes and downstream to HIST1H4C, in agreement with the linkage disequilibrium at this locus. SNP rs1183201 is in high linkage disequilibrium (r2 = 0.97) with rs1165205, a SNP intronic of SLC17A3 gene identified by a previous GWA scan [11].
Fig. 2. Regional association plots of nine urate loci. 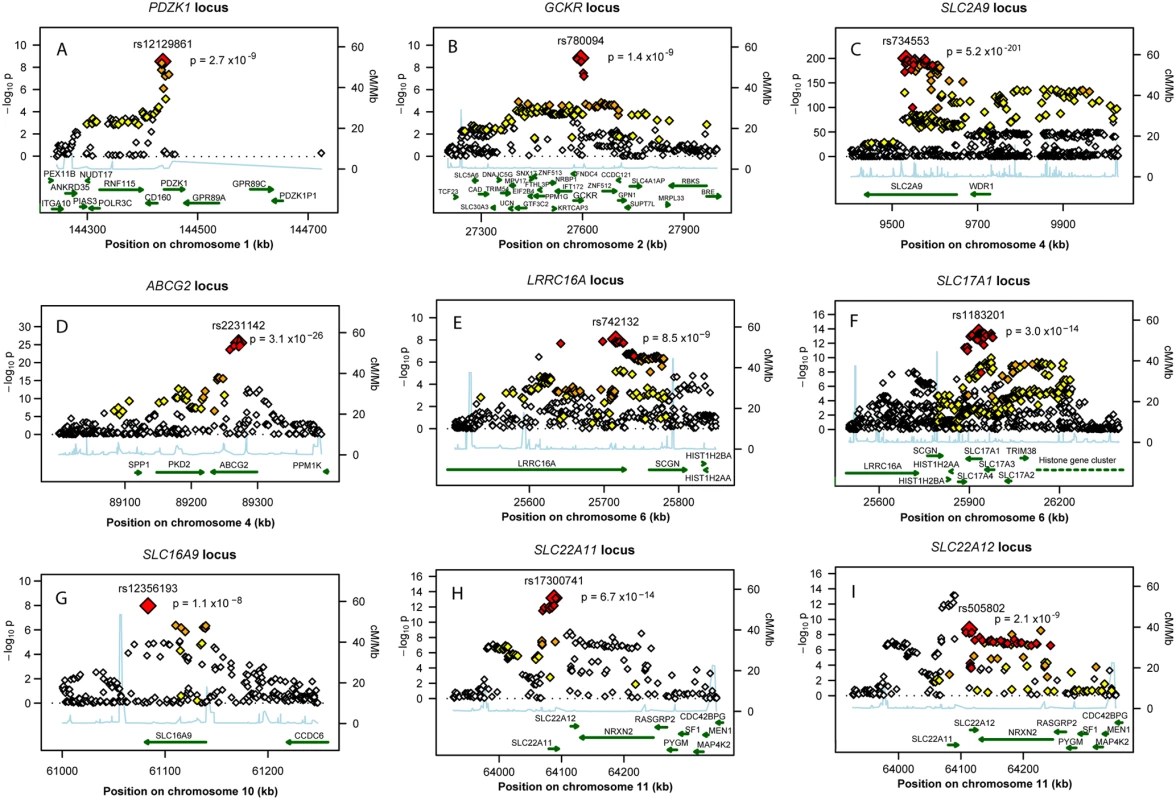
P-value plots showing the association signals in the region of (A) PDZK1 on chromosome 1, (B) GCKR on chromosome 2, (C) SLC2A9 on chromosome 4, (D) ABCG2 on chromosome 4, (E) LRRC16A on chromosome 6, (F) SLC17A1 on chromosome 6, (G) SLC16A9 on chromosome 10, (H) SLC22A11 on chromosome 11, and (I) SLC22A12 on chromosome 11. −log10 p-values are plotted as a function of genomic position (NCBI Build 36). Large diamonds in red indicate the most significant SNP in the region while other SNPs in the region are given as colour-coded smaller diamonds. Red diamonds indicate high correlation with the lead SNP (r2>0.8), orange diamonds indicate moderate correlation with the most significant SNP (0.5<r2<0.8), yellow indicates markers in weak correlation with the most significant SNP (0.2<r2<0.5), white indicates no correlation with the most significant SNP (r2<0.2). Estimated recombination rates (HapMap Phase II) are given in light blue, genes as well as the direction of transcription (NCBI) are displayed by green bars. Tab. 1. Nine loci associated with uric acid concentrations. 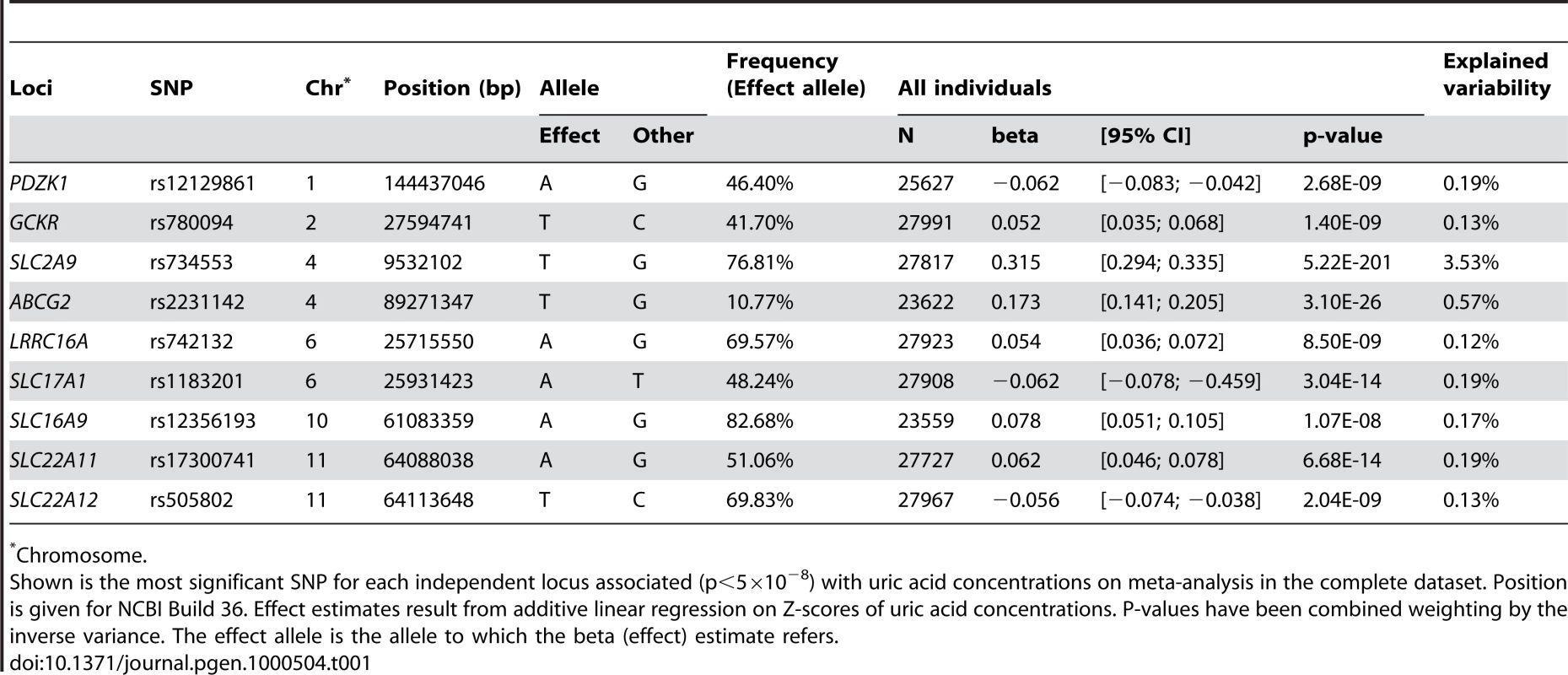
Chromosome. Among the novel loci identified, the strongest was on chromosome 11q13. One locus was localized upstream and within the SLC22A11 gene, and represented by SNP rs17300741 (p = 6.7×10−14, Table 1, Figure 2H). The second signal was SNP rs505802 (p = 2.0×10−9), representative of all associated markers falling within and downstream the extensively studied SLC22A12 gene coding for URAT1 (Figure 2I). The p-value plot as well as the LD block structure (r2 = 0.09) suggested two nearby but independently associated regions, which was verified in multiple regression analysis (Table S5).
The second novel region was on chromosome 2p23.3-p23.2 (Figure 2B). The most significant SNP in this region was SNP rs780094 (p = 1.4×10−9), intronic of GCKR, a glucokinase regulator protein recently associated with several quantitative traits including the regulation of triglycerides levels [12]. We also identified genome-wide significant association on chromosome 1q21 (Figure 2A). The top ranking SNP in this region was rs12129861 (p = 2.7×10−9, Table 1), located 2 kb upstream of PDZK1 coding for PDZ domain-containing 1 reported to interact with URAT1 [13]. The fourth newly detected region was found on chromosome 6p22.2 (Figure 2E), where the association signal spans two genes, LRRC16A and SCGN, within one highly preserved LD block. The strongest p-value was observed for the SNP rs742132, located within an intron of LRRC16A (p = 8.5×10−9, Table 1). Independence of the LRRC16A and the SLC17A1 loci (r2 = 0.07) was verified in multiple regression analysis. P-values and effect estimates only slightly changed between single SNP analysis and multiple regression analysis (Table S5). Finally, we also observed some evidence of association on chromosome 10q21.3 (Figure 2G). One SNP within SLC16A9, rs12356193, reached genome-wide significance (p = 1.1×10−8). However, there were several additional SNPs within this gene with borderline significance, supporting the hypothesis that this locus may be a true signal rather than a false positive result.
Sex-Stratified Meta-Analysis Identifies Male and Female Specific Variants
We have also performed a meta-analysis of sex specific GWA results using all 14 studies (12,328 males, 15,813 females). Although the results did not show any additional genome-wide significant locus (Figure 1B and 1C), we were able to query which of the aforementioned SNPs have sex-specific effects on serum UA levels (Table 2). For the SLC2A9 gene, we found that in males the top ranking SNP was still rs734553 (p = 1.1×10−41), while for women it was the nearby intronic SNP rs12498742 (p = 2.4×10−196). Supporting previously reported results, we found for both markers that the effect size of the minor allele observed in women was twice the effect size observed in men (p<3.8×10−17, Table 2) [6]. The minor allele of rs2231142 in the ABCG2 gene showed a greater effect size in men compared to women (p = 0.01, Table 2). Similar, the effect size of the most significant SNP for males in the ABCG2 gene locus, rs2199936, was greater in men compared to women (p = 0.008, Table 2). The effect sizes of the other SNPs were comparable in men and women (Table 2).
Tab. 2. Gender specific association results at the nine loci. 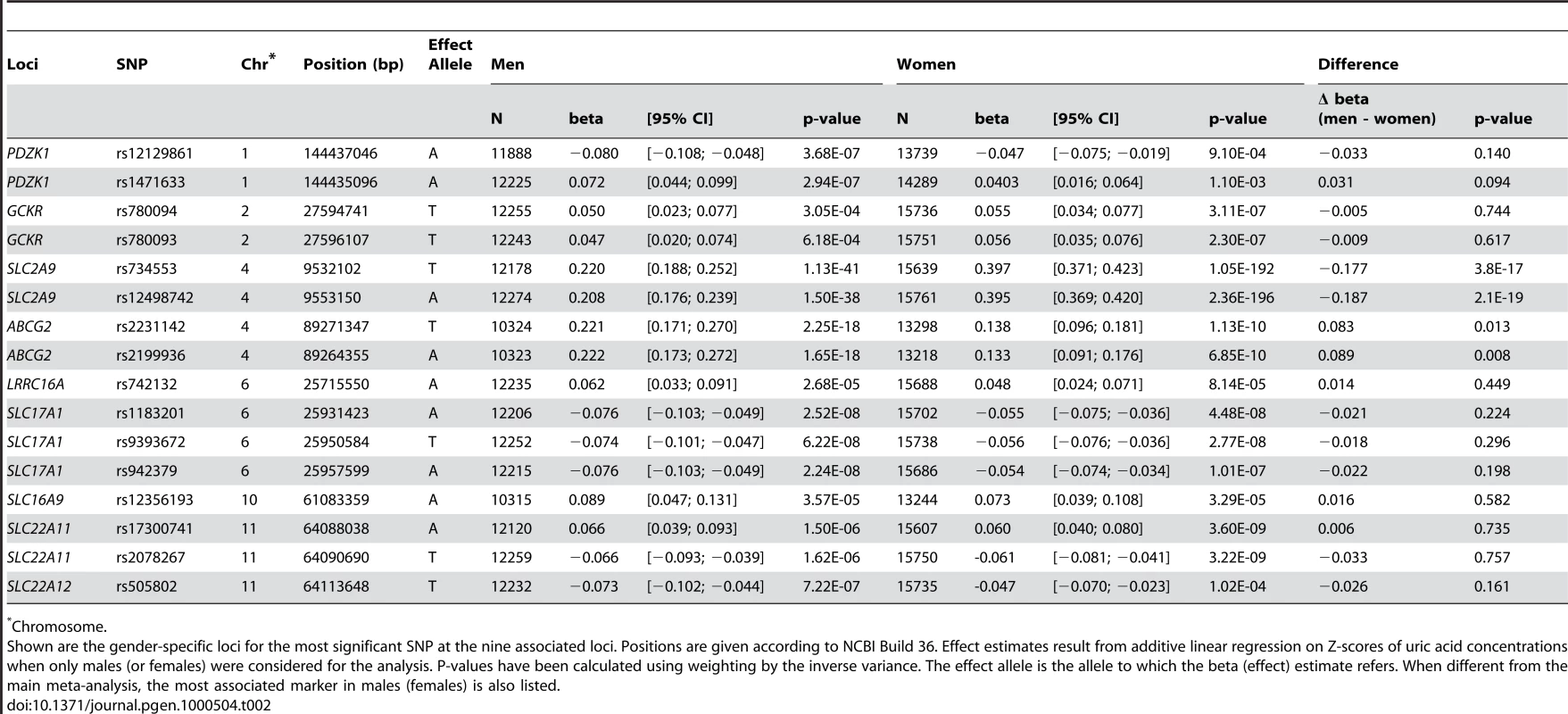
Chromosome. Association of Identified Variants with Metabolite Profiles
To further characterize the identified variants, we analyzed their association with a panel of 163 metabolites measured in the KORA F4 survey. After correction for multiple testing, one SNP within SLC16A9, rs12356193, was associated with DL-carnitine concentrations (β = −3.58, p = 4.0×10−26), which in turn were associated with serum UA levels (β = 0.06, p = 1.4×10−57). In addition, this SNP was associated with propionyl-L-carnitine (β = −0.06, p = 5.0×10−8). Similar to DL-carnitine, propionyl-L-carnitine concentrations were also strongly associated with serum UA levels (β = 1.78, p = 8.1×10−54), forming a triangle between SNP, metabolites and UA levels. None of the other SNPs were significantly associated with the measured metabolites.
Discussion
Based on meta-analysis of GWA studies including 28,141 individuals, we have mapped 5 novel loci and confirmed 4 previously implicated loci that influence serum UA levels. Altogether, these associations highlight biological pathways that are important in the regulation of urate concentrations and may point to novel targets for pharmacological interventions to prevent or treat hyperuricemia.
A genome-wide significant p-value was observed for one SNP within SLC16A9 gene locus, coding for monocarboxylic acid transporter 9 (MCT9). This is a member of the monocarboxylate co-transporter family that has been demonstrated to catalyze transport of monocarboxylic acids across cell membranes [14]. MCT9 is expressed in various tissues including the kidney [15]. As other sodium monocarboxylate transporters have been found to influence urate in knockout models this MCT9 isoform might be a sodium-dependent transporter in the kidney. The second newly identified locus was GCKR (glucokinase regulatory protein) a regulator of glucokinase, the first glycolytic enzyme which serves as a glucose sensor, responsible for glucose phosphorylation in the liver. Recently, GWA studies for type 2 diabetes identified the same rs780094 SNP as a potential marker for modulation of triglyceride levels [16]. Meanwhile, GCKR polymorphisms were also shown to be associated with metabolic traits like fasting glucose and, modestly, type 2 diabetes [12],[17],[18]. Several potential mechanisms have been proposed to link serum UA concentrations with metabolic traits. Exogenous insulin decreases renal sodium and urate excretion [19]. Furthermore, renal clearance of UA is inversely related to the degree of insulin resistance [20]. Finally, insulin resistance is thought to be accompanied by impaired oxidative phosphorylation in hepatic mitochondria, leading to increased concentrations of co-enzyme A esters and thus to increased systemic adenosine concentrations [21]. Increased adenosine, in turn, may result in renal retention of sodium, urate, and water [21],[22]. This provides a putative mechanism for hyperuricaemia via both the break down of adenosine to urate and increased renal urate retention [21],[22].
We also found evidence for association in a region containing two genes, LRRC16A and SCGN. The strongest association was located within LRRC16A coding for CARMIL. This large protein is most abundant in kidney and epithelial tissues and serves as an inhibitor of the heterodimeric actin capping protein (CP), an essential element of the actin cytoskeleton which binds to the barbed ends of actin filaments and regulates their polymerization [23]. The multiple biochemical functions associated with CARMIL raise many possibilities for its mechanism of action in cells, but the relation of CARMIL to UA concentration is thus far unclear. The nearby SCGN is coding for Secretagogin, a calcium-binding protein selectively expressed in neuroendocrine tissue and pancreatic beta-cells. The function of Secretagogin is unknown, but it has been suggested to influence calcium influx and insulin secretion [24].
We also demonstrated association of SNPs in SLC22A11 and SLC22A12 with UA concentrations. SLC22A12 encodes the extensively studied URAT1, a member of the organic anion transporter (OAT) family [25]. URAT1, a well known candidate gene for UA accumulation/transport, mediates the non-voltage-dependent exchange of urate for several organic anions [1]. SLC22A11 codes for OAT4, an OAT isoform which, like URAT1, is localized at the apical membrane of the proximal tubules. OAT4 serves as an organic anion–dicarboxylate exchanger, which mediates urate transport across the apical membrane of kidney [26],[27]. In combination with these findings, we also identified genome-wide significant association of SNPs in and upstream of PDZK1, coding for PDZ domain containing 1, a scaffolding protein reported to interact with OAT4, URAT1 and NTP1 (SLC17A1) via their C-terminal PDZ motifs [13],[28]. It has been proposed that the PDZ scaffold may form a bidirectional transport system by linking URAT1 (reabsorption) and NPT1 (secretion) leading to a functional complex responsible for the balanced urate transport regulation at the apical membrane of renal proximal tubules [1],[28].
In accordance with previous genome-wide studies, the strongest effect on serum UA concentrations was detected for SLC2A9, [6]–[9] coding for GLUT9, which has been shown to be strongly associated with hyperuricemia and gout and to serve as a high-capacity urate transporter in humans [8],[10]. Additional confirmed loci include ABCG2 and SLC17A1 [11]. ABCG2 is a member of the ATP-binding cassette (ABC) superfamily of membrane transporters, while the SLC17A1 locus, located directly downstream of the recently identified SLC17A3 locus (NPT4), encodes NPT1 (renal sodium phosphate transport protein 1). The human NPT1 is localized at the apical membrane of renal proximal tubules and serves as a voltage-driven UA transporter in model systems [28].
Although several of the SNPs associated with uric acid concentrations in this meta-analysis are located within genes that are plausible candidates for influencing uric acid concentrations, our association approach is not able to identify underlying genes or mechanisms in the regions of association signals. Therefore, other genes might be responsible for the observed associations and functional studies are warranted to identify the causal variants and provide insights in the underlying biological mechanisms.
Pronounced gender differences in the regulation of serum UA concentrations have been reported for both humans and animals [1],[6]. In line with our previous findings, [6] the strongest gender-specific effect was observed for the minor allele of rs734553 (SLC2A9), resulting in a 2-fold larger effect size on serum UA concentrations in women compared to men. For ABCG2, the effect of the minor allele of rs2231142 demonstrated a larger effect on UA concentrations in men compared to women. For the other loci, effect sizes did not significantly differ by gender.
The rapidly evolving field of metabolomics aims at a comprehensive measurement of endogenous metabolites in a cell or body fluid [29]. Based on screening of 163 metabolites, we have observed an association of one of the identified variants, rs12356193 within SLC16A9, with DL-carnitine and propionyl-L-carnitine. Moreover, DL-carnitine and propionyl-L-carnitine were strongly correlated with serum UA levels, forming a triangle between SNP, metabolites and UA levels. Carnitine is acquired from diet and endogenous biosynthesis. Its primary function is in the transport of long chain fatty acids. After strenuous physical exercise, both acylcarnitine and UA levels increase in the serum of healthy humans [30]. In spontaneously hypertensive rats, L-carnitine decreases serum UA levels and the age-dependent rise in serum UA [31],[32]. Kidneys absorb 95% of carnitine from the glomerular filtrate via an active Na+-dependent transport mechanism [33]. Impairment of this reabsorptive function can lead to carnitine deficiency, in which hyperuricemia may be present because carnitine competes for renal tubular excretion [34]. Although experimental data are few, currently available data suggest that urinary acylcarnitine, which reflects the balance between dietary intake of carnitine and renal excretion, may be linked to serum UA via oxidative stress pathways [35]. Given that palmitoyl carnitine inhibits binding of Ca2+ channel ligands to rat brain cortical membranes and to inhibit voltage-activated Ca2+ channel currents, acylcarnitines may also have direct influences on MCT9 [36].
Overall, serum UA concentrations are mainly determined by the balance between urate production and renal excretion. We have identified nine loci that are associated with serum UA levels and six of them harbor genes that code for renal transport proteins. Most notably, five of these transport proteins belong to the family–and moreover, to one phylogenetic cluster within this family [37]. These findings strongly support the hypothesis that genetic variation in urate transport proteins are the key influences upon regulation of serum UA levels in humans.
Materials and Methods
Study Participants
The present meta-analysis combined data from 14 GWA scans: British Genetics of Hypertension (BRIGHT), Cohorte Lausannoise (CoLaus), Vis Island Isolate Study (CROATIA), Health 2000 cohort (Health 2000), two surveys of the Cooperative Health Research in the Region of Augsburg (KORA F3, KORA S4), Orkney Complex Disease Study (ORCADES), Precocious Coronary Artery Disease (PROCARDIS), Northern Swedish Population Health Study (NSPHS), SardiNIA Study of Aging (SardiNIA), Study of Health in Pomerania (SHIP), Semi-Structured Assessment for Genetics of Alcoholism (SSAGA), Microisolates in South Tyrol (MICROS), and UK Adult Twin Register (TwinsUK). Altogether, the meta-analysis comprises 28,141 individuals (12,328 males, 15,813 females) of European ancestry with measured serum UA concentrations (Table S1). Approval was obtained by local ethic committees for all studies and informed consent was given from the study participants. A detailed individual description of study designs is provided in Text S1.
Genome-Wide Genotyping and Imputation
Six different platforms/arrays were used for genotyping: the Affymetrix 500 K GeneChip array (4 cohorts, n = 13,103), the Affymetrix 6.0 GeneChip array (2 cohorts, n = 5,901), Illumina HumanHap 300 (5 cohorts, n = 3,609), Illumina Human 610 K Beadchip (1 cohort, n = 2,212), Illumina HumanHap 300-Duo (1 cohort, n = 2,113), and Illumina Human 1 M beadchip (1 cohort, n = 1,203). Imputation of allele dosage of SNPs typed in the HapMap CEU population was performed using either MACH [38] or IMPUTE [39] with parameters and pre-imputation filters as specified in Table S2. All SNPs with a minor allele frequency <0.01 were excluded from analysis. SNPs were also excluded if the cohort-specific imputation quality as assessed by r2.hat (MACH) or .info (IMPUTE) metrics was <0.30 or <0.40, respectively. In total, up to 2,493,963 genotyped or imputed autosomal SNPs were analyzed.
Uric Acid Measurements
Non-fasting blood samples were obtained from study participants of BRIGHT, KORA, NSPHS, SardiNIA, SHIP and SSAGA and fasting samples from those of CoLaus, PROCARDIS, CROATIA, Health 2000, MICROS, ORCADES and TwinsUK. UA analyses were carried out on fresh samples in all studies except from BRIGHT, NSPHS, CROATIA, MICROS and SSAGA, where frozen serum was used that was stored at −20°C (BRIGHT) or −70°C (NSPHS, SSAGA, CROATIA, MICROS). UA concentrations were measured using an uricase/peroxidase method (CROATIA, MICROS, NSPHS and ORCADES: DVIA1650-Autoanalyzer, Siemens Healthcare Diagnostics) or an uricase method (BRIGHT: Hitachi, Roche Diagnostics; CoLaus: uricase PGP, Roche Diagnostics; Health 2000: Thermo Fisher Scientific, Vantaa; KORA F3: URCA Flex, Dade Behring; KORA S4: UA Plus, Roche; PROCARDIS: Hitachi 917, Roche Diagnostics; SardiNIA: Bayer; SHIP: UA PAP, Boehringer; SSAGA: Hitachi 747, Boehringer; TwinsUK: Ektachem/Vitros system, Johnson & Johnson Clinical Diagnostics).
Metabolite Measurements
Metabolomic analyses were conducted in 2020 randomly selected participants (ages 32–81 years) of the KORA F4 survey, a follow-up survey of KORA S4. Genotype information was available for 1814 of these participants. Fasting blood samples were collected in 2006–2008. Blood was drawn into serum gel tube in the morning between 8 and 10 am. The tube was gently inverted two times, followed by 30 minutes resting at room temperature to obtain complete coagulation, and finally centrifugation of blood was performed at 2750 g, 15°C for 10 minutes for serum collection. Serum was aliquoted and kept at 4°C for a maximum of 6 hours, after which it was frozen at −80°C until analyses. Liquid handling of serum samples (10 µl) was performed with Hamilton Star (Hamilton Bonaduz AG, Bonaduz, Switzerland) robot and prepared for quantification with AbsoluteIDQ kit (BIOCRATES Life Sciences AG, Innsbruck, Austria). Sample analyses were done on API4000 Q TRAP LC/MS/MS System (Applied Biosystems, Darmstadt, Germany) equipped with Schimadzu Prominence LC20AD pump and SIL-20AC auto sampler. The complete analytical process (e.g. the targeted metabolite concentration) was performed using the MetIQ software package, which is an integral part of the AbsoluteIDQ Kit. A total of 163 metabolites were measured. The metabolomics dataset contains 14 amino acids, one sugar, 41 acylcarnitines, 15 sphingolipids, and 92 glycerophospholipids.
Statistical Analysis
GWA scans were made using an additive genetic model on Z-scores, calculated by adjusting serum UA levels for age and sex using linear regression and standardizing residuals. In sex-specific association testing Z-scores were calculated in each stratum separately. Study-specific results of the most significant SNP at each locus are presented in Table S3. The results from all 14 GWA scans were combined into a fixed-effects meta-analysis with inverse variance weighting, using the METAL package (www.sph.umich.edu/csg/abecasis/metal). The individual studies were corrected for residual inflation of the test statistic using genomic control methods for genotyped and imputed SNPs combined [40]. For the overall meta-analysis, the inflation factor was 1.028, no further correction was applied. Quantile-quantile plots of the association results are shown in Figure S1, study-specific quantile-quantile plots are illustrated in Figure S2 and S3. Associations were considered genome-wide significant below p = 5×10−8, which corresponds to a Bonferroni correction for the estimated one million independent common variant tests in the human genome of European individuals [41]. We also tested whether the effect estimate resulting from the gender-specific fixed effect meta-analysis differed significantly between men and women by applying a t-test comparing effect and standard error estimates in men with the effect and standard error estimates in women. Genome-wide significant SNPs were tested for independent associations, by including all SNPs in a multiple regression model, and then performing inverse variance weighted meta-analysis, across all cohorts (except for Health 2000), of the coefficient for each SNP. The analysis of metabolites was performed using the same linear regression adjusted by sex and gender as in the genome-wide scan. To specify the dependency of uric acid on metabolite concentration, a univariate regression model without further transformation or adjustment was used. The multiple regression and metabolite analysis were performed using either posterior expected allele dosages, or on best-guess imputed genotypes, with the statistical analysis software R.
Accession Numbers
The OMIM (http://www.ncbi.nlm.nih.gov/OMIM) accession numbers for genes mentioned in this article are PDZK1 (603831), GCKR (600842), SLC2A9 (606142), ABCG2 (603756), LRRC16A (609593), SLC17A1 (182308), SLC22A11 (607097), and SLC22A12 (607096). The HGNC (http://www.gene.ucl.ac.uk) accession number for SLC16A9 is 23520.
Supporting Information
Zdroje
1. TaniguchiA
KamataniN
2008 Control of renal uric acid excretion and gout. Curr Opin Rheumatol 20 192 197
2. KoenigW
MeisingerC
2008 Uric acid, type 2 diabetes, and cardiovascular diseases: fueling the common soil hypothesis? Clin Chem 54 231 233
3. HaydenMR
TyagiSC
2004 Uric acid: A new look at an old risk marker for cardiovascular disease, metabolic syndrome, and type 2 diabetes mellitus: The urate redox shuttle. Nutr Metab (Lond) 1 10
4. CutlerRG
1984 Urate and ascorbate: their possible roles as antioxidants in determining longevity of mammalian species. Arch Gerontol Geriatr 3 321 348
5. WhitfieldJB
MartinNG
1983 Inheritance and alcohol as factors influencing plasma uric acid levels. Acta Genet Med Gemellol (Roma ) 32 117 126
6. DoringA
GiegerC
MehtaD
GohlkeH
ProkischH
2008 SLC2A9 influences uric acid concentrations with pronounced sex-specific effects. Nat Genet 40 430 436
7. LiS
SannaS
MaschioA
BusoneroF
UsalaG
2007 The GLUT9 gene is associated with serum uric acid levels in Sardinia and Chianti cohorts. PLoS Genet 3 e194 doi:10.1371/journal.pgen.0030194
8. VitartV
RudanI
HaywardC
GrayNK
FloydJ
2008 SLC2A9 is a newly identified urate transporter influencing serum urate concentration, urate excretion and gout. Nat Genet 40 437 442
9. WallaceC
NewhouseSJ
BraundP
ZhangF
TobinM
2008 Genome-wide association study identifies genes for biomarkers of cardiovascular disease: serum urate and dyslipidemia. Am J Hum Genet 82 139 149
10. CaulfieldMJ
MunroePB
O'NeillD
WitkowskaK
CharcharFJ
2008 SLC2A9 is a high-capacity urate transporter in humans. PLoS Med 5 e197 doi:10.1371/journal.pmed.0050197
11. DehghanA
KottgenA
YangQ
HwangSJ
KaoWL
2008 Association of three genetic loci with uric acid concentration and risk of gout: a genome-wide association study. Lancet 372 1953 1961
12. SparsoT
AndersenG
NielsenT
BurgdorfKS
GjesingAP
2008 The GCKR rs780094 polymorphism is associated with elevated fasting serum triacylglycerol, reduced fasting and OGTT-related insulinaemia, and reduced risk of type 2 diabetes. Diabetologia 51 70 75
13. AnzaiN
MiyazakiH
NoshiroR
KhamdangS
ChairoungduaA
2004 The multivalent PDZ domain-containing protein PDZK1 regulates transport activity of renal urate-anion exchanger URAT1 via its C terminus. J Biol Chem 279 45942 45950
14. HalestrapAP
PriceNT
1999 The proton-linked monocarboxylate transporter (MCT) family: structure, function and regulation. Biochem J 343 Pt 2 281 299
15. HalestrapAP
MeredithD
2004 The SLC16 gene family-from monocarboxylate transporters (MCTs) to aromatic amino acid transporters and beyond. Pflugers Arch 447 619 628
16. SaxenaR
VoightBF
LyssenkoV
BurttNP
de BakkerPI
2007 Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 316 1331 1336
17. VaxillaireM
Cavalcanti-ProencaC
DechaumeA
TichetJ
MarreM
2008 The common P446L polymorphism in GCKR inversely modulates fasting glucose and triglyceride levels and reduces type 2 diabetes risk in the DESIR prospective general French population. Diabetes 57 2253 2257
18. Orho-MelanderM
MelanderO
GuiducciC
Perez-MartinezP
CorellaD
2008 Common missense variant in the glucokinase regulatory protein gene is associated with increased plasma triglyceride and C-reactive protein but lower fasting glucose concentrations. Diabetes 57 3112 3121
19. Ter MaatenJC
VoorburgA
HeineRJ
Ter WeePM
DonkerAJ
1997 Renal handling of urate and sodium during acute physiological hyperinsulinaemia in healthy subjects. Clin Sci (Lond) 92 51 58
20. FacchiniF
ChenYD
HollenbeckCB
ReavenGM
1991 Relationship between resistance to insulin-mediated glucose uptake, urinary uric acid clearance, and plasma uric acid concentration. JAMA 266 3008 3011
21. ChoiHK
MountDB
ReginatoAM
2005 Pathogenesis of gout. Ann Intern Med 143 499 516
22. BakkerSJ
GansRO
Ter MaatenJC
TeerlinkT
WesterhoffHV
2001 The potential role of adenosine in the pathophysiology of the insulin resistance syndrome. Atherosclerosis 155 283 290
23. YangC
PringM
WearMA
HuangM
CooperJA
2005 Mammalian CARMIL inhibits actin filament capping by capping protein. Dev Cell 9 209 221
24. SkovhusKV
BergholdtR
ErichsenC
SparreT
NerupJ
2006 Identification and characterization of secretagogin promoter activity. Scand J Immunol 64 639 645
25. EnomotoA
KimuraH
ChairoungduaA
ShigetaY
JutabhaP
2002 Molecular identification of a renal urate anion exchanger that regulates blood urate levels. Nature 417 447 452
26. AnzaiN
EnomotoA
EndouH
2005 Renal urate handling: clinical relevance of recent advances. Curr Rheumatol Rep 7 227 234
27. EkaratanawongS
AnzaiN
JutabhaP
MiyazakiH
NoshiroR
2004 Human organic anion transporter 4 is a renal apical organic anion/dicarboxylate exchanger in the proximal tubules. J Pharmacol Sci 94 297 304
28. AnzaiN
KanaiY
EndouH
2007 New insights into renal transport of urate. Curr Opin Rheumatol 19 151 157
29. GiegerC
GeistlingerL
AltmaierE
HrabedA
KronenbergF
2008 Genetics meets metabolomics: a genome-wide association study of metabolite profiles in human serum. PLoS Genet 4 e1000282 doi:10.1371/journal.pgen.1000282
30. NozakiS
TanakaM
MizunoK
AtakaS
MizumaH
2009 Mental and physical fatigue-related biochemical alterations. Nutrition 25 51 57
31. NakazonoK
WatanabeN
MatsunoK
SasakiJ
SatoT
1991 Does superoxide underlie the pathogenesis of hypertension? Proc Natl Acad Sci U S A 88 10045 10048
32. RauchovaH
DobesovaZ
DrahotaZ
ZichaJ
KunesJ
1998 The effect of chronic L-carnitine treatment on blood pressure and plasma lipids in spontaneously hypertensive rats. Eur J Pharmacol 342 235 239
33. HuangW
ShaikhSN
GanapathyME
HopferU
LeibachFH
1999 Carnitine transport and its inhibition by sulfonylureas in human kidney proximal tubular epithelial cells. Biochem Pharmacol 58 1361 1370
34. RoschingerW
MuntauAC
DuranM
DorlandL
IJlstL
2000 Carnitine-acylcarnitine translocase deficiency: metabolic consequences of an impaired mitochondrial carnitine cycle. Clin Chim Acta 298 55 68
35. LootsDT
MienieLJ
BerghJJ
Van der SchyfCJ
2004 Acetyl-L-carnitine prevents total body hydroxyl free radical and uric acid production induced by 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP) in the rat. Life Sci 75 1243 1253
36. StapletonSR
CurrieKP
ScottRH
BellBA
1992 Palmitoyl-DL-carnitine has calcium-dependent effects on cultured neurones from rat dorsal root ganglia. Br J Pharmacol 107 1192 1197
37. FredrikssonR
NordstromKJ
StephanssonO
HagglundMG
SchiothHB
2008 The solute carrier (SLC) complement of the human genome: phylogenetic classification reveals four major families. FEBS Lett 582 3811 3816
38. LiY
AbecasisGR
2006 Mach 1.0: rapid haplotype reconstruction and missing genotype inference. Am J Hum Genet S79 2290
39. MarchiniJ
HowieB
MyersS
McVeanG
DonnellyP
2007 A new multipoint method for genome-wide association studies by imputation of genotypes. Nat Genet 39 906 913
40. BacanuSA
DevlinB
RoederK
2000 The power of genomic control. Am J Hum Genet 66 1933 1944
41. DudbridgeF
GusnantoA
2008 Estimation of significance thresholds for genomewide association scans. Genet Epidemiol 32 227 234
Štítky
Genetika Reprodukční medicína
Článek vyšel v časopisePLOS Genetics
Nejčtenější tento týden
2009 Číslo 6- Akutní intermitentní porfyrie
- Růst a vývoj dětí narozených pomocí IVF
- Vliv melatoninu a cirkadiálního rytmu na ženskou reprodukci
- Intrauterinní inseminace a její úspěšnost
- Délka menstruačního cyklu jako marker ženské plodnosti
-
Všechny články tohoto čísla
- Cytoplasmic Streaming in : Disperse the Plug To Increase the Flow?
- The Mysteries of Chromosome Evolution in Gibbons: Methylation Is a Prime Suspect
- Meta-Analysis of 28,141 Individuals Identifies Common Variants within Five New Loci That Influence Uric Acid Concentrations
- The Impact of Divergence Time on the Nature of Population Structure: An Example from Iceland
- Is a Novel Locus for Waist Circumference: A Genome-Wide Association Study from the CHARGE Consortium
- Asymmetric Strand Segregation: Epigenetic Costs of Genetic Fidelity?
- PLOS Genetics
- Archiv čísel
- Aktuální číslo
- Informace o časopisu
Nejčtenější v tomto čísle- Cytoplasmic Streaming in : Disperse the Plug To Increase the Flow?
- Meta-Analysis of 28,141 Individuals Identifies Common Variants within Five New Loci That Influence Uric Acid Concentrations
- Is a Novel Locus for Waist Circumference: A Genome-Wide Association Study from the CHARGE Consortium
- Asymmetric Strand Segregation: Epigenetic Costs of Genetic Fidelity?
Kurzy
Zvyšte si kvalifikaci online z pohodlí domova
Autoři: prof. MUDr. Vladimír Palička, CSc., Dr.h.c., doc. MUDr. Václav Vyskočil, Ph.D., MUDr. Petr Kasalický, CSc., MUDr. Jan Rosa, Ing. Pavel Havlík, Ing. Jan Adam, Hana Hejnová, DiS., Jana Křenková
Autoři: MUDr. Irena Krčmová, CSc.
Autoři: MDDr. Eleonóra Ivančová, PhD., MHA
Autoři: prof. MUDr. Eva Kubala Havrdová, DrSc.
Všechny kurzyPřihlášení#ADS_BOTTOM_SCRIPTS#Zapomenuté hesloZadejte e-mailovou adresu, se kterou jste vytvářel(a) účet, budou Vám na ni zaslány informace k nastavení nového hesla.
- Vzdělávání



